External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G51680 390 / 6e-137
AtSDR2
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G26770 237 / 9e-77
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G29250 230 / 2e-74
AtSDR4
short-chain dehydrogenase reductase 4, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT4G03140 231 / 4e-74
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47140 228 / 6e-74
AtSDR5
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G26760 229 / 2e-73
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47130 226 / 4e-73
AtSDR3
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47120 213 / 6e-68
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G29260 212 / 2e-67
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G52340 204 / 2e-64
SIS4, SDR1, ISI4, GIN1, ATABA2, SRE1, ABA2, ATSDR1
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022756
572 / 0
AT3G51680 397 / 2e-139
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026419
397 / 3e-139
AT3G51680 367 / 9e-128
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10021314
260 / 4e-86
AT3G51680 238 / 6e-78
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10021312
258 / 3e-85
AT3G51680 238 / 8e-78
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016995
257 / 5e-85
AT3G51680 243 / 7e-80
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10021313
257 / 8e-85
AT3G26770 232 / 3e-75
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10021318
256 / 2e-84
AT3G51680 238 / 9e-78
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016991
255 / 4e-84
AT2G47140 238 / 2e-78
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016997
252 / 4e-83
AT1G52340 249 / 2e-82
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G105900
422 / 9e-150
AT3G51680 412 / 4e-146
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G129100
422 / 2e-149
AT3G51680 383 / 3e-134
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.019G125600
411 / 3e-145
AT3G51680 394 / 8e-139
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G199900
273 / 2e-91
AT2G47140 256 / 1e-85
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G106100
270 / 6e-90
AT2G47140 234 / 9e-77
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G200100
268 / 2e-89
AT3G26770 257 / 2e-85
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G073900
260 / 5e-86
AT3G51680 252 / 2e-83
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G207100
258 / 2e-85
AT4G03140 241 / 2e-78
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G074000
253 / 2e-83
AT1G52340 246 / 5e-81
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G206500
251 / 1e-82
AT1G52340 246 / 4e-81
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10029438 pacid=23139823 polypeptide=Lus10029438 locus=Lus10029438.g ID=Lus10029438.BGIv1.0 annot-version=v1.0
ATGACTGCACAAGTTATTCCGGCGGAGACAACCATCCAACAGCCGGGCATTCATGTTTTGTCCGGCCGAGACCAACAATCCCATCTCCCTCCCCGAAGAT
TGGAAGGGAAGATTGCGGTAGTGACAGGTGGCGCACGGGGAATCGGGGAAGCGACTGCGCGTCTTTTCGCAAAGCTTGGAGCCAAAGTTGTGATTGCCGA
CGTGGACGACAAACTCGGCAACGCGTTGGCCGACTCTTTATCATTGTCGTTGACAACAATGAATCATTCGCCTCCTTCTGTATCGTACGTGCACTGCGAT
GTCAGCTCTGAATCGGATGTCGAAAACTTGATAAGCTCCACAGTGTCACGATACGGGAGGTTAGACATACTCTTCAACAACGCCGGTGTCTTGGGGTCTC
AGACGAAGAAAAAAAAGAGCATCATCAACTTTGATGCGAGTGAGTTTGACCGGGTCATGCGTGTCAACGTCCTGGGAATGGCGCTAGGGATCAAGCACGC
AGCTCGTGCGATGATCCCACGTGGCGGTGGGTGCATAGTGTCCACCGCCAGCGTCGCGGGGGTAATGGGAGGGATGGGGCCCCATGCTTACACGGCGTCA
AAGCACGCTATAGTGGGGCTGACAAAGAACGCAGCGTGCGAGTTGGGAAAGTATGGGATCCGTGTCAATTGCATTTCTCCGTTCGGGGTCGCCACTTCGA
TGTTGGTTAACGCATGGCGCGACGACGAGAATGCAAATGACGACGAGGATAAGATCATGTTACCCTGCGAGGAGGAGGTGGAGGAGATCGAGGAGTTTGT
GAGAGGGTTGGGGAATTTGAAAGGAGGAGCAACGCTAAAGGCCAAAGATATAGCGGAGGCAGCGGTGTATTTGGTTAGCGACGAGTCTAAGTACGTTAGT
GGGCATAACCTAGTGGTTGATGGCGGTGTAACAACTTCGAGGAATTGTGTTGGGTTATAA
AA sequence
>Lus10029438 pacid=23139823 polypeptide=Lus10029438 locus=Lus10029438.g ID=Lus10029438.BGIv1.0 annot-version=v1.0
MTAQVIPAETTIQQPGIHVLSGRDQQSHLPPRRLEGKIAVVTGGARGIGEATARLFAKLGAKVVIADVDDKLGNALADSLSLSLTTMNHSPPSVSYVHCD
VSSESDVENLISSTVSRYGRLDILFNNAGVLGSQTKKKKSIINFDASEFDRVMRVNVLGMALGIKHAARAMIPRGGGCIVSTASVAGVMGGMGPHAYTAS
KHAIVGLTKNAACELGKYGIRVNCISPFGVATSMLVNAWRDDENANDDEDKIMLPCEEEVEEIEEFVRGLGNLKGGATLKAKDIAEAAVYLVSDESKYVS
GHNLVVDGGVTTSRNCVGL
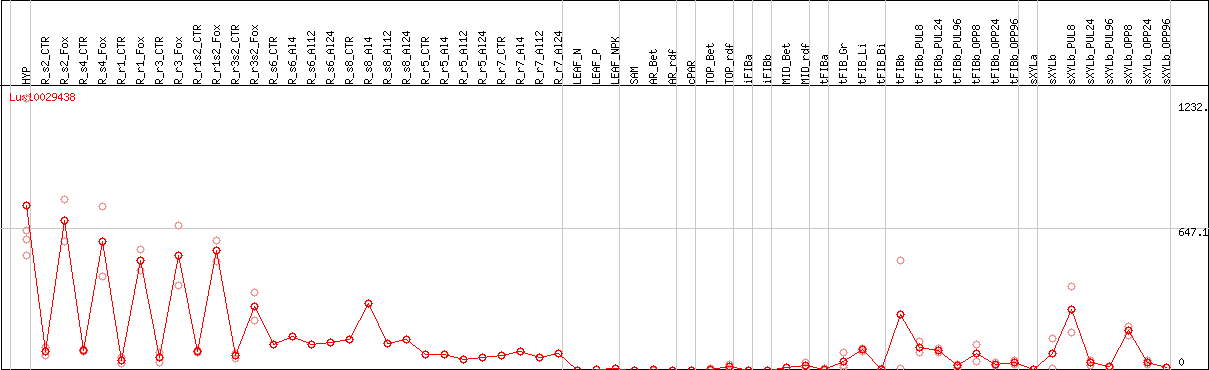
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029438 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.