Lus10029628 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10029628 pacid=23152036 polypeptide=Lus10029628 locus=Lus10029628.g ID=Lus10029628.BGIv1.0 annot-version=v1.0
ATGGACGAAGTGCTGCTCAAGTGCTCCACTGAAGGAACAACAGTCTCCAATGCTTTTCCAGAATCACAAACATCAGCTGCAGAACGGCAATCTGTCGCCA
GTACTTTGGAGTACGAGGTCTTCTTGAGCTTTAGAGGGTCTGATGTCCGCAAAAGCTTTGCGGACTTTCTCTACAGCTATTTGGTCCGTTCCAAAATCCG
CACCTTTAGGGACGAGGAAGAGCTTCGGAAGGGAGAGAATATTGCACCTTCTCTCGTCCAGGCTATCACGGAATCGAAGATTTACATCCCAATTTTATCA
AAGAACTATGCTTCTAGCAAATGGTGCCTCCAGGAACTAGCTAAGATGGTTGAGTGTTGCAGGAAAGGGAAGGGACAAATCATATTGCCCGTTTTCTATT
TTGTAGACCCCAGAGATGTGCGGCATCAAGCGGGTCCATACGAGAAGGCATTTGAACTGCATAGTAAGAAACATGGCCCCATAATTGTACAAGAATGGAG
GACAGCTCTCCAAGAAGTTGGACAAATGAAAGGATGGCATGTTACCGAAATAGACGGTCAGGGAGCTGTTATAGATCATGTATTTTCTACGGTCGAGTTG
CATCTGAGGAGTAACTACACTTTAGTGACTGATGAGCTGGTTGGGATTGATTCTCATGTGCGAGAAGTGACGAGATTGTTGAACTTAGACTCTAGAGATG
CGAGAATTGTTGGGATTCAAGGCATTGGCGGCATGGGTAAAACTACTATAGCCAAGGCAGTCTATGACGATATCTGCACAAGCTTCGACCGTTGCTGTTT
CTTAGAAAATATACGAGAAGTATTGTCGAGGAGTGATGGCATTATGTCTTTACAGAATAAGTTCATCTCTAGCATTTTGAGGGATGAAGTTCAGGTTAAA
GATCCTAGCGAAGGGGTCAACATGATTAGAGAGAGAGTCTGCAAATATAAACTCTTGGTTGTTCTTGATGATGTAGACGATAAGTTTGAGTTTGATCAGA
TATTGGGAAAATTAGGTCACTTCTACTCGGAAAGTAGATTCATCATCACAACAAGAGACAAAAGAGTTTTAGAGCTCCTTCAAGAATACAAGTTGTATGA
ACCGGAAGAAATGAGTTATGATCATTCTCTCCAACTTTTTAGCAAACATGCTTTTCGTATGGATTACCCTCCTGGAGATTATGCTGCTCTATGTGATGAT
TTCGTAAAAGTTGCATCAAGGCTTCCTTTGGCTCTTAAGGTCATAGGATCACTCCTGTTTTGTAGAGAGAAACAGTTCTGGGAAGTAAAGTTGATGGAGC
TGAAAAATTTACCAGCTGCCACTAGCAATAAGGTGCAGCAGAGATTGAAGATTAGCTTCAATGAGTTGACCCCTAATGAAAAACAGATCTTTCTTGATAT
AGCCTGCTTTTTTATTGGAGAGGATAAAGAATCGCCCGTTCATATGTGGAGTGATTGTAAAGTTTTTCCGGAAACTGGGATAAGTACTCTTGTTCTTAGA
TCCTTAATAAAAATTGATGAGGAAAATCGATTCTGGATGCATGATCTTCTTAGAGACCTTGGGAGAGCTATTGTTATTGAGGAAGATGTTAAAAATCCCT
GCAAGCGCAGCAGGATTTGGTCCAATGGAGATGCTTGGAAAATATTGAAAAACGTAAAGGGAGCTGATCAAGTGGAAATGCTCAGGATTGACATGAGACA
TGGTGACAGAAATCTTAAGCTCACAGCCAAAGCATTTGAAAAGCTGGCAGGGTTAAGGTATCTAGACCTGGCATTTTTCACGCTTACTAGGGACTTCGGT
AGGATTCTTCCAGATATATCGTTGCGAAAAATGAATCTGTGTGGTTGTTGGTCAGTTGAGGAGTTGGATGCAAGTAAATGCACAGAGTTGATTAAGGTTG
