External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G29350 251 / 1e-83
SAG13
senescence-associated gene 13 (.1.2.3)
AT5G06060 249 / 5e-83
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29290 246 / 9e-82
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G29150 245 / 2e-81
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29320 243 / 1e-80
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29360 240 / 3e-79
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29340 239 / 4e-79
NAD-dependent epimerase/dehydratase family protein (.1.2.3)
AT2G29300 238 / 1e-78
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G07450 236 / 9e-78
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29330 234 / 6e-77
TRI
tropinone reductase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010875
323 / 4e-112
AT5G06060 146 / 1e-42
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10010877
282 / 2e-90
AT2G29150 191 / 2e-55
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10010873
265 / 3e-89
AT5G06060 308 / 2e-106
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040271
253 / 4e-84
AT5G06060 388 / 2e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10004704
250 / 5e-83
AT5G06060 387 / 3e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040737
247 / 6e-82
AT2G29350 332 / 3e-115
senescence-associated gene 13 (.1.2.3)
Lus10004984
243 / 1e-80
AT5G06060 343 / 6e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016494
241 / 1e-79
AT5G06060 333 / 7e-116
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040736
241 / 1e-79
AT5G06060 382 / 3e-135
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G089700
277 / 6e-94
AT5G06060 320 / 6e-111
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G245000
254 / 9e-85
AT2G29150 343 / 9e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.013G026000
253 / 2e-84
AT2G29290 362 / 2e-127
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.010G142600
251 / 8e-84
AT2G29290 354 / 3e-124
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.005G039500
248 / 2e-82
AT2G29150 351 / 5e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G039300
245 / 3e-81
AT2G29150 356 / 7e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.008G059100
243 / 1e-80
AT5G06060 416 / 8e-149
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G089800
239 / 4e-79
AT2G29150 307 / 2e-105
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G244900
236 / 7e-77
AT2G29260 387 / 4e-135
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.013G026100
213 / 6e-69
AT2G29150 284 / 5e-97
NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10024358 pacid=23174889 polypeptide=Lus10024358 locus=Lus10024358.g ID=Lus10024358.BGIv1.0 annot-version=v1.0
ATGGAAGAAGTTGCAGCAGCAGCAGTAGAAGAAGCTAACAATATTGGAGAAATCAACAGGTGGTCTCTGAAGGGTATGAGTGCACTTGTGACCGGAGGAA
ACAGAGGAATTGGGTATGCAATTGTTGAGGAACTTTGCAGAAAGGGTGCCAGAGTTCACATATGCTGCCGTGGCCAGGAAGACATCGACCGGAGAATTCA
AGAATGGAAAGCCAAAGGGTTTCAAGTCAGCGGCTCTGTCTGCGACTTGACTTCCAGTTCCGAACGCCGGAAGCTCATCGACGCAGTATCCTCTGTTTTT
CATGGCTCGCTTAATATCCTCGTCAACAATGCTGGAATCACTTCCTACAACAAATTCCTGAACTGGAGCCCTGAAGATTACTCAAACGTGATGACCACAA
ACCTGGAGGCACCTTTCCATCTCTGCCAACTCGCACATCCACTCCTGACAGCATCCGGAAACGGAAGACGGAAATGGAAGCGTCGTGTTCGTCTCCTCAC
GGGAATCAACCAATTGACGAAGAACTTGGCGTTCGAATGGGCCAAAGACAACATCCGAGTCAACACCGTCGCTCCTTCTGCAACCAACACCGGGATACTT
CCATCGGGTATGGAGGAAGTTACGAAAGACTATGTAGAGGTGTGTAAGAGGATCCCAATCGGTAGAATTGCTGAATCGGACGAGGTGTCACCGTTGGTCG
CGTTCCTCTGTATGCCTGCCGCCTCTTACATATCCGGCCAAGTCATTGCCGTTGATGCTGGACTTACAGTTACTGCTGCTTAA
AA sequence
>Lus10024358 pacid=23174889 polypeptide=Lus10024358 locus=Lus10024358.g ID=Lus10024358.BGIv1.0 annot-version=v1.0
MEEVAAAAVEEANNIGEINRWSLKGMSALVTGGNRGIGYAIVEELCRKGARVHICCRGQEDIDRRIQEWKAKGFQVSGSVCDLTSSSERRKLIDAVSSVF
HGSLNILVNNAGITSYNKFLNWSPEDYSNVMTTNLEAPFHLCQLAHPLLTASGNGRRKWKRRVRLLTGINQLTKNLAFEWAKDNIRVNTVAPSATNTGIL
PSGMEEVTKDYVEVCKRIPIGRIAESDEVSPLVAFLCMPAASYISGQVIAVDAGLTVTAA
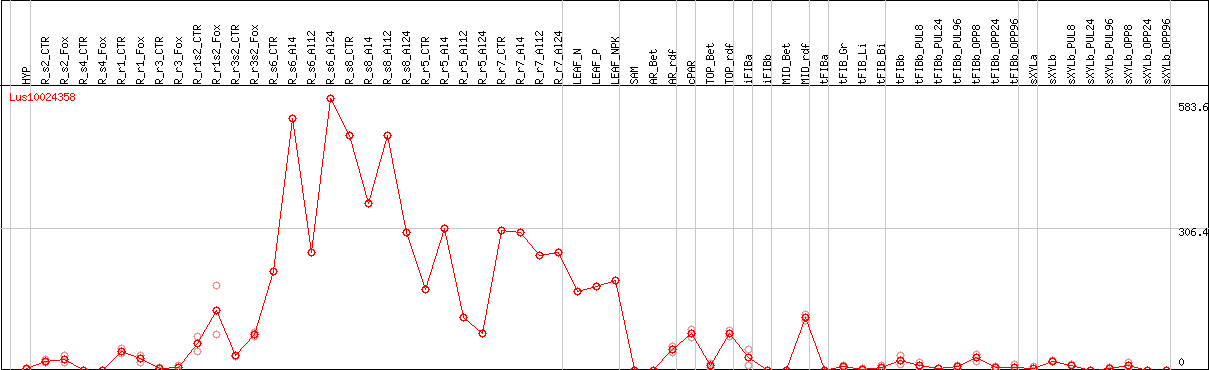
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024358 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.