External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G49330 228 / 7e-73
MYB
PFG3, ATMYB111
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
AT2G47460 207 / 2e-64
MYB
PFG1, ATMYB12
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
AT3G62610 196 / 2e-60
MYB
PFG2, ATMYB11
PRODUCTION OF FLAVONOL GLYCOSIDES 2, myb domain protein 11 (.1)
AT3G23250 187 / 1e-57
MYB
ATMYB15, ATY19
myb domain protein 15 (.1.2)
AT4G38620 186 / 1e-57
MYB
AtMYB4
myb domain protein 4 (.1)
AT4G09460 185 / 1e-57
MYB
ATMYB6, ATMYB8
myb domain protein 6 (.1)
AT2G16720 184 / 6e-57
MYB
AtY49, AtMYB7
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
AT1G35515 182 / 1e-56
MYB
MYB8, HOS10
high response to osmotic stress 10 (.1)
AT2G31180 182 / 2e-56
MYB
ATMYB14, Myb14at
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 14, myb domain protein 14 (.1)
AT1G22640 181 / 7e-56
MYB
AtMYB3
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016855
225 / 4e-72
AT5G49330 242 / 3e-78
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Lus10010273
201 / 2e-62
AT2G47460 221 / 3e-69
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
Lus10036336
199 / 1e-61
AT2G47460 225 / 9e-71
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
Lus10001458
191 / 6e-59
AT2G47460 212 / 5e-66
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
Lus10001548
189 / 1e-58
AT4G09460 291 / 2e-99
myb domain protein 6 (.1)
Lus10033889
189 / 4e-58
AT3G23250 239 / 1e-78
myb domain protein 15 (.1.2)
Lus10009448
188 / 4e-58
AT4G09460 290 / 2e-99
myb domain protein 6 (.1)
Lus10002435
188 / 1e-57
AT2G47460 206 / 9e-64
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
Lus10000411
187 / 1e-57
AT4G09460 300 / 5e-103
myb domain protein 6 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G141000
219 / 2e-69
AT5G49330 248 / 7e-80
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.002G198100
206 / 1e-63
AT5G49330 225 / 2e-70
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.014G122700
194 / 3e-59
AT5G49330 214 / 4e-66
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.019G081500
190 / 7e-59
AT1G22640 277 / 1e-93
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
Potri.005G224100
189 / 2e-58
AT3G23250 255 / 1e-84
myb domain protein 15 (.1.2)
Potri.019G018200
186 / 4e-58
AT2G31180 223 / 1e-73
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 14, myb domain protein 14 (.1)
Potri.003G144200
187 / 6e-58
AT5G49330 205 / 2e-64
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.014G100800
187 / 6e-58
AT2G16720 207 / 3e-66
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Potri.004G174400
186 / 1e-57
AT4G38620 305 / 2e-104
myb domain protein 4 (.1)
Potri.001G086700
186 / 1e-57
AT5G49330 203 / 6e-64
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10036453 pacid=23174544 polypeptide=Lus10036453 locus=Lus10036453.g ID=Lus10036453.BGIv1.0 annot-version=v1.0
ATGGGGAGAGCTCCTTGCTGCGAGAAAGTAGGGTTGAAGAGAGGAAGGTGGGCGGCGGAGGAGGACCAGATCTTGACTGCTTACATCCGCCTGCACGGAG
AAGGAAAATGGCGATCCCTCCCCAAGGACGCCGGGTTGTTGAGGTGCGGGAAGAGCTGCAGACTGAGGTGGATTAACTACTTGAGAGATGACTTGAAGCG
TGGCAATATTACTAAAGATGAAGAAGAAGCCATTGTTAAGCTCCACAGCTCCCTTGGCAATAGGTGGTCGTTGATAGCAGCCCAATTATCTGGAAGAACA
GACAATGAGATCATGAACTACTGGAATTCTCACCTTCGCCGGAAAATCTACACCTTCCACACCGTCGGCCAACCCCATTCTCCCTGTCTGGAATCATCTT
CATCTGCCTCTTCGTCCTCCACCTCTCCCTCGTCCACACGCAGTAGAAACAGTAAGAGGCCGCCATCCCTGGTCACCGCCGGCGGCGGCAGTAAAGAAGA
TCACGGCGAAATCGCAGCCTGTGGGGACCAACTACAGGAATTAAGAATGGATGAAGAGAAAGTTGCAGAGGATCATGCGGCGGGACCATATGAATGGCTT
GACAGCGAGATGAACCGCCTAATTAAAAGCAGAGTAGTACTCCATGAAGATTCGCCCAGTGGGGAAATGAAGGATGCTGATGATCATGGAGGAGGAGGAG
GAGGAGGGAGCAGCTGCAACAGTAATAGTAGCGATAATCAGCGCTCTTCCTCTGTTTCGCAACAATGGCTGGATTGGGAACAAATTACCGGAGGAGGGGA
GATTAATTATAGTAGTAATCATCATCATCATCATCATAATGATGTTGTGTGGGACGAAGGAGAGGATGACGTCCTTTTGTCTTGGCTATTGGATGGTGTT
GATCCATAA
AA sequence
>Lus10036453 pacid=23174544 polypeptide=Lus10036453 locus=Lus10036453.g ID=Lus10036453.BGIv1.0 annot-version=v1.0
MGRAPCCEKVGLKRGRWAAEEDQILTAYIRLHGEGKWRSLPKDAGLLRCGKSCRLRWINYLRDDLKRGNITKDEEEAIVKLHSSLGNRWSLIAAQLSGRT
DNEIMNYWNSHLRRKIYTFHTVGQPHSPCLESSSSASSSSTSPSSTRSRNSKRPPSLVTAGGGSKEDHGEIAACGDQLQELRMDEEKVAEDHAAGPYEWL
DSEMNRLIKSRVVLHEDSPSGEMKDADDHGGGGGGGSSCNSNSSDNQRSSSVSQQWLDWEQITGGGEINYSSNHHHHHHNDVVWDEGEDDVLLSWLLDGV
DP
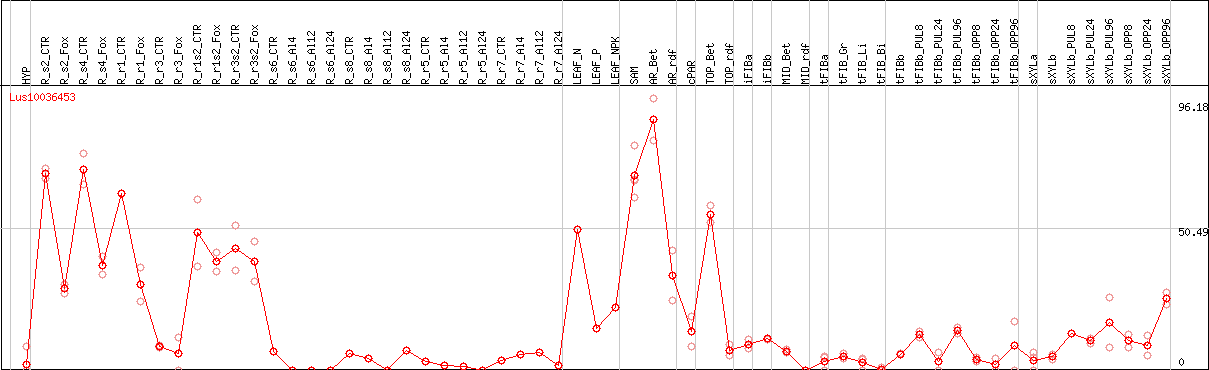
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036453 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.