Lus10016316 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016316 pacid=23161838 polypeptide=Lus10016316 locus=Lus10016316.g ID=Lus10016316.BGIv1.0 annot-version=v1.0
ATGTTTACGGTGCGGCCTTCAAGTGATTTTGCTCGCTTGATCAAGTCCCGCAAGGTTGATGAAGGACTGCTGAAGAAGTTGATTGCGGCGATGACAACTA
TTAATCAGCTTATTGACAATGCATCGGAGAAGCAGCTTACTGATGATGCAATCAGAAACTGGGTTGCTGACCTTAAAGATGCTGCCTATGATGCCGATGA
CTTGGTTGATGAGCTTGGCTATGAACAACTGCGGTCACTGGCTGAATCTTCATCCAAAACTACCACAGGAAAGATGATAAGTTATATAAACAATCGTAAC
CCTTTCAAGGATGGGATTAATGGAAAGCTGAAGAAGCTTCTTGAGCGGCTTGATGCCGTGATAAAACTGAAGGATGTCCACGGGTTGAGTTTAGATAGTG
GAGGGAAACAGTCGTCGGTTAAAACTACTGCAGCTTCTCATGTGAATGAATCTTGTGTTTATGGCAGGGAAGGTGATAAGGAAGCTATTATGAAGTTGCT
TCTCTTGAATGACCAAACAAGTGATGAGTTAGGTGTCATTGTGATCGTGGGAACTGGTGGTGCGGGTAAAACTGCACTTTCTAAGATGATATACAATGAT
AGCAAGGTTCAAGAGCACTTTCAACTAAATGCTTGGATATGTGTTTCGGAGAAGTTTGATGCTAACAGAATTACAAAGCACATCCTTGAGGGGCTTAGCG
GTAAGGCGATTGATAATCGTACTTTAGATCAGCTACAAATAGAGCTTAAAGGAAGATTGACACAGAAAAAGTTCCTTCTAATTCTAGATGATTTGTGGAA
TGAGAATGATGGTGATTGGGATTTACTACAAACACCTCTCAAACATGGTGTAAAAGGAAGTAAGATTATAGTTACAACACGGCATCCGAATGCAGCATCA
GCACTGCCAACTAGTGAGACGTTTCATTTAAGCCCCTTGACAAATGATGATTGCTGGACGTTGTTCTCAAACTGTGCCTCTGTTGATGGAAATTTAAGTG
CAAGTCCAGAGCTGGAATCGGTTGGCAGAGAACTCATGGAGAAATGCCAGGGTCTACCTTTGTCTGCAAAAATATTAGGAGCTCTACTGCGCTCTGAACA
GGATGCTGAGAAATGGGGGAATATACTGAGGAGTACTCTATGGAATCTCCAAGACAAGATTCTTCCTGCTCTGATTTTGAGCTATTATTATCTTCCATCA
TATTTGAAACCATGCTTTGCTTACTGTGCAATATTTCCAAGAGATTATGCATTTGAAAGGGACGAACTTGTGGCAATCTGGTGGAATGTTGATTACAAAT
CGATTGACAATGATGTGCTCAATGGTCTTTTGAGTTCACTTAGCAACTTACATGTACTGTCCTTCTACGGGTACAAGTTAGTCGAGTTGCCTATTTCAAT
TGGTGGCTTAAAACACTTGCGGTATTTGAATCTCTCACTGACTTGTGTAAAAGAGTTGCCAGCAACAATCTGCATCCTTTACAATCTACAGATTTTAATT
ATGCAGTCCTGCAAGGAGCTTGAAGTATTGCCCGACTCGATTGGCAAGTTGAAGCAATTAAAACATCTAGATCTCTGCCAAACAGCAGTTGGAAGGTTGC
