External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G66900 114 / 2e-29
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT5G66630 110 / 3e-28
DAR5
DA1-related protein 5 (.1)
AT5G66910 97 / 2e-23
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G58410 47 / 2e-06
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G33560 47 / 2e-06
ADR1
ACTIVATED DISEASE RESISTANCE 1, Disease resistance protein (CC-NBS-LRR class) family (.1)
AT5G47280 47 / 3e-06
ADR1-L3
ADR1-like 3 (.1)
AT5G04720 47 / 4e-06
PHX21, ADR1-L2
PHOENIX 21, ADR1-like 2 (.1)
AT5G43470 46 / 9e-06
HRT, RCY1, RPP8
RECOGNITION OF PERONOSPORA PARASITICA 8, RESISTANT TO CMV\(Y\) 1, HYPERSENSITIVE RESPONSE TO TCV, Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
AT5G48620 45 / 1e-05
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G53350 44 / 3e-05
Disease resistance protein (CC-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034761
369 / 8e-128
AT5G66900 197 / 1e-54
Disease resistance protein (CC-NBS-LRR class) family (.1)
Lus10022464
167 / 3e-48
AT5G66900 531 / 2e-177
Disease resistance protein (CC-NBS-LRR class) family (.1)
Lus10020779
130 / 4e-35
AT5G66900 434 / 5e-141
Disease resistance protein (CC-NBS-LRR class) family (.1)
Lus10007359
125 / 2e-33
AT5G66900 473 / 3e-154
Disease resistance protein (CC-NBS-LRR class) family (.1)
Lus10021769
66 / 1e-12
AT4G33300 583 / 0.0
ADR1-like 1 (.1.2)
Lus10026765
39 / 0.0002
AT3G07040 71 / 2e-15
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10016316
40 / 0.0008
AT3G14470 494 / 6e-156
NB-ARC domain-containing disease resistance protein (.1)
Lus10032759
40 / 0.001
AT1G33560 543 / 0.0
ACTIVATED DISEASE RESISTANCE 1, Disease resistance protein (CC-NBS-LRR class) family (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G039000
167 / 2e-48
AT5G66900 343 / 2e-107
Disease resistance protein (CC-NBS-LRR class) family (.1)
Potri.007G039100
164 / 4e-47
AT5G66900 474 / 3e-156
Disease resistance protein (CC-NBS-LRR class) family (.1)
Potri.007G038700
141 / 7e-39
AT5G66900 481 / 3e-158
Disease resistance protein (CC-NBS-LRR class) family (.1)
Potri.007G039300
132 / 7e-36
AT5G66900 540 / 0.0
Disease resistance protein (CC-NBS-LRR class) family (.1)
Potri.014G035700
59 / 4e-10
AT4G33300 889 / 0.0
ADR1-like 1 (.1.2)
Potri.002G129300
57 / 9e-10
AT4G33300 902 / 0.0
ADR1-like 1 (.1.2)
Potri.003G201900
48 / 2e-06
AT3G14470 567 / 0.0
NB-ARC domain-containing disease resistance protein (.1)
Potri.003G149800
46 / 5e-06
AT5G43470 421 / 4e-133
RECOGNITION OF PERONOSPORA PARASITICA 8, RESISTANT TO CMV\(Y\) 1, HYPERSENSITIVE RESPONSE TO TCV, Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
Potri.003G150200
44 / 3e-05
AT5G43470 420 / 2e-132
RECOGNITION OF PERONOSPORA PARASITICA 8, RESISTANT TO CMV\(Y\) 1, HYPERSENSITIVE RESPONSE TO TCV, Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
Potri.006G014400
44 / 3e-05
AT3G50950 435 / 1e-139
HOPZ-ACTIVATED RESISTANCE 1 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Lus10033299 pacid=23172052 polypeptide=Lus10033299 locus=Lus10033299.g ID=Lus10033299.BGIv1.0 annot-version=v1.0
ATGCAGATCATTCAATTTGCAGACCAGAAAGAAACCTTGTTTGAAGTCAAACAGCTGAACAGCCAAATTAGGAAGTCCCTCCCTCGATCACTTTTAGGTC
TCTGCTCCCCTTCTGATTTGAAAGTTGATCCAGTAGGGTTAGAAACTCCGTTGATGGAGTTGAAAGAACAGCTGCTCGGAGATGAGCTCCCACTTGTTGT
CATTTCTGCTCCTGATGGATGGGGGAAAACCACTCTCGCTACAGCAATCTGCCAGGATACACAAGTTAAAGATAGATTTAAGGGGAATATATTATTTGTC
ACTGCTGGGAGAAGTCCTAACATGATGGCTATAGTGGGTAGACTTCTTGAGCACTACAATTGCCCACTACCTGAACTACAGAGTGAAGATGAAGCTATCT
TTGAACTAGAGAACCTTATGAGGAAGATACAATCTCAGCCGGTTCTTTTGGTAATTGATGATCTTTCGGCTGGATCAGAATCTCTTCTTCTGAGACTCAA
GTTCCAAATACAGAATTACCAGATTTTGGTGACCTCTAGTTCCGAGTTCCCCAGAATTAGTTCCACATACAAAATGTAA
AA sequence
>Lus10033299 pacid=23172052 polypeptide=Lus10033299 locus=Lus10033299.g ID=Lus10033299.BGIv1.0 annot-version=v1.0
MQIIQFADQKETLFEVKQLNSQIRKSLPRSLLGLCSPSDLKVDPVGLETPLMELKEQLLGDELPLVVISAPDGWGKTTLATAICQDTQVKDRFKGNILFV
TAGRSPNMMAIVGRLLEHYNCPLPELQSEDEAIFELENLMRKIQSQPVLLVIDDLSAGSESLLLRLKFQIQNYQILVTSSSEFPRISSTYKM
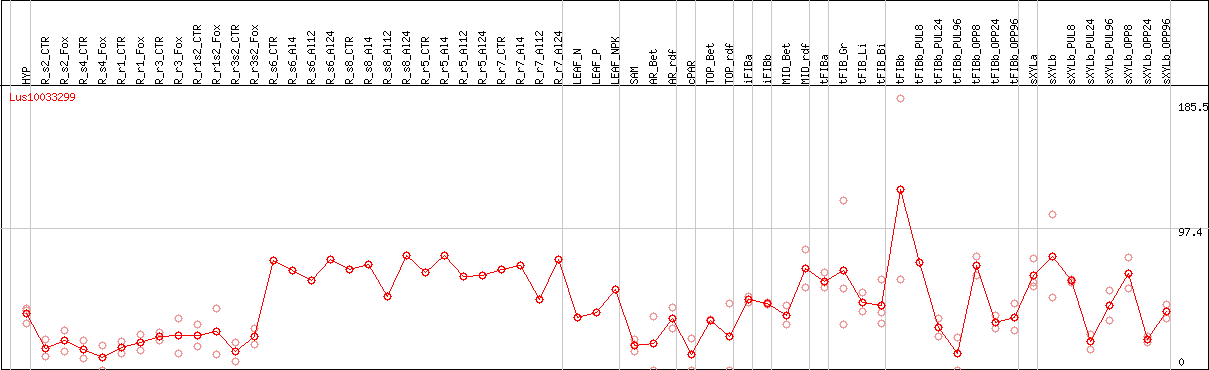
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033299 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.