TAAATTTTGGGAAATTGGAAAAGTTACGGGAGCTGAACTTGTTTGGTTGCAAGTCAACTACGGAGTTGTTTGATTTGTCCGGTCTCGTGAGCTTAGATGG
TTTAAATTTGGAGGGATGCACGGAGTTGATTAAGGTCACAAGTTTCGAGAAACTGGAGATGTTACGAAAATTGAACATGCTTGGTTGCAAGTCTATAAGG
GAGTTGCCAGATATGTCGTGTCTCAAGAACCTGCATGATTTAAACTTGGAGGGCTGCGCAAAGTTGATTAAGGTTACATGTTTTGGGAAATTGGAGAAGT
TACGGAAGCTGAACTTGTTTGGTTGCAAGTCTATTAGGGAGTTGCCAGATCTGTCCGGCCTCGTAAGCTTGAATGATTTGAATTTGGAGGGATGCACGAA
GTTGGTTAAGATCACAAGTTACCAGAAATTGGAGATGCTACGAAAGTTGAACATGGCTGGTTGCGAGTCTATTAAGGAGTTGTCAAATATGTCTGATCTC
AAGAACCTGCAGGAGTTAAACATGGAGGGATGCATAAAGTTGAGTAAAGTTAGAAGTTTTGAAAAATTGGAGACGTTACGAAAGTTGAACATGTCTGGTT
GCGAGTCTATTAAGGAGTTTCCTGATATGTCTGGTCTCAAGAACCTGCATGATTTAAACTTGGAGGGATGCACAAAGTTGATTGAGGTTACAAGTTTTGA
GAAATTGGAAAAGTTACGGAAGTTGAACATGTCTGGTTGCAAGTCTATTAAGGAGTTGCCAGATATGTCCGGTCTCATGAGCATAGATGATTTAAATTTT
GAGGGATGCACAGAGTTGGTTAAGGTCACAAATTTTGAAAAAATGGAGATGTTACAGAAGTTGAACATGTCTGGTTGCAAGTCTATTAAGGAGTTGCCAG
ATCTATCGCGTCTCAAGAACCTACATGTTTTAAACTTGGAAGGATGCACAGAGTTGATTAAGGTTACAAGTTTCGGGAGATTGGAGATGTTACAGAAGTT
GAAGATGTCTGGTTGTCAGTCTATTAAGGAGTTGCCAGATATGTCCGGTCTCATGAGCATAGATGATTTAAATTTTGAGGGATGCACAGAGTTGGTTAAG
GTCACAAATTTTGAAAAAATGGAGATGTTACAGAAGTTGAACATGTCTGGTTGCAAGTCTATTAAGGAGTTGCCAGATCTATCGCGTCTCAAGAACCTAC
ATGTTTTAAACTTGGAAGGATGCACAGAGTTGATTAAGGTTACAAGTTTCGGGAGATTGGAGATGTTACAGAAGTTGAAGATGTCTGGTTGTCAGTCTAT
TAAGGAGTTGCCAGATATGTCCGGTCTCATGAGCATAGATGATTTAAATTTTGAGGGATGCACAGAGTTGGTTAAGGTCACAAATTTTGAAAAAATGGAG
ATGTTACAGAAGTTGAACATGTCTGGTTGCAAGTCTATTAAGGAGTTGCCAGATCTATCGCGTCTCAAGAACCTACATGTTTTAAACTTGGAAGGATGCA
CAGAGTTGATTAAGGTTACAAGTTTCGGGAGATTGGAGATGTTACAGAAGTTGAAGATGTCTGGTTGTCAGTCTATTAAGGAGTTGCCAGATATGTCCGG
TCTCATGAGCATAGATGATTTAAATTTTGAGGGATGCACAGAGTTGGTTAAGGTCACAAATTTTGAAAAAATGGAGATGTTACAGAAGTTGAACATGTCT
GGTTGCAAGTCTATTAAGGAGTTGCCAGATCTATCGCGTCTCAAGAACCTACATGTTTTAAACTTGGAAGGATGCACAGAGTTGATTAAGGTTACAAGTT
TCGGGAGATTGGAGATGTTACAGAAGTTGAAGATGTCTGGTTGTCAGTCTATTAAGGAGTTGCCAGATATGTCCGGTCTCATGAGCATAGATGATTTAAA
TTTTGAGGGATGCACAGAGTTGGTTAAGGTCACAAATTTTGAAAAAATGGAGATGTTACAGAAGTTGAACATGTCTGGTTGCAAGTCTATTAAGGAGTTG
CCAGATCTATCGCGTCTCAAGAACCTACATGTTTTAAACTTGGAAGGATGCACAGAGTTGATTAAGGTTACAAGTTTCGGGAGATTGGAGATGTTACAGA
