External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G61830 431 / 5e-153
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G51030 416 / 5e-147
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G59710 226 / 1e-72
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G24190 176 / 4e-53
SDR2
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT3G61220 175 / 9e-53
SDR1
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G01800 169 / 2e-50
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G63380 69 / 4e-13
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G62610 67 / 1e-12
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
AT4G23430 67 / 3e-12
AtTic32-IVa
translocon at the inner envelope membrane of chloroplasts 32-IVa, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
AT4G09750 66 / 4e-12
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032524
437 / 2e-155
AT5G51030 493 / 2e-177
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10043023
437 / 4e-155
AT5G51030 495 / 3e-178
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10034835
221 / 2e-70
AT3G59710 335 / 2e-115
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10035736
212 / 6e-67
AT3G59710 355 / 2e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040933
176 / 3e-53
AT3G61220 342 / 1e-118
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10008413
171 / 7e-51
AT2G24190 273 / 3e-91
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014694
170 / 1e-50
AT3G61220 369 / 4e-129
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10037320
166 / 9e-50
AT3G59710 277 / 2e-93
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10014693
166 / 3e-49
AT3G61220 347 / 3e-120
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G110200
442 / 3e-157
AT5G51030 460 / 1e-164
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.015G108400
432 / 2e-153
AT5G51030 468 / 2e-167
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.013G127700
244 / 1e-79
AT3G59710 326 / 4e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.002G157000
184 / 2e-56
AT3G61220 406 / 9e-144
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.004G235500
176 / 5e-53
AT3G61220 313 / 7e-107
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.014G080000
164 / 2e-48
AT3G61220 332 / 2e-114
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156600
164 / 2e-48
AT3G61220 348 / 7e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.014G080100
164 / 3e-48
AT3G61220 350 / 3e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156800
163 / 6e-48
AT3G61220 363 / 1e-126
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156300
162 / 8e-48
AT3G61220 367 / 6e-128
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10032525 pacid=23146393 polypeptide=Lus10032525 locus=Lus10032525.g ID=Lus10032525.BGIv1.0 annot-version=v1.0
ATGGGAAAGGGAACCAAAGAGCGTGCGAAAGAGCGACGAGAAAAGATACGCCAAGAGATTTCCCTTCTCCGTACCATTCCCTACCCCGATCACCAAAGGT
GGTGGAACTCTGATACTGTGGCCGTAGTTACTGGTTCAAACAGAGGAATAGGGTTTGAGATTGCGAAACAGCTTTCCGGCCATGGAATTACTGTTATACT
GACATCCAGAAACCCTAATTCTGGAATTGAAGCTGCTAAGTTTTTGCAAGAATTAGGCCTATCTGTAGTGTTTCACGAGCTTGATGTCCTGCACACTTCG
TCTATCGATCGTTTTGCTGAGTGGATAAAACACGAGTATGGTGGTATCGACATTCTGGTGAATAATGCAGGGGTTAACTTCAATCTTGGTTCTGATAACT
CTGTCGAATTCGCCCAGCTGGTCATTGATACCAATTATCAGGGAACTAAGAACATGACCAAAGCCATGATCCCATTGATGAGACATTCTCCAGCTGGTTC
GCGAATCGTTAATGTGTCCTCCAGGCTGGGTAGAGTGAACGGGAGACGAAACAGGATTGGAGATGTAACGCTAAGAGAACAGCTAAGTGGTGCCGAGACA
CTTTCGGAGGAGTTAATTGACACGATGTTATCGAGTTTCCTCCAGGAAGTTGAAGACGGGACTTGGGAATCAAGCGGGAGGTGGCCTCAGGTGTTCACGG
ACTATGCACTGTCGAAGCTCGGAGTTAATTGTTTCACCAGGTTGATGGCTAAGTTGCTCTCTGACAGGCCCGATGGCGAGAAGATTTCCGTGAACAGTTT
CTGCCCCGGCTGGGTGAAAACTGCCATGACGGGTTGGAATGGTAACATATCGCCGGAGGAAGCTGCAGATACCGGTGTGTGGCTGTCGTTTCTGTCGGAG
GTTTCGACGGGGAAGTTCTATGCAGAGAGGCGTGAGATAAGCTTTTGA
AA sequence
>Lus10032525 pacid=23146393 polypeptide=Lus10032525 locus=Lus10032525.g ID=Lus10032525.BGIv1.0 annot-version=v1.0
MGKGTKERAKERREKIRQEISLLRTIPYPDHQRWWNSDTVAVVTGSNRGIGFEIAKQLSGHGITVILTSRNPNSGIEAAKFLQELGLSVVFHELDVLHTS
SIDRFAEWIKHEYGGIDILVNNAGVNFNLGSDNSVEFAQLVIDTNYQGTKNMTKAMIPLMRHSPAGSRIVNVSSRLGRVNGRRNRIGDVTLREQLSGAET
LSEELIDTMLSSFLQEVEDGTWESSGRWPQVFTDYALSKLGVNCFTRLMAKLLSDRPDGEKISVNSFCPGWVKTAMTGWNGNISPEEAADTGVWLSFLSE
VSTGKFYAERREISF
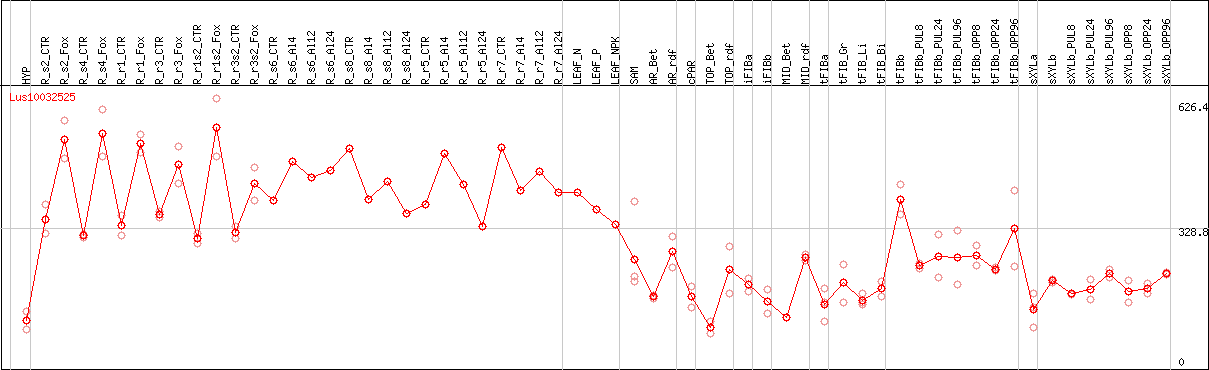
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032525 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.