Lus10026516 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10026516 pacid=23164226 polypeptide=Lus10026516 locus=Lus10026516.g ID=Lus10026516.BGIv1.0 annot-version=v1.0
ATGGATACAGACACTGGAAATGTCCATATACCAGTTGTTCTCCGGAGGCTACTTCCTGCTGTTGGACCTGTGCTTTTGATTTCAGTAGCGTATGTCGACC
CTGGGAAGTGGGCAGCCACTGTGGAAGGCGGTGCCCGTTTTGGAATTGATCTGGTCCTGCCAATGCTCATTTTTAATTTTGCAGCAATACTGTGCCAGTA
CCTGTCCGCTAAAGTTGGTATTGTCACCGGTAGAGATCTTGCTCAGATTTGCAGTGAGGAGTATGACAAGGGAACATGCATGCTCTTGGGAATTCAAACA
GCAGTTTCAGTGATGGCATTGGACCTCAGCATGGTACTTGGCACTGCACATGGGCTTAATCTTCTTCTGGGGCTGGACTTGCCCACCTGCATCTTTTTGA
CTGCTGTGGATGCTGTATTATTTCCATTTTTCGGTGCCTTTCTGGAGAAACACAAGGGATATTACCCATCCATATGTGTGGCAGGCTCTTTGTTACTCTT
CTATTTTCTTGGGGTGCTCATTAGTCAACCAGAGGTTCCTCTTTCATTGAATGGGATGCTGATGAAGTTAAGCGAGGAAAGTGCATTTGCTCTGATGAGT
CTTCTTGGTGCAAGCATTATGCCTCACAATTTCTATATGCATTCTTCCATTGTTCTGCATCATCCGCGATCTCCCAATGTTTCTAAGAGTTCCTTGTGCT
ATGACAGCTTTTTTGCCATTTTATGCATCTTTAGTGGCATATATTTGGTGAACTATGTGCTCATGAACTCGGCTGCGAATGTTTTCCACAGTACTGGCCT
TGTTTTGGTTACTTTTCCAGATGCTTTGTCACTAATGGAGCAGGTGTTTCGCAGTCCAGTGGCACCCTTGTTCTTTGTAGTGATTTTGTACCTTTCGAAT
CAAATCTCAGCATTAACTTGGAACCATGGCGGGCAAGTAGTTGTGCATGATTTCCTCCTACTTGACATACCTAACTGGCTCCAGCGAGCTACAACCAGGA
TTGTTGCGATTGTTCCAGCCCTTTATTGTGTATGGACTTCTGGAGTTGAAGGAGCATACCAGTTGCTCATATTTTCTCAAGTTATGGTAGCCCTTCTACT
ACCATCTTCTGTTTCTTCTGTCATTCCACTATTCCGGGTGGCATCATCAAGGGCCATAATGGGTGCCCATAAAATATCTCAAATCCTGGAGTTCTTGGCA
GTTATCACATTCATGGGGATGCTAGGGTTGAAGATCATATTTGTGGTAGAAATGATTTTTGGGGATAGTGATTGGATCGGAAATTTGCAATGGAATATGG
GCAGCAGTTCGTCTGTTCCTTCTGTTGTTCTTATTCTAACGGCTTTTACGTCATTCTGTTTGATGCTTTGGCTAGCAGCTACACCTTTGAAATCTGCGAC
CGCCACCCATGCAGACCCTCAGACTTGGAACTGGGATGCAAAGCAAACTTTGCCAGAGCCATCCATGCGCAGAGATGAAACTTATTTAAGTGAGACTAGG
TATGAAGGAGAAAGGCGTGCTGATCCAGTGAAGCCATTTGAGGGTTACACAGATAGAAGTATTGTACATGCTGATCCACATTTACCTGAAACTATTTTGG
ATTCTCACAATTTGAGACCTGTAGCAGAACATCATTCTGATGTGAAATTTTTGAGTCCTCTGAGACCAAATCAGTCTGATATGAAGCTTCCTAGTCCTGT
GGAACCAAATCAGTCTGATATGAAGCTTCCTAGTCCTCTGAGATTAAATCAATCTGATATGAAGCTTCCAAGTCCTCTGATACCAAATCAGTCTGATATG
AAGCTTCCAAGTCCTCTGAGATCAAAGCAGTCTGATCTGAAGCTTCCGAGTCCCCCAAGACCAAATCAGTCTGATCTGAATCTTCCTAGACCTCTGGAAC
