External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G07040 71 / 1e-15
RPS3, RPM1
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
AT1G59780 57 / 1e-10
NB-ARC domain-containing disease resistance protein (.1)
AT1G58807 55 / 4e-10
Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
AT1G59124 55 / 4e-10
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G59218 55 / 5e-10
Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
AT1G58848 55 / 5e-10
Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
AT3G46530 55 / 6e-10
RPP13
RECOGNITION OF PERONOSPORA PARASITICA 13, NB-ARC domain-containing disease resistance protein (.1)
AT1G50180 53 / 3e-09
NB-ARC domain-containing disease resistance protein (.1)
AT1G58390 53 / 3e-09
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G58400 52 / 5e-09
Disease resistance protein (CC-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002945
78 / 4e-18
AT3G07040 580 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10020016
75 / 4e-17
AT3G07040 563 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10041243
75 / 6e-17
AT3G07040 526 / 1e-172
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10014922
74 / 1e-16
AT3G07040 523 / 1e-171
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10028639
74 / 2e-16
AT3G07040 579 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10018937
73 / 2e-16
AT3G07040 587 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10019530
69 / 1e-14
AT3G07040 311 / 4e-95
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10014712
62 / 2e-12
AT3G07040 492 / 3e-159
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10024173
50 / 2e-08
AT3G14460 681 / 0.0
LRR and NB-ARC domains-containing disease resistance protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G134601
123 / 4e-34
AT3G07040 443 / 2e-142
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134300
122 / 1e-33
AT3G07040 487 / 2e-157
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134450
122 / 1e-33
AT3G07040 488 / 1e-157
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134701
120 / 6e-33
AT3G07040 464 / 6e-149
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.003G099100
115 / 2e-31
AT3G07040 482 / 8e-156
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134400
114 / 9e-31
AT3G07040 427 / 3e-134
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.003G099000
113 / 1e-30
AT3G07040 479 / 2e-154
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134475
113 / 1e-30
AT3G07040 413 / 1e-129
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134425
102 / 1e-26
AT3G07040 347 / 6e-107
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134500
102 / 1e-26
AT3G07040 422 / 1e-132
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Lus10026765 pacid=23176624 polypeptide=Lus10026765 locus=Lus10026765.g ID=Lus10026765.BGIv1.0 annot-version=v1.0
ATGGATCAAGGATCAAGCTCCTTTAGTGAAATGCACTTGGCTAGAAGAGGAGATGACCTTTTCCTTGAAGATGTTGATATGGTTGGTATTGAAAATCCAA
ACAGGCAACTGATCAGATGGCTAGTTGGGGGCAAATCTGAACGTGAAATGGTGTCTATCACCGGGATGGGTGGTCGGGGAAAAACCACCTTGGCCAAGAA
GTTATATGATGATGCAGATATGAAGAAACACTTCAAATTCTGTGTTTGGATTACTCTCTCTCCACACGTAATGATAGAGGATGTGTTGAAGGACATGATT
TAG
AA sequence
>Lus10026765 pacid=23176624 polypeptide=Lus10026765 locus=Lus10026765.g ID=Lus10026765.BGIv1.0 annot-version=v1.0
MDQGSSSFSEMHLARRGDDLFLEDVDMVGIENPNRQLIRWLVGGKSEREMVSITGMGGRGKTTLAKKLYDDADMKKHFKFCVWITLSPHVMIEDVLKDMI
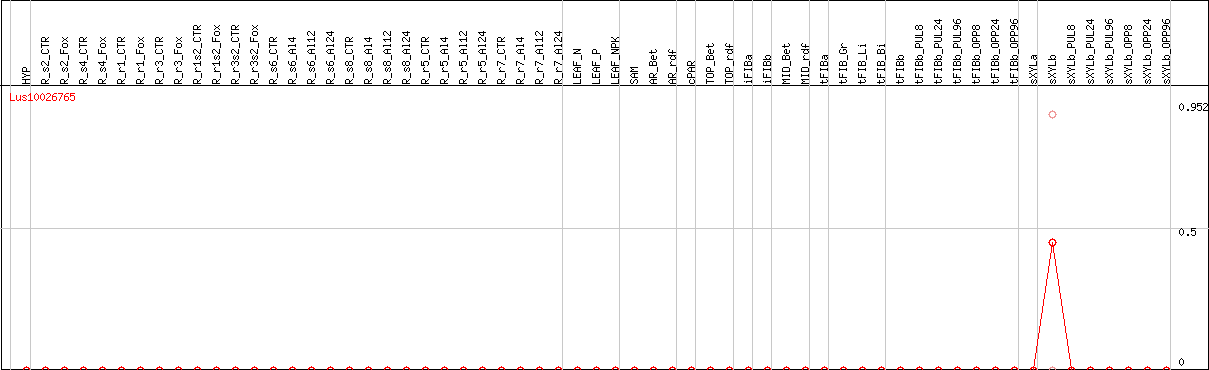
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026765 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.