AGTTGAAGATGTCTGGTTGTCAGTCTATTAAGGAGTTGCCAGATATGTCCGGTCTCATGAGCATAGATGATTTAAATTTTGAGGGATGCACAGAGTTGGT
TAAGGTCACAAATTTTGAAAAAATGGAGATGTTACAGAAGTTGAACATGTCTGGTTGCAAGTCTATTAAGGAGTTGCCAGATCTATCGCGTCTCAAGAAC
CTACATGTTTTAAACTTGGAAGGATGCACAGAGTTGATTAAGGTTACAAGTTTCGGGAGATTGGAGATGTTACAGAAGTTGAAGATGTCTGGTTGTCAGT
CTATTAAGGAGTTGCCAGATATGTCCGGTCTCATGAGCCGTGTGGAGCGCGGCGGTGAAACTCAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10029628 pacid=23152036 polypeptide=Lus10029628 locus=Lus10029628.g ID=Lus10029628.BGIv1.0 annot-version=v1.0
MDEVLLKCSTEGTTVSNAFPESQTSAAERQSVASTLEYEVFLSFRGSDVRKSFADFLYSYLVRSKIRTFRDEEELRKGENIAPSLVQAITESKIYIPILS
KNYASSKWCLQELAKMVECCRKGKGQIILPVFYFVDPRDVRHQAGPYEKAFELHSKKHGPIIVQEWRTALQEVGQMKGWHVTEIDGQGAVIDHVFSTVEL
HLRSNYTLVTDELVGIDSHVREVTRLLNLDSRDARIVGIQGIGGMGKTTIAKAVYDDICTSFDRCCFLENIREVLSRSDGIMSLQNKFISSILRDEVQVK
DPSEGVNMIRERVCKYKLLVVLDDVDDKFEFDQILGKLGHFYSESRFIITTRDKRVLELLQEYKLYEPEEMSYDHSLQLFSKHAFRMDYPPGDYAALCDD
FVKVASRLPLALKVIGSLLFCREKQFWEVKLMELKNLPAATSNKVQQRLKISFNELTPNEKQIFLDIACFFIGEDKESPVHMWSDCKVFPETGISTLVLR
SLIKIDEENRFWMHDLLRDLGRAIVIEEDVKNPCKRSRIWSNGDAWKILKNVKGADQVEMLRIDMRHGDRNLKLTAKAFEKLAGLRYLDLAFFTLTRDFG
RILPDISLRKMNLCGCWSVEELDASKCTELIKVVNFGKLEKLRELNLFGCKSTTELFDLSGLVSLDGLNLEGCTELIKVTSFEKLEMLRKLNMLGCKSIR
ELPDMSCLKNLHDLNLEGCAKLIKVTCFGKLEKLRKLNLFGCKSIRELPDLSGLVSLNDLNLEGCTKLVKITSYQKLEMLRKLNMAGCESIKELSNMSDL
KNLQELNMEGCIKLSKVRSFEKLETLRKLNMSGCESIKEFPDMSGLKNLHDLNLEGCTKLIEVTSFEKLEKLRKLNMSGCKSIKELPDMSGLMSIDDLNF
EGCTELVKVTNFEKMEMLQKLNMSGCKSIKELPDLSRLKNLHVLNLEGCTELIKVTSFGRLEMLQKLKMSGCQSIKELPDMSGLMSIDDLNFEGCTELVK
VTNFEKMEMLQKLNMSGCKSIKELPDLSRLKNLHVLNLEGCTELIKVTSFGRLEMLQKLKMSGCQSIKELPDMSGLMSIDDLNFEGCTELVKVTNFEKME
MLQKLNMSGCKSIKELPDLSRLKNLHVLNLEGCTELIKVTSFGRLEMLQKLKMSGCQSIKELPDMSGLMSIDDLNFEGCTELVKVTNFEKMEMLQKLNMS
GCKSIKELPDLSRLKNLHVLNLEGCTELIKVTSFGRLEMLQKLKMSGCQSIKELPDMSGLMSIDDLNFEGCTELVKVTNFEKMEMLQKLNMSGCKSIKEL
PDLSRLKNLHVLNLEGCTELIKVTSFGRLEMLQKLKMSGCQSIKELPDMSGLMSIDDLNFEGCTELVKVTNFEKMEMLQKLNMSGCKSIKELPDLSRLKN
LHVLNLEGCTELIKVTSFGRLEMLQKLKMSGCQSIKELPDMSGLMSRVERGGETQ
|
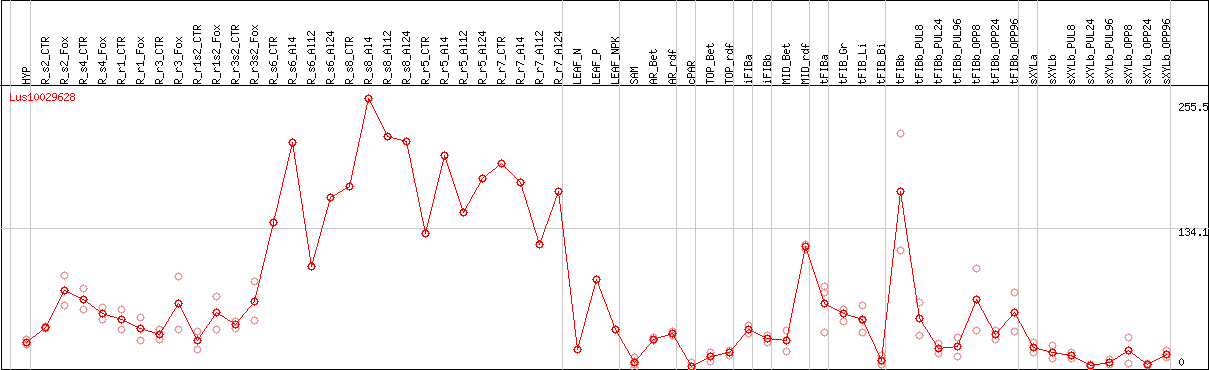
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029628 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
