Lus10022464 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10022464 pacid=23171720 polypeptide=Lus10022464 locus=Lus10022464.g ID=Lus10022464.BGIv1.0 annot-version=v1.0
ATGTCGTCTTCTTTCCTCCAATCCACTGGCTCTGCTGCTGGTGGAGCTGGATTCAACATTCTTTTCGATGCCTTTCTCAAGGCAATCCTCAATGTCTCCA
AGAAAAATACAATCTTCGAGCCCACCTTCCGCAGGCTTGAGAAGACCTTGCAGGCAATGGCTCCCGCAATCAAGAAGATCGATGTTTACAACATGGACTT
CGATCGCCCTGACGAGATTGACGGGCTCAAGGCCATACTCAAGAAAGGTGCTGAGTTGGTCTCCAAGTGCTCCAAGATTAGCCGCTATGACATCATCAGG
AAGCCCATGTACGCCAGTAAGCTTCTCAAGCTGGAAGACAAGCTTAAGAGGTACAACACCACTGTTCTTCAGGTTCATCAGGTCGCCGATCAGAAGGAGA
TTCTGTTTGAAGTCAAGCAGCTCGGAAACCAGGTCAGGCAGATGAGCATGGGGGGAGCTTCGGCTTACGGTTTTTCTCCCGGTAACTTCGGCTTCTCACC
TCCTTTCCTTAAGGTGGTTCCTGTTGGTTTGGAGATCCCATTGAAGGAACTTAAGAAAAAACTCTTGACGGATGCCCTTCCATTTCTTGTCTTGTCTGCT
CCTGGAGGATGTGGAAAGACCACATTGGCCACTGCTTTTTGCCATGATCCACAAGTTAAAGTGAGGTTTAAGGACAACATCATCTATATCACCGTCTCCA
GAGCGGCCAACCTGCTTACAATTGGGCAGAACATCTTTCAACGTTTGGGTCGCCGAGTGCCTTGTTTTCAAAGTGACGAAGATGCAGTAAACCAACTTGA
ATGTCTGCTCAAGGATATGAATGGAGACCCTATACTTTTAGTTCTTGATGATGTTTGGTCTGGAGCAGAGTTTCTTCTTGAAAGGCTCTGCTTTAACTTT
AAGATAACAAACTACCAGATATTAGTGACGTCAAGAGATGAATTTAAATCATTTGAATCTACATACAAATTACAGACATTGAGCGATGCAGATGCCATGA
AACTCTTCCGTCAGTCAGCTTTTCCTACAGATGGACGCTCCTCATATGAGCGAGATGCGGAAGTTCAGGAGAAGATAGTTAAAAGCTGCAAGGGTTTTCC
ACTACTGATTTCTGTGGTTGGCAAATCCTTACTCGGGAAACCTGCAGCAGACTGGCGTAATAGAGCCAAAGAATGTTCTAAGACTGCTTCCTTTCTTGAT
AACGATGAGGTTCTCAATTGTCTCAAGAAAAGTGTTGATGCCTTGGATAACAAACAATCTATCAAAGAGTGTTACATGGATTTAGGATTGTTCCCTGAGG
ACACTAAAATGCCTGCCACAGCCCTCCTTGATATATGGGTAGAACTACACGGGTTGGATGATGATTATGCTATGTCTAATCTCTATCAGCTCTCTTTCCG
AAATCTAGTTGACCTCGTAGTCATCAGGAAAGATGCCAGTGAAAATGATGGCTTCCACAATGAGGATTTTGTGACGCAGCATGATCTGCTGCGAGATTTA
GCTATCCGTCAAAGCAATTCCAGTAGCATTGAGGAAAGAGAAAGACTCTTTGTAGAAATAACTGGTAACAAGTTTCCTCATTGGTGGTTCGAAAGCACGG
AGAAAGTCTTTAAGAGCAAACTCTTGTCCATTACCACAGATGAAAATTTCTCAGCAAGTTGGCTTGAGATTCATGCTCCTGAAGTTGAGGTGTTGGTTCT
