Lus10036347 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10036347 pacid=23174634 polypeptide=Lus10036347 locus=Lus10036347.g ID=Lus10036347.BGIv1.0 annot-version=v1.0
ATGGTGGAGACTCGCCGCAGCTCTTCCGCCTCCAAGCGCCCAATTCATTCCTCCCCTCCTCCACCATCCAAACGATCCAAGTCGGCGGCGGCTGTTGCTG
CGGCGGCGGCGGAGGCTTCTTCATCGACGAACGATGACGCACTGGTGTCTGTGCCGGTTGAGACCCTAGGCCTGGCTAAGGATTCTGACACTGTTGTTGT
CCCTCCGGAGCTGCAATCCTCTGATCCGCAGGTCACTACTCCTGCTTTAAAAGATAGTGCTGCAGCAGCTGCTAAATCCCCTGATGCGGATGTTGTTGAT
GTGGAAGGTGAGGATTTGATCTGCCCGCCGCCTCAAGGTGGAGCTGCAGTGGAGATGGAGAGAGCCAAGTTTGTGAGTGGACCGGCTACAACTACCCGGT
CTAAGAAGGGGCAAAAGAAGTCGGAGAAGCTGGATCCTCAAGTTGCATGGGCAAAGCTTCTTTGCCAGTATTCTAAGTGTGGAGGTGTTGTTGCTGCCAT
GTTGGAAGTTGTTGGGGGTAAAGGTACTGTTCATGTCAACAATAAGATTCAGCGGAAAGGCACCCGTGTAATGTTGAATGCAGGAGATGAGTTGGTTTTC
AACGCTTCTTTGGAGCATGCTTATATCTTTCAGCAGCGCCCCAATGATACTTTGGGAGGTCGAGGAATAGCTGCATTGAGCATTTTAGAAGCTCCGGGTG
CTGCAGGCAAAGGGATCCCTCTGGAGGCGAGATCTGGAGATACGTCAGCCTTCGCAGGCGCATCAATATTGGCATCTTTGTCGAATTTTCGCAAAGATTT
ATCTCTTCCACCTCCTTCTAAAGCTGTCGAGGACACACAAAAGAAACCTGATAATTCTATTCTATCCCCTCTCAGTGTAGTGCCCGGTGATTGTATTCCT
GAAGTTGGCATGAAGGATAACACAAACAATAATGATCAACCTGTGGTTTCCTCAAGGAAGGACAATGCTGTTACATCTGGCAATTGTGCCTCTGAAAGCC
CCAATCTTGAACCTCTTCCATTAGACAGTTGCACAGATGCAGAAACTGGGAATAACCCAGCCTCTAATTATGAATTAAGGCCTCTTCTGCAGATGCTTTC
TGGTTCTTCTTGTGAACTAGATATAAGTGGAAGCATTTCAAAGATACTTGAAGGACGCAGGGAGTTGCTCAAAGATATTGATCCCTCTGCCGTTTTGCTT
TCTTCTGGTAGGCAGGCATACAAAGACAGTCTAGAACAAGGAATCTTAAGTCCTGAGAACATCGAGTTCTCATTTCAAAATTTCCCTTATTACCTTGGTG
ATACAACAAAGAAAGTCCTTCTGGGTGCTGCATTTGTTCACTTGAAGTGTAATAACAAAGTAGCTAAATTTGCTTCTGATCTCCCCACTATATCACCTCG
AATGCTGTTGTCTGGTCCTGCAGGCTCAGAGATATACCAGGAGAGACTTGTGAAGGCCCTTGCGAAAGAGCTTGGAGCTAGACTACTGGTTGTTGATTCT
CTACAATTGCCTGGCGGATCAACCCCCAAAGAAGCTGATAATGTGAAAGAAACTTTTAGACCGGAAAGAGCATTATTTGCCAAAAGAGTAATGCAATCAG
CACTACACCATAAGAAACCAGCTTCTAGTGTGGAAGCTGGTATTACAGGTGGATCCGTCATAAGTTCTCACGCGCTGCCGAAGCAGGAAACTTCTACTGC
