External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G70300 249 / 2e-78
KUP6
K+ uptake permease 6, K+ uptake permease 6 (.1)
AT5G14880 232 / 3e-72
Potassium transporter family protein (.1)
AT2G40540 212 / 2e-64
ATKUP2, ATKT2, TRK2, SHY3, KT2
potassium transporter 2 (.1.2)
AT2G30070 172 / 2e-50
ATKUP1, ATKT1P, ATKT1
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
AT3G02050 154 / 2e-43
ATKT4, ATKUP3, KUP3
K+ uptake transporter 3, ARABIDOPSIS THALIANA POTASSIUM TRANSPORTER 4, K+ uptake transporter 3 (.1)
AT4G23640 146 / 7e-41
ATKT3, KUP4, TRH1
TINY ROOT HAIR 1, Potassium transporter family protein (.1)
AT4G13420 146 / 9e-41
HAK5, ATHAK5
high affinity K+ transporter 5, ARABIDOPSIS THALIANA HIGH AFFINITY K+ TRANSPORTER 5, high affinity K+ transporter 5 (.1)
AT1G31120 137 / 9e-38
KUP10
K+ uptake permease 10, K+ uptake permease 10 (.1)
AT2G35060 137 / 9e-38
KUP11
K+ uptake permease 11, K+ uptake permease 11 (.1), K+ uptake permease 11 (.2)
AT1G60160 137 / 1e-37
Potassium transporter family protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G147400
249 / 2e-78
AT1G70300 1221 / 0.0
K+ uptake permease 6, K+ uptake permease 6 (.1)
Potri.010G094300
246 / 2e-77
AT1G70300 1228 / 0.0
K+ uptake permease 6, K+ uptake permease 6 (.1)
Potri.013G083400
215 / 9e-66
AT2G40540 1225 / 0.0
potassium transporter 2 (.1.2)
Potri.019G056500
214 / 3e-65
AT2G40540 1272 / 0.0
potassium transporter 2 (.1.2)
Potri.009G073500
183 / 3e-54
AT2G30070 1092 / 0.0
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
Potri.014G144900
175 / 4e-51
AT3G02050 1175 / 0.0
K+ uptake transporter 3, ARABIDOPSIS THALIANA POTASSIUM TRANSPORTER 4, K+ uptake transporter 3 (.1)
Potri.002G237500
170 / 3e-49
AT3G02050 1179 / 0.0
K+ uptake transporter 3, ARABIDOPSIS THALIANA POTASSIUM TRANSPORTER 4, K+ uptake transporter 3 (.1)
Potri.003G109800
153 / 4e-43
AT2G35060 985 / 0.0
K+ uptake permease 11, K+ uptake permease 11 (.1), K+ uptake permease 11 (.2)
Potri.001G123800
151 / 1e-42
AT2G35060 1149 / 0.0
K+ uptake permease 11, K+ uptake permease 11 (.1), K+ uptake permease 11 (.2)
Potri.003G109700
150 / 3e-42
AT2G35060 1155 / 0.0
K+ uptake permease 11, K+ uptake permease 11 (.1), K+ uptake permease 11 (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0062
APC
PF02705
K_trans
K+ potassium transporter
Representative CDS sequence
>Lus10029173 pacid=23151689 polypeptide=Lus10029173 locus=Lus10029173.g ID=Lus10029173.BGIv1.0 annot-version=v1.0
ATGGATCTGGAGACTGGTGCTCATCATAAAAATGTCCAGGTTAGAATCGTGGCAATCTTTTTGTTGCAGAAGGAGTCATGGAGGACGGTATTGACACTAG
TGTACCAGAGCTTAGGAGTCGTCTATGGCGATTTAAGCATTTCCGCTTTGTACGTTTTCAAGAGCACGTTTGCTGATGAGGACATAATCCATTCTGAGAC
CAACGAGATCTTTGGGGTTCTGTCGTTTGTATTCTGGACTCTCACTTTAGTTCCCTTATTGAAGTATGTGTTCATAGTTCTTAGAGCTGATGATAATGGG
GAAGGAGGCACATTTGCTCTATTCTCTTTGCTGTATCGACACGCCCGAGTCAACTCGCTCCCCAATCGCCAGGTGGCAGATGGGGAGCTGTCTGAGTATA
AGAAGAAGGAGAGCAGTGATGGATCATCAGTTCATCCGGGTTTTCCTTCAGGGTTGAAATCAACGTTGGAGAAGCACAGGGTTCTGCAGAGGATCTTTTT
GGCCTTGGTTGGGACTTGTATGGTGATTGGTGATGGCATTCTGACTCCTGCCCTTTCTGGTAAAACATAA
AA sequence
>Lus10029173 pacid=23151689 polypeptide=Lus10029173 locus=Lus10029173.g ID=Lus10029173.BGIv1.0 annot-version=v1.0
MDLETGAHHKNVQVRIVAIFLLQKESWRTVLTLVYQSLGVVYGDLSISALYVFKSTFADEDIIHSETNEIFGVLSFVFWTLTLVPLLKYVFIVLRADDNG
EGGTFALFSLLYRHARVNSLPNRQVADGELSEYKKKESSDGSSVHPGFPSGLKSTLEKHRVLQRIFLALVGTCMVIGDGILTPALSGKT
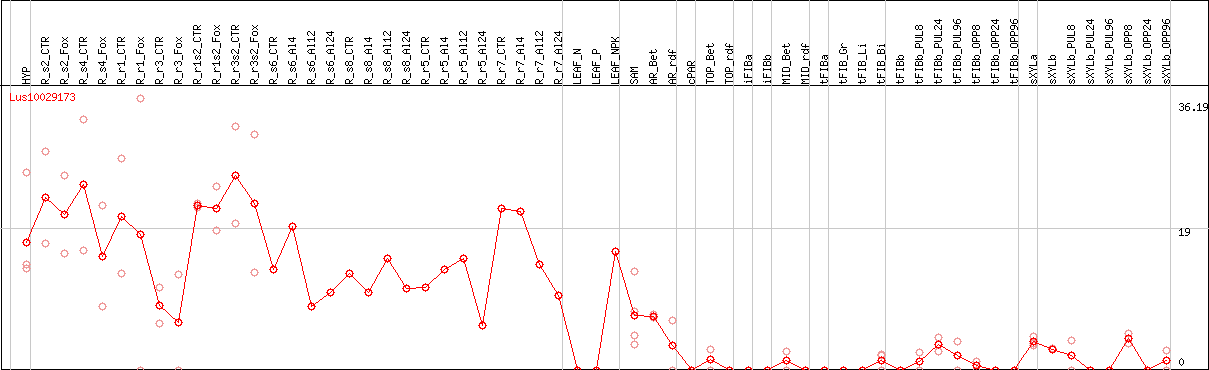
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029173 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.