CAACTTTAAGACCCAGAACTATGCCTTACCCAAGTTTATTGAAAGAATGGAGAAGCTGAAGGTTCTGATAGTTACAAATCATGGGATGTGTCCAGCTAAC
TTGAGTAACATCCATCATCTTGGTTCCCTATCTAATCTTAAGAGAATTAGGCTCGAGAAGATTTGGATCCCACCAGATTTCCTTCCAGCCATTGCATTAA
GGAAGCTAGAAAAGTTAGCCTTGTTCATGTGCAACATTGGTGAAGCTTTTAGCAATGTAGACCTTGAGGCAGACTCATTCCCAAACATGGACGTGATCGA
TATAGACTACTGTAGCGATTTGATGGCGTTACCAGTTTGGTTGTTTGAATTACACCAAATGAGGAAGCTGAGCATCAGCAACTGTCAAAACCTGACAGAA
CTGCCAGAAGAACTAGGCAACTTGGGAAATTTGGAGGAGTTGAGGCTTAGTTCCTGTATAGAGTTGTCAGAGTTGCCAGAGACTATTGGAAACCTACGGA
AGTTGACGACTCTGGACATATCCGATTGTGCAAGCATAAAGAAGCTGCCTGAACAAATTGGTGAGCTTTGCAACCTGAGGAAGCTGTACATGATAGAATG
TGACCGCTGCGAACTACCGGCATCGGTAATGGAACTGAGCAAGCTGGAGGTAGTGATAGGGGATGAAGAAACAGCCAATTCATGGGAAGTTTATAAGCCT
TACCTCCCAGGTTTGAAGGTCATAAAGCACAAACAAGTCAGCTTAGACTGGCTACAATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10022464 pacid=23171720 polypeptide=Lus10022464 locus=Lus10022464.g ID=Lus10022464.BGIv1.0 annot-version=v1.0
MSSSFLQSTGSAAGGAGFNILFDAFLKAILNVSKKNTIFEPTFRRLEKTLQAMAPAIKKIDVYNMDFDRPDEIDGLKAILKKGAELVSKCSKISRYDIIR
KPMYASKLLKLEDKLKRYNTTVLQVHQVADQKEILFEVKQLGNQVRQMSMGGASAYGFSPGNFGFSPPFLKVVPVGLEIPLKELKKKLLTDALPFLVLSA
PGGCGKTTLATAFCHDPQVKVRFKDNIIYITVSRAANLLTIGQNIFQRLGRRVPCFQSDEDAVNQLECLLKDMNGDPILLVLDDVWSGAEFLLERLCFNF
KITNYQILVTSRDEFKSFESTYKLQTLSDADAMKLFRQSAFPTDGRSSYERDAEVQEKIVKSCKGFPLLISVVGKSLLGKPAADWRNRAKECSKTASFLD
NDEVLNCLKKSVDALDNKQSIKECYMDLGLFPEDTKMPATALLDIWVELHGLDDDYAMSNLYQLSFRNLVDLVVIRKDASENDGFHNEDFVTQHDLLRDL
AIRQSNSSSIEERERLFVEITGNKFPHWWFESTEKVFKSKLLSITTDENFSASWLEIHAPEVEVLVLNFKTQNYALPKFIERMEKLKVLIVTNHGMCPAN
LSNIHHLGSLSNLKRIRLEKIWIPPDFLPAIALRKLEKLALFMCNIGEAFSNVDLEADSFPNMDVIDIDYCSDLMALPVWLFELHQMRKLSISNCQNLTE
LPEELGNLGNLEELRLSSCIELSELPETIGNLRKLTTLDISDCASIKKLPEQIGELCNLRKLYMIECDRCELPASVMELSKLEVVIGDEETANSWEVYKP
YLPGLKVIKHKQVSLDWLQ
|
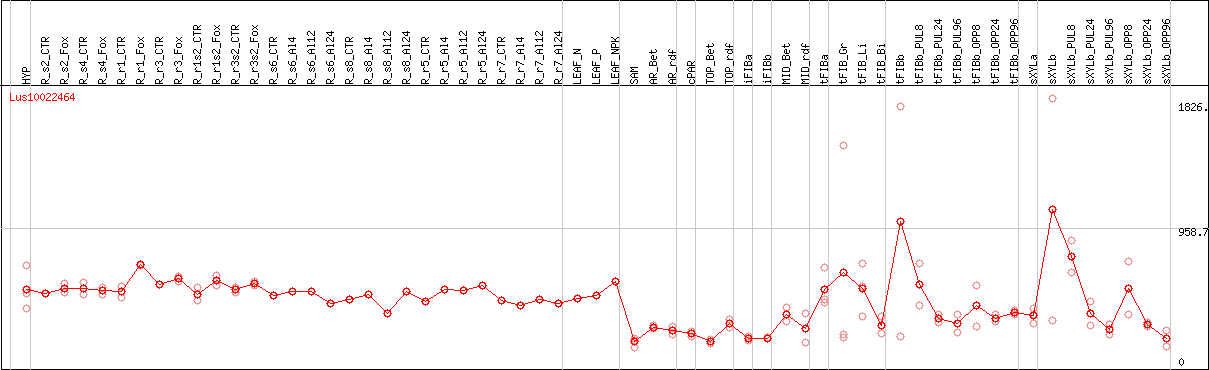
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022464 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