CTCATCTAAACATTCCGCATTTAAAACAGGCATATTAACTTTCCTATGTGATAGAGTGAAGTATGTTGGTTCCTCTTCACCGTCTTCTCTGTCTCCTCTT
CAACCTCCATTGAGGGGACCTTCAATTGGGTTACGCGGCAAGGTACTACTTGCTTTTGAAGAAAACGGGTCGTCTAAGATTGGTGTGAGGTTTGATAGGC
CCGTCCAAGAAGGAAATGATCTTGGTGGTCTTTGTGAAGAAGAACGTGGTTTCTTTTGTGCTGCCAATTCACTTCGCTTGGATGGCTCTGGAGGTGAAGA
GTCTGACAGGCTTGCTGTTAATGAACTCTTCGAGGTTGTTTTAAGTGAAAGCAAAAGTGGTCCATTGATAGTTTTTCTGAAAGACATAGAAAAGTCTGTG
GCGGGCAATCCAGATGCATATACTGCCTTGAAAAGCAAGATGGAAAATTTACCAGATAAGGTTATTGTGATTGGATCCCATGCTCAGATGGACAGTCGAA
AAGAAAAGTCTCAGCCTGGTGGTCTTCTGTTTACCAAGTTTGGAAGCAACCACACTGCTTTGCTTGATTTTGCATTTCCAGAATACATAATGAAAATTGA
TCCTATGATCTCTTCTCGTCAGGATCATTTAGGTAGACTACATGAAAAGAGCAAGGAGACTCCTAAAACAATGAAGCAGCTTTCAAGACTTTTTCCCAAC
AAAGTGGCCATTCAGATACCTCAGGATGAAGGATTGCTTTTGGACTGGAAGCAGCAGTTGGATCGTGATATTGAAACTCTCAAAGTACAGACTAATATTG
TGACTATTCGTTCAGTCCTTACTCGAGTTGGCCTGCAATGCCCTGATCTCGAGTCCCTGTCCATTAATGACCAGGCCTTGACAAGCGAAAGTGTGGAGAA
AATAGTAGGTTGGGCATTAAGCCACCACTTCATGCATTGTTCCTCGACTTCACCGACAGAAGCTAAATTATCAATATCCACAGAAAGTATTAACTATGGA
TTGAGTATTCTACATGGTATACTTAATGAAAACAAGAGCTCGAAGAAATCTCTCAAGGATGTCGTCACTGAGAATGAATTTGAGAAGAAACTTCTAGGAG
ATGTCATTCCTCCCAACGACATTGGAGTTTCATTTGACGACATTGGTGCCTTGGAAAATGTGAAGGAAACATTAAAGGAGCTCGTCATGCTTCCTCTTCA
GAGGCCTGGATTGTTCTGCAAAGGGCAGCTTACTAAGCCTTGCAAGGGGATATTGCTGTTTGGACCTCCTGGTACTGGGAAAACAATGCTGGCGAAGGCT
GTGGCAACTGAAGCTGGTGCAAATTTCATCAACATATCGATGTCAAGCATTACATCGAAGTGGTTTGGCGAAGGAGAGAAGTACGTCAAGGCCGTGTTCT
CTTTAGCTAGCAAAATTGCTCCTAGTGTTATATTCGTCGACGAGGTGGATAGTATGTTAGGAAGGCGTGAGAATCCTGGAGAACATGAAGCAATGCGTAA
AATGAAGAACGAATTCATGGTGAACTGGGACGGTCTGCGTACGAAAGATAAAGAAAGAGTGTTGGTCCTTGCTGCCACAAATCGACCGTTCGATCTTGAT
GAAGCTGTTATTCGGAGACTTCCTCGAAGGTTGATGGTAAACTTGCCTGATGCATCAAATAGAGGAAAGATTCTGAAGGTGATCTTAGCTAAAGAGGAAC
TGGCAGGAGATGTGGATCTGGAAGCAGTTGCAAGTATGACTGATGGCTATTCCGGAAGTGACTTGAAGAACCTGTGTGTGACTGCAGCTCATTGTCCCAT