CGGACTCTGTGGTACTGTTAAACAACTTGTATCATCTGGACCTTACAGATACAAAATTGCAAGAGATGCCATTGCAAATAGGTAAACTGACAAAACTTCA
GTTCTTAACTGATTTTGTTGTGGGGAGTCACAGTGGTTCTACGATCAAGGAGTTGGAGGAGCTTCATCATCTCCGAGAAACACTCCGTATTTGGAATCTT
CAAAACGTTGAAGATTCGGTAGATACACTTGCAGCAGTTTTGAAGACTAAGAAGCACCTTAGGAGGTTAGAGTTAAGGTGGGATGGTGATGCTGATGACT
CATCACATGAAGTAATGCTAATGGACAAGTTGCAACCATATGTGCAGTTGAATAGCTTGAAGATTGATGGGTATAGAGGAACAACATTTCCACCATGGTT
TGGGGATTCTACATTTACAAATATAGTTGCTATGGAACTCTCAGGGTGTAAGAACTGCTACTGCTTGCCCCCACTTGGACAGTTGGTGTCTCTAGAAAAG
CTATCTATCAAAGCATTTGACAATATACGAACTGTGGGTGTGGAGTTTTGTGGGAGTGCCACATCCACAGAGAAGCCATTTAGATCTCTCAAAACGTTGG
AGTTTGAAAACATGCCAGAATGGGAGGAATGGACTCCTGTGGTAGAAAATGACGGTGGAAATTACCCTCTTCTCGTGGAGCTTTACTTGACTGATTGTCC
CAAACTGAAGCAAACGCTACCTAGCAACCTTCCTTCATTAAAAACACTTCGGTTAATGGGATGCCAGCTTCTTCGAGTTGCTTCACTTCCAAGTAATTCT
GCAGCTTTTAAAATGGAAGTGAGGGATACTTTTCGTAAGGTACTGCTCGAGACATCAACTGAGGGGATGCATTCTGTTCATGTCACTGGGTGTGGCTCCG
AAAACGCAGGAACAGATGAAATGGAGCCAGTCCATGGCCTTAGTGCCACTTTAGAAGAGATTGAGATTGCAAAATGCATTTCACTCAGCTGCTTCCCTCT
CGATTTGTTCCCCAAATTGAAAAGTCTTTCTATCGTGAACTGTCAAAGCCTGGAATCTCTGTCAAGTAAAGAGGTCATTGAGGAGGATTTGGCTTCTCTG
CGCTCTTTGAGAATTTCTGAGTGCCCGAAATTTGTCTCATTTCCAACAGGAGGGTTGCCTGCCCCAGATTTGACTGATCTTGTTTTATTGACATGCTCAA
ATCTGACTTATCTACCAGAATATATAAACGAGCTTCCATCCCTTGTGGAGTTGACAGTGGGTCACTGTCCTGAAATCGAGTTTTTCCCTGAAGGTGGTTT
GCCTTCCAAATTAGAGAAACTGGAGATCTGTGATTGTGATAAACTCATAGCTGAGCATATGCAATGGAACTTGCAGGCACTCCCCTCTCTTTTTCACTTC
AGTATTGATTCATACGAAGGACTGGAATCTTTCCCTCACGAGACACTGCTCCCTTCCACGCTCACAACACTGTCGATTTTGAGCATTCAAAATCTGAAGT
CTCTGGATAACAAAGGGATCCAACACTTAACTGCTCTTAAGGAGTTGAAGATCAGTGCTTGCCGGAGTTTGGAATCATTTCCCATAGGGAATTGGTTTCC
CACTTTGGAATCACTCGAGATTAATAGCTGCGACAAACTCATAGCCGGTCGATCACAATGGAACATACAGGCAATGACGTCTCTCTTGCACTTGAAATTG
TATGAAAATGAAGAAGTTGAATCATTTACAGGGGAGATGCTGCTACCTTCCTCCCTAATCTCCCTTGACATATGGGATCTGTATGGTCTGAAATCACTGG
