External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G37630 360 / 5e-124
MYB
AtPHAN, AtMYB91, MYB91, AS1
ARABIDOPSIS PHANTASTICA-LIKE 1, MYB DOMAIN PROTEIN 91, ASYMMETRIC LEAVES 1, myb-like HTH transcriptional regulator family protein (.1)
AT5G56110 100 / 9e-24
MYB
MS188, ATMYB80, AtMYB103
MALE STERILE 188, ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 80, myb domain protein 103 (.1)
AT5G59780 97 / 2e-23
MYB
ATMYB59, ATMYB59-1, ATMYB59-3, ATMYB59-2
myb domain protein 59 (.1.2.3)
AT3G46130 97 / 2e-23
MYB
PFG3, ATMYB111 ATMYB48, ATMYB48-3, ATMYB48-1, ATMYB48-2
myb domain protein 48 (.1.2.3.4)
AT1G25340 97 / 6e-23
MYB
ATMYB116
myb domain protein 116 (.1.2)
AT1G66370 96 / 7e-23
MYB
ATMYB113
myb domain protein 113 (.1)
AT5G52600 95 / 7e-23
MYB
AtMYB82
myb domain protein 82 (.1)
AT5G14750 93 / 2e-22
MYB
WER1, WER, AtMYB66
WEREWOLF 1, WEREWOLF, myb domain protein 66 (.1)
AT3G01530 92 / 4e-22
MYB
ATMYB57
myb domain protein 57 (.1)
AT1G68320 94 / 5e-22
MYB
BW62C, BW62B, AtMYB62
myb domain protein 62 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024392
475 / 4e-169
AT2G37630 456 / 3e-161
ARABIDOPSIS PHANTASTICA-LIKE 1, MYB DOMAIN PROTEIN 91, ASYMMETRIC LEAVES 1, myb-like HTH transcriptional regulator family protein (.1)
Lus10040063
101 / 9e-24
AT2G32460 229 / 1e-69
ARABIDOPSIS THALIANA MYB 1, myb domain protein 101 (.1.2)
Lus10009037
97 / 3e-23
AT1G68320 231 / 1e-75
myb domain protein 62 (.1)
Lus10009522
98 / 5e-23
AT1G08810 284 / 9e-96
myb domain protein 60 (.1.2)
Lus10035275
100 / 6e-23
AT2G32460 233 / 5e-70
ARABIDOPSIS THALIANA MYB 1, myb domain protein 101 (.1.2)
Lus10005886
96 / 8e-23
AT3G46130 265 / 1e-89
myb domain protein 48 (.1.2.3.4)
Lus10021539
99 / 1e-22
AT2G32460 226 / 5e-67
ARABIDOPSIS THALIANA MYB 1, myb domain protein 101 (.1.2)
Lus10041435
97 / 1e-22
AT1G68320 262 / 2e-86
myb domain protein 62 (.1)
Lus10027189
98 / 2e-22
AT2G32460 243 / 1e-73
ARABIDOPSIS THALIANA MYB 1, myb domain protein 101 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G085900
388 / 1e-134
AT2G37630 414 / 2e-144
ARABIDOPSIS PHANTASTICA-LIKE 1, MYB DOMAIN PROTEIN 91, ASYMMETRIC LEAVES 1, myb-like HTH transcriptional regulator family protein (.1)
Potri.004G102600
230 / 4e-73
AT2G37630 243 / 9e-78
ARABIDOPSIS PHANTASTICA-LIKE 1, MYB DOMAIN PROTEIN 91, ASYMMETRIC LEAVES 1, myb-like HTH transcriptional regulator family protein (.1)
Potri.017G112300
196 / 1e-59
AT2G37630 188 / 2e-56
ARABIDOPSIS PHANTASTICA-LIKE 1, MYB DOMAIN PROTEIN 91, ASYMMETRIC LEAVES 1, myb-like HTH transcriptional regulator family protein (.1)