AAGAGAGATATTGGAGAAAGAGAAGAAGGAAAAAGACGTGGCACTTTCTGAGAAAAGACCTGTGCCTGACTTGTACGGCGGTACAGATGTTCGCCCGATG
AAAATGGAGGATTTCAGATATGCTCATGAACAGGTTTGTGCAAGTGTGTCATCTGAGTCGAGTAACATGAGTGAGTTGCTACAATGGAATGATCTGTATG
GAGAAGGTGGATCGAGGAAGAAGAAGTCATTAAGCTATTTCATGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10036347 pacid=23174634 polypeptide=Lus10036347 locus=Lus10036347.g ID=Lus10036347.BGIv1.0 annot-version=v1.0
MVETRRSSSASKRPIHSSPPPPSKRSKSAAAVAAAAAEASSSTNDDALVSVPVETLGLAKDSDTVVVPPELQSSDPQVTTPALKDSAAAAAKSPDADVVD
VEGEDLICPPPQGGAAVEMERAKFVSGPATTTRSKKGQKKSEKLDPQVAWAKLLCQYSKCGGVVAAMLEVVGGKGTVHVNNKIQRKGTRVMLNAGDELVF
NASLEHAYIFQQRPNDTLGGRGIAALSILEAPGAAGKGIPLEARSGDTSAFAGASILASLSNFRKDLSLPPPSKAVEDTQKKPDNSILSPLSVVPGDCIP
EVGMKDNTNNNDQPVVSSRKDNAVTSGNCASESPNLEPLPLDSCTDAETGNNPASNYELRPLLQMLSGSSCELDISGSISKILEGRRELLKDIDPSAVLL
SSGRQAYKDSLEQGILSPENIEFSFQNFPYYLGDTTKKVLLGAAFVHLKCNNKVAKFASDLPTISPRMLLSGPAGSEIYQERLVKALAKELGARLLVVDS
LQLPGGSTPKEADNVKETFRPERALFAKRVMQSALHHKKPASSVEAGITGGSVISSHALPKQETSTASSKHSAFKTGILTFLCDRVKYVGSSSPSSLSPL
QPPLRGPSIGLRGKVLLAFEENGSSKIGVRFDRPVQEGNDLGGLCEEERGFFCAANSLRLDGSGGEESDRLAVNELFEVVLSESKSGPLIVFLKDIEKSV
AGNPDAYTALKSKMENLPDKVIVIGSHAQMDSRKEKSQPGGLLFTKFGSNHTALLDFAFPEYIMKIDPMISSRQDHLGRLHEKSKETPKTMKQLSRLFPN
KVAIQIPQDEGLLLDWKQQLDRDIETLKVQTNIVTIRSVLTRVGLQCPDLESLSINDQALTSESVEKIVGWALSHHFMHCSSTSPTEAKLSISTESINYG
LSILHGILNENKSSKKSLKDVVTENEFEKKLLGDVIPPNDIGVSFDDIGALENVKETLKELVMLPLQRPGLFCKGQLTKPCKGILLFGPPGTGKTMLAKA
VATEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKIAPSVIFVDEVDSMLGRRENPGEHEAMRKMKNEFMVNWDGLRTKDKERVLVLAATNRPFDLD
EAVIRRLPRRLMVNLPDASNRGKILKVILAKEELAGDVDLEAVASMTDGYSGSDLKNLCVTAAHCPIREILEKEKKEKDVALSEKRPVPDLYGGTDVRPM
KMEDFRYAHEQVCASVSSESSNMSELLQWNDLYGEGGSRKKKSLSYFM
|
DESeq2's median of ratios [FLAX]
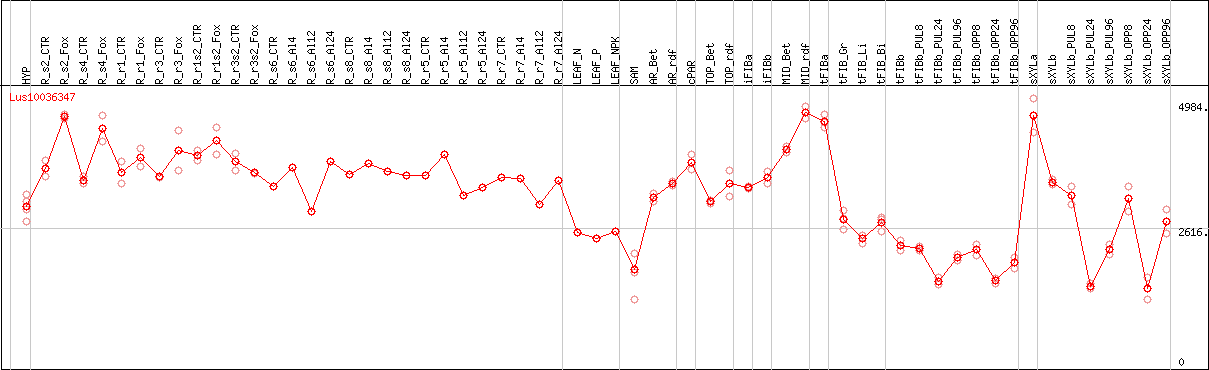
Coexpressed genes
Lus10036347 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