CAAATCTGGAATCAGCACCTTCAGTAGAGCCAGTGTCAGTGCCAAAGGCAGAAAATTTAGCTGTTACTGAATTTGATTTACCGGACTCGAAGATGACGGT
TGAATCACTTGAACCAATCGAAAAGACCGTGAGCATGGAAGGTGATTTGCTGCCAGAAAAGGAGCATGACAAGGGAGAATCATGGGAGCCTGAAGAGCCC
TCCAAAGGAGTTCCTGCGAGTGTTTCGTCTTTCTCATCTGATGGTCCATCGTCATTCAGGAGTCTCAGTGGAAAAAGTGACGATGGTGGAAATGGTGCTG
GAAGTCTTTCAAGACTGGCAGGGCTTGGTAGGGCTGCCAGACGTCAATTAGCTACAGTTCTTGATGAATTCTGGGGACAGTTGTATGATTTCCATGGTCA
ACCAACGCAGGATGCCAAGAATAGAAGACTCGATCTGTTACTTATCGATCCAAAGCTTTCGTTTTCCTTCCTGAAAGTTGACCCTCCTTCAACTGAATAC
AATGGGTATCCATCAAGAACAGACAGTAAAGGATCTGAATCTTTCATGGGTTCAAGTTTCTGTGATTCTCCCCGGCAGGTGACGCTGCAGAGTACTCTTG
ACTCTCCATATAATGGGGGTCGAAGTGGACCTTCCTCGTTATTATCCAACCGCTCTCAGTTACTTGATAAATATTCTTTACCCAGCTCGAGCGTGGTGGA
CCCTTTTGAGAGGCGGTATTCTAGTGTGCGAAATCTACCATCTACTGATGGTTGGGGAAATAGTCAGCCAATTACAAATATGATGGACCCTTCTGAAAGG
CGTTACTCAAGTGTGCGAAATTTACCATCTTCTGATGGTTGGGGAAATGGCCAGCTAGCTACAAATGTGATGGACCCTTCTGAAAGGCGTTACTCAAGTG
TGCGAAATCTACCATCTTCTGATGGTTGGGGGAATACTCAGCCAGCTACAATCCATGGTTATCAGATTGCTTCCATTGCTAACCGGATTGCCAAGGAGAG
GAGTTCCAGTGGCCTGAATGGTGGAGTAGAGCCCATGTCGCCAACTTCTTCAGCCATGGGCATGCAGAGTTACAGGGATCCACAAGCAAGTGCTTTGGGC
CAGAGGATACAGAGTGGATTGAGCTCCCCGGGATATCAGAATTTCGCAGTATCCAGAAATAGCTCATTACAGTCTGAAGCACAATACTACGATCTTGATT
CGGCTTCTGCTGCTAACGATGGAGGACAGACAACCAATACAAAGAAGTACCACAGTTTGCCTGACATTTCAGGAATTGCTGGGCCTTACAGAGAGCTATA
TCTATCCAGAAAACCTACTCAGCGCGACAATTCTGGTGGTTTTGCTTCCTCCTTAGGGAGAACGAGTTACGAGTCTTCTTTGTATCCAAATACTGGATCA
GGAAATGCAAGTTCTTTGGGTTTTAATGACATTGGCACAAGTTATGGAGATGCTTTCTCATTTCCTACAACAGTTGCTGCTACTACTAGTACAGCTGGCG
CAGGATCTCTTTGGTCCAGACAGCCCTTTGAGCAGTTTGGTGTTGGGGATAAAAACTGTGCAGTTGGAAGTTCTGTAGGAAGTAGTAGGTCTAATCCGAT
TTCTCGTGAAGCTAGTTCCATAGTGGATGGAGAGGCGAAACTTCTTCGGTCCTTCAGATGCTGCATTGTCAAGCTACTGAAACTGGAAGGCTCGGATTGG
TTGTTCAGGCAGAATGATGGAGCTGATGAGGACTTGATTGATCGTGTGGCCGCACGTGAGAAGTGCTTATATGAAGCTGAGGCGAGAGAGATGGGCGGCC
AGGGGGTTCACTTGGGTGAGTCTCAATCCAGCAGCGATAAGAAGCAGAAGAATGACGAGGCCAGCATTGCTAGTATCTTGGTTTCGTCAATTCCTCACTG
CGGAGAAGGCTGTGTTTGGAAGGCGGATTTGATTGTCAGCTTTGGAGTTTGGTGCATTCACAGGATTCTCGATTTGTCGCTAATGGAAAGCCGGCCTGAG
