External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G05530 366 / 3e-129
SDRA, IBR1
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
AT2G29150 115 / 7e-31
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G06060 109 / 1e-28
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29290 107 / 7e-28
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G29260 104 / 2e-26
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29360 103 / 2e-26
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29370 101 / 2e-25
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G07440 99 / 8e-25
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT3G26760 100 / 9e-25
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29340 97 / 4e-24
NAD-dependent epimerase/dehydratase family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015542
457 / 7e-165
AT4G05530 385 / 2e-136
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Lus10001430
401 / 6e-143
AT4G05530 369 / 2e-130
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Lus10001636
200 / 2e-65
AT4G05530 174 / 9e-56
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Lus10040735
120 / 2e-32
AT5G06060 341 / 4e-119
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016498
119 / 3e-32
AT5G06060 351 / 7e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040271
117 / 3e-31
AT5G06060 388 / 2e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040737
115 / 6e-31
AT2G29350 332 / 3e-115
senescence-associated gene 13 (.1.2.3)
Lus10016499
114 / 2e-30
AT5G06060 337 / 2e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040733
112 / 2e-29
AT5G06060 346 / 5e-121
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G022500
392 / 2e-139
AT4G05530 373 / 5e-132
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Potri.011G022400
374 / 2e-132
AT4G05530 330 / 4e-115
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Potri.006G089800
110 / 7e-29
AT2G29150 307 / 2e-105
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.014G135510
110 / 1e-28
AT4G03140 400 / 8e-141
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G074000
108 / 5e-28
AT1G52340 246 / 5e-81
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G206900
104 / 1e-26
AT2G47140 229 / 3e-75
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.010G142600
100 / 2e-25
AT2G29290 354 / 3e-124
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.006G025400
100 / 4e-25
AT3G12800 427 / 3e-152
short-chain dehydrogenase-reductase B (.1)
Potri.005G106100
99 / 1e-24
AT2G47140 234 / 9e-77
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.013G026000
99 / 2e-24
AT2G29290 362 / 2e-127
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10020019 pacid=23156141 polypeptide=Lus10020019 locus=Lus10020019.g ID=Lus10020019.BGIv1.0 annot-version=v1.0
ATGAGCAAGATCGGAAGACGATTCGAGGGTAAGGTCGCGATCGTCACCGCTTCCACTCAGGGTATTGGGTTTTCCATCGCTCAGCGAGTTGGACTTGAAG
GCGCCTCTGTTGTTATCTCCTCCCGCAAACAGCATAATGTAGATGAGGCAGTTGCGAAGCTCCGGGAACAAGGTATCCAAGCGTTGGGTGTTGTTTGCCA
TGTCTCTAATGCGGAACAGAGAAAGAATCTCATCGACAAAACTGTTGAGAAATATGGAAAAATAGATGTGGTGGTATCCAATGCTGCTGCCAACCCCTCG
ACCGAGTCCATCGTAGAAACCAAAGAATCAGTCCTCGACAAGTTGTGGGAAATCAACGTCAAAGCATCTATCCTTCTGTTGAAGGATGCAGCTCCATACA
TGCAAAGGGGTTCTTCAGTTGTTCTGGTTGCCTCAATTGCTGGCTATGACCCACCAGCGTCAATGGCTATGTATGGGGTCACGAAGACTGCCCTTCTTGG
CCTCACTAAGGCACTTGCAGCCGAGATGGCGCCTGAGGTTCGAGTAAACTGTATAGCCCCGGGTTTTGTGCCAACACATTTTGCTTCATTCCTTACCAAG
AACGATGAATTGAGGAAGTCCCTAGAAGAGAAGACCTTGCTGAATAGGTTGGGAACCACAGAAGACATGGGAGCCGCAGTAGCTTTTTTGGCATCAGACG
ATGCTTCTTACATAACTGGTGAAACATTAGTGGTGGCTGGTGGGATGTCATCTCGGCTTTAA
AA sequence
>Lus10020019 pacid=23156141 polypeptide=Lus10020019 locus=Lus10020019.g ID=Lus10020019.BGIv1.0 annot-version=v1.0
MSKIGRRFEGKVAIVTASTQGIGFSIAQRVGLEGASVVISSRKQHNVDEAVAKLREQGIQALGVVCHVSNAEQRKNLIDKTVEKYGKIDVVVSNAAANPS
TESIVETKESVLDKLWEINVKASILLLKDAAPYMQRGSSVVLVASIAGYDPPASMAMYGVTKTALLGLTKALAAEMAPEVRVNCIAPGFVPTHFASFLTK
NDELRKSLEEKTLLNRLGTTEDMGAAVAFLASDDASYITGETLVVAGGMSSRL
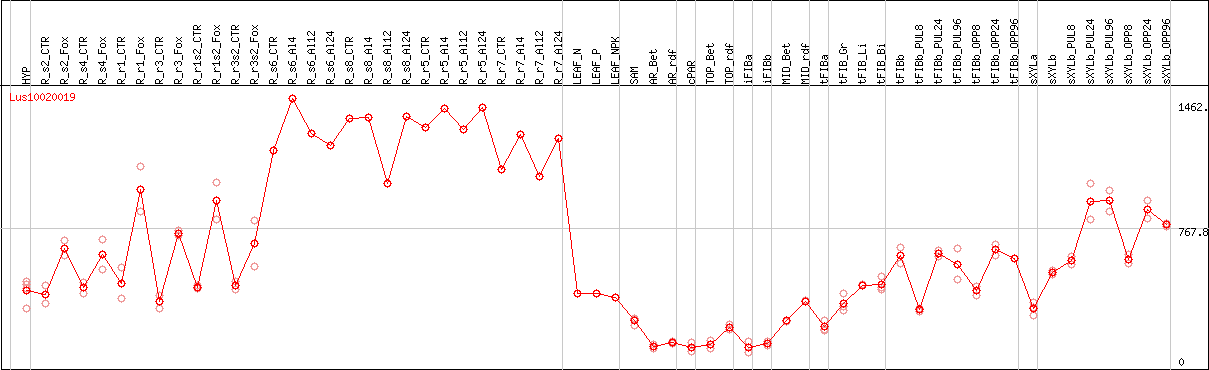
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020019 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.