External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G52020 114 / 1e-31
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT1G12630 94 / 4e-24
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT2G36450 83 / 4e-20
AP2_ERF
HRD
HARDY, Integrase-type DNA-binding superfamily protein (.1)
AT4G16750 82 / 2e-19
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT5G11590 81 / 1e-18
AP2_ERF
DREB3, TINY2
TINY2, Integrase-type DNA-binding superfamily protein (.1)
AT5G25810 80 / 1e-18
AP2_ERF
TNY, TINY
TINY, Integrase-type DNA-binding superfamily protein (.1)
AT2G44940 81 / 3e-18
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT2G35700 78 / 3e-18
AP2_ERF
ATERF38
ERF family protein 38 (.1)
AT4G32800 79 / 4e-18
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT4G25480 77 / 2e-17
AP2_ERF
CBF3, DREB1A, ATCBF3
C-REPEAT BINDING FACTOR 3, dehydration response element B1A (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031652
159 / 5e-50
AT5G52020 150 / 5e-46
Integrase-type DNA-binding superfamily protein (.1)
Lus10027413
155 / 2e-48
AT1G12630 146 / 3e-44
Integrase-type DNA-binding superfamily protein (.1)
Lus10031655
154 / 6e-48
AT5G52020 143 / 7e-43
Integrase-type DNA-binding superfamily protein (.1)
Lus10017830
98 / 2e-25
AT5G52020 137 / 7e-40
Integrase-type DNA-binding superfamily protein (.1)
Lus10024492
96 / 1e-24
AT5G52020 140 / 3e-41
Integrase-type DNA-binding superfamily protein (.1)
Lus10027415
88 / 1e-22
AT5G52020 78 / 2e-18
Integrase-type DNA-binding superfamily protein (.1)
Lus10023873
83 / 3e-20
AT2G36450 121 / 3e-35
HARDY, Integrase-type DNA-binding superfamily protein (.1)
Lus10014376
84 / 4e-20
AT2G36450 167 / 2e-52
HARDY, Integrase-type DNA-binding superfamily protein (.1)
Lus10002801
81 / 7e-19
AT5G11590 209 / 4e-68
TINY2, Integrase-type DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0081
MBD-like
PF00847
AP2
AP2 domain
Representative CDS sequence
>Lus10031656 pacid=23180652 polypeptide=Lus10031656 locus=Lus10031656.g ID=Lus10031656.BGIv1.0 annot-version=v1.0
ATGGCAGCCAACCAACCTCAAGACAATGCTCCTCCTCCTTCTCCAACCCAACAACAATCGCCGCCGTCCATCAGAAAGCAACCAAAGTACAGAGGAATCC
GTAGCCGGAGCGGGAAATGGGTGTCGGAGATCCGAGAACCCAAAAAAACGACCCGAATATGGCTTGGCACTTACCCAACCCCGGAAAAGGCTGCCGCAGC
ATACGACGTCGCAGCAATCGCCCTCAAGGGTCCCCACACCCCTCTAAACTTCCCAACCTCCTTCTACTCTTACCCTATCCCGGCCACTACCTCCCCTTCC
GACATCCGCTCTGCAGCAGCTCGTGCGGCTGCTCAAATCCAAGCATCCTCCTCGCAGGGCCCGGTGGTCCGGGCCCAGGAGGAGGAGTACGTGGACATGG
AGGACATGGAGGACATGCTGAACATGCCGAACTTGCTATTAGAAATGGCGGGAGGCATGATGGTTAGCCCGCCAAGGATGCACGATACCGCACCCTATGG
AGATTCTCCTGAAAATTCGGAGGGCGAATCCGTGTGGTCGTATCGACAGTTTTGA
AA sequence
>Lus10031656 pacid=23180652 polypeptide=Lus10031656 locus=Lus10031656.g ID=Lus10031656.BGIv1.0 annot-version=v1.0
MAANQPQDNAPPPSPTQQQSPPSIRKQPKYRGIRSRSGKWVSEIREPKKTTRIWLGTYPTPEKAAAAYDVAAIALKGPHTPLNFPTSFYSYPIPATTSPS
DIRSAAARAAAQIQASSSQGPVVRAQEEEYVDMEDMEDMLNMPNLLLEMAGGMMVSPPRMHDTAPYGDSPENSEGESVWSYRQF
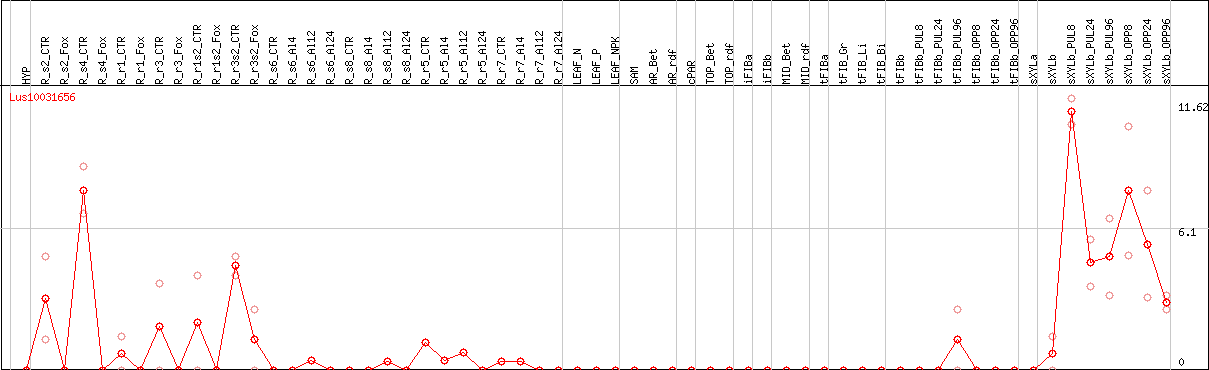
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031656 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.