External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G03980 359 / 5e-126
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G13180 355 / 9e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G18210 325 / 1e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G04000 323 / 4e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G06060 115 / 6e-31
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G24360 116 / 1e-30
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G54870 109 / 6e-28
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G07440 107 / 1e-27
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G29360 107 / 1e-27
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G52340 104 / 2e-26
SIS4, SDR1, ISI4, GIN1, ATABA2, SRE1, ABA2, ATSDR1
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002628
476 / 1e-172
AT3G03980 358 / 1e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10036980
130 / 1e-35
AT1G24360 437 / 3e-155
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10015821
127 / 3e-33
AT1G24360 427 / 5e-148
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040271
117 / 3e-31
AT5G06060 388 / 2e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10004704
114 / 3e-30
AT5G06060 387 / 3e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040736
109 / 2e-28
AT5G06060 382 / 3e-135
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016499
107 / 1e-27
AT5G06060 337 / 2e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10029368
105 / 5e-27
AT3G51680 231 / 3e-75
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040733
105 / 1e-26
AT5G06060 346 / 5e-121
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G059100
395 / 1e-140
AT3G03980 360 / 1e-126
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.019G033500
379 / 4e-134
AT3G03980 321 / 3e-111
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.013G059000
357 / 4e-125
AT4G13180 359 / 4e-126
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.019G033600
298 / 8e-102
AT3G03980 279 / 4e-94
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.010G056100
134 / 3e-37
AT1G24360 421 / 6e-149
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.008G178700
133 / 7e-37
AT1G24360 385 / 1e-134
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.012G105700
117 / 1e-31
AT2G47140 251 / 9e-84
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.015G104800
117 / 1e-31
AT2G47140 243 / 2e-80
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G039300
114 / 2e-30
AT2G29150 356 / 7e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.013G026000
114 / 3e-30
AT2G29290 362 / 2e-127
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10020265 pacid=23144036 polypeptide=Lus10020265 locus=Lus10020265.g ID=Lus10020265.BGIv1.0 annot-version=v1.0
ATGGCTACCACCGCATCCCAATCGTCTCCCTCACTCCCACTCGTCGACAAGGTGGCGATCGTCACTGGATCCTCTCGTGGCATCGGAAAAGCGATCGCTC
TCCACCTGGGATCACTCGGTGCTCGAGTCGTCGTCAATTACTCTTCCAACAAGGAACAAGCAGACCTCGTTGCCGCCGAACTCAACAAATCCTCGTCCGA
TTCCCTCCGCGCCATTTCTGTACTCGCTGACGTATCTGATCCAGCTCATATGAAGTCCTTGTTCGACGAGGCCGAGAGAGCGTTCGGGTCCGAGGTCCAT
ATCCTGGTGAATTCCGCTGGGGTGTTGGATTCGAAGTATCCTTCTGTAGCCGATTCTTCCTTGGAGGATTTCGATCGTACTTTCGGTGTGAATACTAGAG
GTGCGTTCCTATGCTGCAAGGAGGCAGCGAATCGGGTGAAACGTGGAGGTGGAGGCAGGATAATCTTGCTAACTTCGTCCTTGGTGGCTTCATTGAGACC
AGGGTTTGGGGTCTATGTTGCTTCCAAGGCTGCTGTTGAGGCGATGGTGAAGGTTCTGGCAAAGGAGCTTAAGGGAACTGGGATCACTGCCAACTGTGTG
GCGCCTGGTCCGATTGCGACTGAAATGTATTTTGCAGGGAAGTCCGAGGAGGTGATTCGTAGGAACATTGATGAATGCCCGTTAAGCCGACTTGGGGAAC
CTGAGGACATTGCTAATGTTGTTGGGTTTTTGGCTAGTGATGCTGGTGAGTGGGTCAATGGACAGGTGATTCGTGCCAATGGTGGGTATGTTTAA
AA sequence
>Lus10020265 pacid=23144036 polypeptide=Lus10020265 locus=Lus10020265.g ID=Lus10020265.BGIv1.0 annot-version=v1.0
MATTASQSSPSLPLVDKVAIVTGSSRGIGKAIALHLGSLGARVVVNYSSNKEQADLVAAELNKSSSDSLRAISVLADVSDPAHMKSLFDEAERAFGSEVH
ILVNSAGVLDSKYPSVADSSLEDFDRTFGVNTRGAFLCCKEAANRVKRGGGGRIILLTSSLVASLRPGFGVYVASKAAVEAMVKVLAKELKGTGITANCV
APGPIATEMYFAGKSEEVIRRNIDECPLSRLGEPEDIANVVGFLASDAGEWVNGQVIRANGGYV
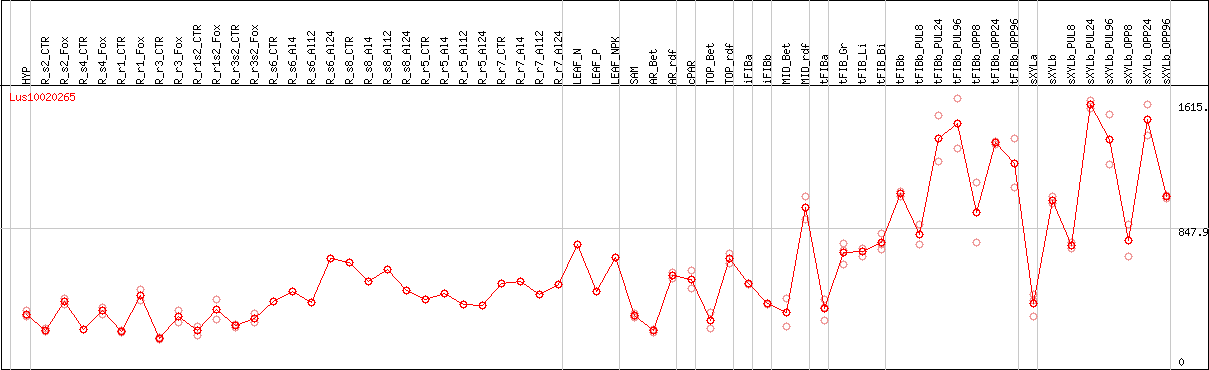
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020265 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.