AAGGACTTCAACAGCTTACTTCTCTTAGAGAATTGAAGATTTCTCGCTGCCCCGACCTTCAGACCATTCCAGAAGAGGGTCTTCCATCTTCCTTATCTTC
TCTGTCCATTCTCCGCTGCCCATTCCTGGAACAAAGGTGTCAGCAAGAGAAAGGTGAAGACTGGGCCAAGATTTCTCACATCCCCAACCTAGAAATCGGT
TGCTACAATGTTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10016316 pacid=23161838 polypeptide=Lus10016316 locus=Lus10016316.g ID=Lus10016316.BGIv1.0 annot-version=v1.0
MFTVRPSSDFARLIKSRKVDEGLLKKLIAAMTTINQLIDNASEKQLTDDAIRNWVADLKDAAYDADDLVDELGYEQLRSLAESSSKTTTGKMISYINNRN
PFKDGINGKLKKLLERLDAVIKLKDVHGLSLDSGGKQSSVKTTAASHVNESCVYGREGDKEAIMKLLLLNDQTSDELGVIVIVGTGGAGKTALSKMIYND
SKVQEHFQLNAWICVSEKFDANRITKHILEGLSGKAIDNRTLDQLQIELKGRLTQKKFLLILDDLWNENDGDWDLLQTPLKHGVKGSKIIVTTRHPNAAS
ALPTSETFHLSPLTNDDCWTLFSNCASVDGNLSASPELESVGRELMEKCQGLPLSAKILGALLRSEQDAEKWGNILRSTLWNLQDKILPALILSYYYLPS
YLKPCFAYCAIFPRDYAFERDELVAIWWNVDYKSIDNDVLNGLLSSLSNLHVLSFYGYKLVELPISIGGLKHLRYLNLSLTCVKELPATICILYNLQILI
MQSCKELEVLPDSIGKLKQLKHLDLCQTAVGRLPDSVVLLNNLYHLDLTDTKLQEMPLQIGKLTKLQFLTDFVVGSHSGSTIKELEELHHLRETLRIWNL
QNVEDSVDTLAAVLKTKKHLRRLELRWDGDADDSSHEVMLMDKLQPYVQLNSLKIDGYRGTTFPPWFGDSTFTNIVAMELSGCKNCYCLPPLGQLVSLEK
LSIKAFDNIRTVGVEFCGSATSTEKPFRSLKTLEFENMPEWEEWTPVVENDGGNYPLLVELYLTDCPKLKQTLPSNLPSLKTLRLMGCQLLRVASLPSNS
AAFKMEVRDTFRKVLLETSTEGMHSVHVTGCGSENAGTDEMEPVHGLSATLEEIEIAKCISLSCFPLDLFPKLKSLSIVNCQSLESLSSKEVIEEDLASL
RSLRISECPKFVSFPTGGLPAPDLTDLVLLTCSNLTYLPEYINELPSLVELTVGHCPEIEFFPEGGLPSKLEKLEICDCDKLIAEHMQWNLQALPSLFHF
SIDSYEGLESFPHETLLPSTLTTLSILSIQNLKSLDNKGIQHLTALKELKISACRSLESFPIGNWFPTLESLEINSCDKLIAGRSQWNIQAMTSLLHLKL
YENEEVESFTGEMLLPSSLISLDIWDLYGLKSLEGLQQLTSLRELKISRCPDLQTIPEEGLPSSLSSLSILRCPFLEQRCQQEKGEDWAKISHIPNLEIG
CYNV
|
DESeq2's median of ratios [FLAX]
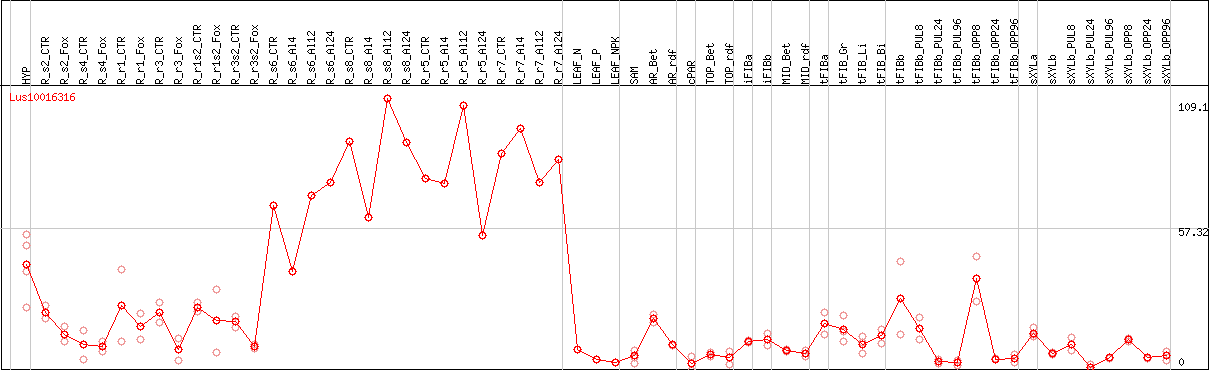
Coexpressed genes
Lus10016316 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