CTATGGGGGAAGTATACCTATGTACTCAATCGCCTTCAGGGGATTGTGGAGCCGGCATTCTTGAGACCAAGGACTCCAATGCAGCCATGCTTCTGCCTGC
AACTCTCTGCAGCGCACCAGCAGAGGTTGTTGTCCCAGCCACCATCGAATGGTATGCTACCGCCCACTGTGAAACCAGGGAGAGGGAAATGCACAACTGG
TGCAATGGTCCTGGAGATGATAAAGGATGTAGAGATAGCGATTTCATGCAGGAAAGGCCGGTCAGGGACTGCTGCCGGGGATGTAGCTTTCCCAAAGGGA
AAAGAGAACCTGGCGTCCGTCCTGAAACGGTACAAGCGGCGGCTTTCAAACAAGAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10026516 pacid=23164226 polypeptide=Lus10026516 locus=Lus10026516.g ID=Lus10026516.BGIv1.0 annot-version=v1.0
MDTDTGNVHIPVVLRRLLPAVGPVLLISVAYVDPGKWAATVEGGARFGIDLVLPMLIFNFAAILCQYLSAKVGIVTGRDLAQICSEEYDKGTCMLLGIQT
AVSVMALDLSMVLGTAHGLNLLLGLDLPTCIFLTAVDAVLFPFFGAFLEKHKGYYPSICVAGSLLLFYFLGVLISQPEVPLSLNGMLMKLSEESAFALMS
LLGASIMPHNFYMHSSIVLHHPRSPNVSKSSLCYDSFFAILCIFSGIYLVNYVLMNSAANVFHSTGLVLVTFPDALSLMEQVFRSPVAPLFFVVILYLSN
QISALTWNHGGQVVVHDFLLLDIPNWLQRATTRIVAIVPALYCVWTSGVEGAYQLLIFSQVMVALLLPSSVSSVIPLFRVASSRAIMGAHKISQILEFLA
VITFMGMLGLKIIFVVEMIFGDSDWIGNLQWNMGSSSSVPSVVLILTAFTSFCLMLWLAATPLKSATATHADPQTWNWDAKQTLPEPSMRRDETYLSETR
YEGERRADPVKPFEGYTDRSIVHADPHLPETILDSHNLRPVAEHHSDVKFLSPLRPNQSDMKLPSPVEPNQSDMKLPSPLRLNQSDMKLPSPLIPNQSDM
KLPSPLRSKQSDLKLPSPPRPNQSDLNLPRPLEPNLESAPSVEPVSVPKAENLAVTEFDLPDSKMTVESLEPIEKTVSMEGDLLPEKEHDKGESWEPEEP
SKGVPASVSSFSSDGPSSFRSLSGKSDDGGNGAGSLSRLAGLGRAARRQLATVLDEFWGQLYDFHGQPTQDAKNRRLDLLLIDPKLSFSFLKVDPPSTEY
NGYPSRTDSKGSESFMGSSFCDSPRQVTLQSTLDSPYNGGRSGPSSLLSNRSQLLDKYSLPSSSVVDPFERRYSSVRNLPSTDGWGNSQPITNMMDPSER
RYSSVRNLPSSDGWGNGQLATNVMDPSERRYSSVRNLPSSDGWGNTQPATIHGYQIASIANRIAKERSSSGLNGGVEPMSPTSSAMGMQSYRDPQASALG
QRIQSGLSSPGYQNFAVSRNSSLQSEAQYYDLDSASAANDGGQTTNTKKYHSLPDISGIAGPYRELYLSRKPTQRDNSGGFASSLGRTSYESSLYPNTGS
GNASSLGFNDIGTSYGDAFSFPTTVAATTSTAGAGSLWSRQPFEQFGVGDKNCAVGSSVGSSRSNPISREASSIVDGEAKLLRSFRCCIVKLLKLEGSDW
LFRQNDGADEDLIDRVAAREKCLYEAEAREMGGQGVHLGESQSSSDKKQKNDEASIASILVSSIPHCGEGCVWKADLIVSFGVWCIHRILDLSLMESRPE
LWGKYTYVLNRLQGIVEPAFLRPRTPMQPCFCLQLSAAHQQRLLSQPPSNGMLPPTVKPGRGKCTTGAMVLEMIKDVEIAISCRKGRSGTAAGDVAFPKG
KENLASVLKRYKRRLSNKK
|
DESeq2's median of ratios [FLAX]
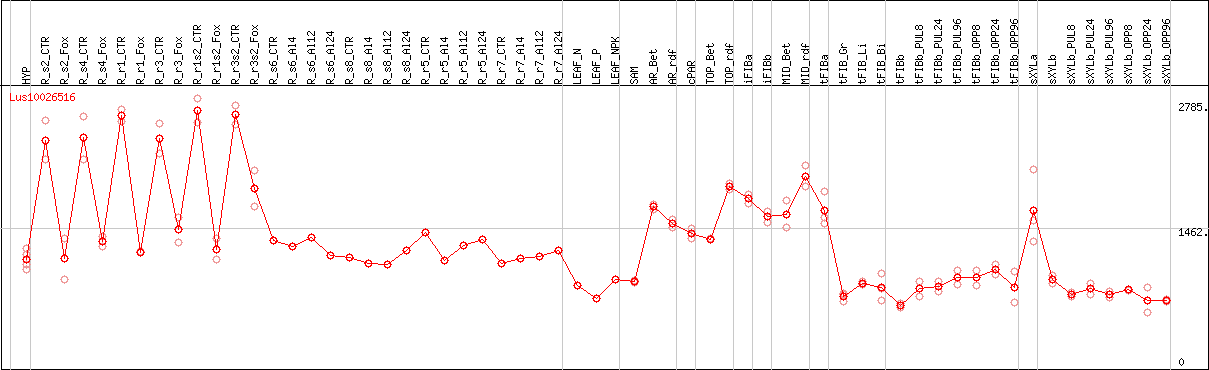
Coexpressed genes
Lus10026516 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