Potri.011G040400
103 / 2e-25
AT2G31180 163 / 6e-49
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 14, myb domain protein 14 (.1)
Potri.015G046200
103 / 6e-25
AT1G25340 241 / 2e-78
myb domain protein 116 (.1.2)
Potri.012G055600
102 / 2e-24
AT1G68320 244 / 1e-79
myb domain protein 62 (.1)
Potri.008G122100
100 / 5e-24
AT1G68320 274 / 1e-91
myb domain protein 62 (.1)
Potri.017G126000
95 / 2e-22
AT1G66370 188 / 5e-59
myb domain protein 113 (.1)
Potri.009G027300
94 / 2e-22
AT3G46130 279 / 3e-95
myb domain protein 48 (.1.2.3.4)
Potri.011G167600
96 / 3e-22
AT5G56110 338 / 3e-116
MALE STERILE 188, ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 80, myb domain protein 103 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10025355 pacid=23157364 polypeptide=Lus10025355 locus=Lus10025355.g ID=Lus10025355.BGIv1.0 annot-version=v1.0
ATGAAAGAGAGGCAGCGTTGGAGAGCTGAAGAGGACGCCTTACTACGTTCTTACGTCAAACGATATGGACCAAGGGAATGGAACCTCGTCTCCCAGCGCA
TGAACACACCCTTAAACCGCGACGCCAAATCCTGCTTGGAACGCTGGAAGAACTACCTCAAACCAGGAATCAAGAAAGGATCTCTCACCGAAGAAGAGCA
GCATCTAGTCATCAAACTACAAGAGAAATACGGCAACAAGTGGAAGAAAATCGCAGCCGAAGTCCCCGGAAGAACAGCCAAACGACTCGGGAAGTGGTGG
GAAGTCTTCAAGGAGAAGCAACATAGAGAGAAGAAGGAAACTAGCAAGACTCCACTCGAGCCTATCGACGAGTCCAAGTACGATCGCATCCTCGAGACCT
TCGCCGAGAAGCTGGTCAAGGAAAGAACAACACCACCTCCTCCTACAGGATTCGCTATGGCAGCTTCTAATGGAAGCAACTATCTTCACACTACTGATCA
TCATCATCTTCATCAACCTGCAATGCTGCCACCGTGGCTTTCTAGTTCCAACGGTGCAGTAAGGCCTCCTCCTGCTCCATCGGTGACATTGAGTCTTTCG
AGCTCGACTGTTGCTGCTGGTGGCGCTTCTCCATCGATACCATGGCTGCAGCCTGCTGACCTCCCCCCACCACCACCACTACCACCACCGGCAGGATTTA
GCAATGAGAGTTTGGTGGTGAACGACCTGGTGGAATGCTGCAGGGAGCTGGATGAAGGGCATCGGGCATGGGCGGCGCATAAGAAGGAAGCTGCTTGGAG
GCTGAGGAGGGTTGAGTTGCAGTTGGAGTCGGAAAAGTCATGTATGATGAAGGAGAAATCGGAGGAGATAGAGGCGAAGATAAGAGCGATGAGGGAAGAG
CAAAAGGCTGCACTCGATGCTATTGAAGAGGAGTATAGAGAGCAGCTGGCTGGGTTGAGGAGGGACGCTGAAGCTAAGGAGCAGAAATTGGCTGATCAGT
GCAGAAGTTGA
AA sequence
>Lus10025355 pacid=23157364 polypeptide=Lus10025355 locus=Lus10025355.g ID=Lus10025355.BGIv1.0 annot-version=v1.0
MKERQRWRAEEDALLRSYVKRYGPREWNLVSQRMNTPLNRDAKSCLERWKNYLKPGIKKGSLTEEEQHLVIKLQEKYGNKWKKIAAEVPGRTAKRLGKWW
EVFKEKQHREKKETSKTPLEPIDESKYDRILETFAEKLVKERTTPPPPTGFAMAASNGSNYLHTTDHHHLHQPAMLPPWLSSSNGAVRPPPAPSVTLSLS
SSTVAAGGASPSIPWLQPADLPPPPPLPPPAGFSNESLVVNDLVECCRELDEGHRAWAAHKKEAAWRLRRVELQLESEKSCMMKEKSEEIEAKIRAMREE
QKAALDAIEEEYREQLAGLRRDAEAKEQKLADQCRS
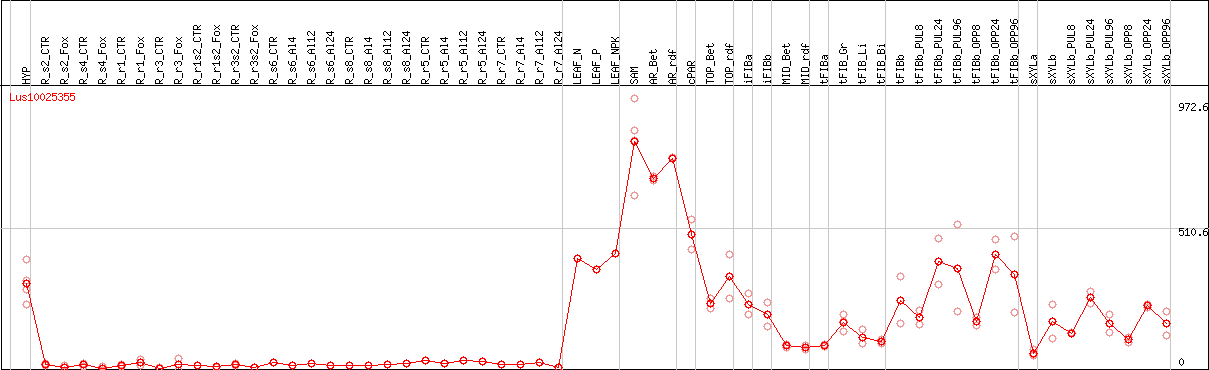
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025355 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.