Lus10036515 [FLAX]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Lus10036515 pacid=23174444 polypeptide=Lus10036515 locus=Lus10036515.g ID=Lus10036515.BGIv1.0 annot-version=v1.0
ATGCCGCCGGAGCCATTGCCCTGGGACCGGAAGAAGCACGAGTCGTCTTCTTCTTCATTAGGGTCAACCCCTCGCTGGAGGGACTGCCCCGCGGGTCACT
ATGGTTCCCACCGTGACTTTTCTCCTCGCTGGGCTGGAGGATCCACTGAGTTCCGCAGACCTTCCGGTCATGGTAAGCAAGGCGGCTGGCACCTGTATGC
TGATGAATCTGGTTATGGGTATGGCCCTTCCAGGTCCAACGACAAGATGAAGGAAGCTACTAATGGAAGGCCCCTATTTTCACGAGGGGATGGAAAGTGT
GGCAGGAGTGGTAGAGATAACAGAGGGCCTTATGGTCAGAGAGACTGGGGAGCATATTCATGGGATATGAGCAATGGATCCACGAATGGACCTAGAAGAC
CCTATGATGTGAGTAATGATCAGATGTCAGTTGATGATATGCTAACATGTCCCCCTTCACATCCTCCTAACTCTGACTTTGGAAGCACATGGGATCAGCT
TCAGTTAAGACAGCAGCATGGCAATAATGAGGATGGAGGCAACGCCTTGGGCTCTGGCCAAAGGGGTGATAAAGATAACTCATTGGATTGGAGGCCTCTA
AAGTGGACTCGTACAGGAAGCTTGTCTTCCAGGGGGTCTGGATATAGCCACTCCAGCAGTTCAATGAATCTTGGGGCTGTGAAGGCTAATGGTGGAAAGA
ATGAGATGATACCGAAGAATGGGACCCCAATTCATTCCACTTCAGGAGATGTGACTGTTGTCCGTGCTGATTCAGTGACATTGTCTGATGAGACTATGTC
AAGAAAAAAGCCACGCCTTAAATGGGGTGAGGGACTGGCGAAGTATGAGAAGAAAAGAGTTGAAGGACCTGAGACTAGTGTGACCAGAGATGATACCACC
TTGTTTACTAATAGCCTGGAACCTGTTCACTCGAGAAGCTCAAGCCTGCCTGACAAAAGCCCTAGGCAGATGGTTTATCCAGAATGTGTTTCTCCTGCAA
CTCCTTCCTCCATTGCTTGTAGTTCCTCGCCAGGTGTTGAAGATAAAGTTTTTTGTCCAGTAGTCAGTGTTGATAATGATGCTAGTAATTTATGTGGTTC
TCCTAGTGTTGGCTCTCAAGGCCGTGAGGAGTTTTCTTTCAGTTTAGAAAAGCTGGATATGGGTTCTATTCCTAATTTGGGGTCCTCACTCATTGAGATG
ATTCAGTCAGAAGATTTATGTATAGTGGACTCTTGTTTTGAGAAGTCAACTGGTATGAATAAGTTGCTTGTTTGGAAAGGCAATGCTTTGAAGGCATTGG
AAACAACAGAGACTGAGATAGATATTCTTGAAAATGAACTAAAGTCACTGAAGTCTGATTCAAAAAGCAAGCTTTTGGGCTCATCGCCGTCTGATTCCCT
AACAATAAGTGGTCATGCCATTGCTTGCAGTGAACAGGGAACTTTATGCCGTGATGACCTATGTCGTCCTGTATCCAGTATGGATGAAGCTGTGGATGGA
ATTCCTAATCCGGATGATGCTTTCGAAGATGGTTGTGCCATGGATGACGACATTGATAGTCCTGGGACTGCTACATCTAAGTTTGTTGAACCTTTGATAT
CTGTGTCTGATATGGTGAGATGCACTCAACATTCTGGTGTTGCTGCAGATGAATCTTCAAATTTGATGGTGGAGAAAGGAAGTGCGCTTGCAGCATGCAA
TGTTAGTAGCATACACTGCAACAATAATGCACTTCTCTCTGCTAGTAAGAGTTCTGTGGAAGACGGATACTCTAACCTATGTACTGCTATATTAGCCTCA
AATAAGGAATCTGCTAGTACAGTAGGTGAAGAATTTTATAGGTTATTGCCTAGGGATCGTTGCTCGGTTAATCATTCAGGAGTTGCGAATATATTATCCG
GGCAAACTGATGCCTTGGTTATGAAAAAATTCTTAGCGAGGAGACGCTTGATGAAGATTAAGGAGAAAATTGTGGCCCTGAAGTTCAAATCATTACAACA
TTTGTGGAGGGAGGATATGCACTTGCTCTCTGTAAATAAGTTCAAGGCAAAATCTCAGAAGAAATTTGAGTCAAGCTTGCGGCCTATTTATAGTATCCAT
AAGAAGCACCGGTCTTCATCTCGCATGCGAATTTGTTCTCCTGGTAGTATGGGCCTGGTTCTTTCTCCGGAGATGCTAAATTTCAGTAGAAAGTTGCTTG
CAGATTGCAGTGTCAAAGTTTACAGGGACAAGTTGAAAATGCCAATACAGATTTTGGATGACAGCGAGAGGAGAGTATCAATGTTCATTTCTGGCAATGG
ATTAGTTGAAGATCCCTTTGCTGTTGAGAGGGAAAGAGCAATGATAAATTGTTGGAGTTTGGAAGAGCGTGAAATCTTTATTGATAAGCTAGCCATATGT
GGAAAGGATTTTAAGAAAATTTCCTTCTATCTTGAACATAAGACTACTGCAGACTGTATTGAATTTTACTACAAAAATCATAAGTCTGACTCTTTTGGGA
AAGTCAAGAAGAACAAGCAATACAAGTCTTTAACAAATTACTTGATGTCATCCGGTAAGAAGTGGAATCGTGAGATGGGCACAGCTCCACTTGATACGTT
AGGTGCTGCCTCGGTGGTGGCTGCTCATCCTGACAGGAAAATGGGAAGTAGGAAAGTGTCTTCTAGTAAAATCTTTATCAGAGGCTATGATAAATGCAAA
ATCTTGCAAGGTGATGATGGTAATTCAGAAAGGTTAAACAACTATGATGTCCTTGTGAATGAAAGAGAAACTGCTGCTGCTGATGTGCTAGCTGGTATCA
GTGGTTCTATGAGTTCTTGTATCACTACTTCTGCGGATCGTGCTGAGGGGTTTTACGACTATAATGCACGGAGGACAGATACCACTGATAGACGTCCCTC
AATATCTGATGTAACCCAGAATGTCGATGAAGAGACATGTTCTGATGAGAGCTGTGGCGAAATGGATCCTGCTGATTGGACAGATGAGGAGAAAACAGCT
TTTGTTCGGGCCGTTTCTTTGTACGGAAAGGATTTTGCATTTATTTCCCGTTGTGTACGGACAAGGTCTCAGGAGCAGTGTAAGGTGTTCTTTAGTAAAG
CAAACAAGTGTCTTGGGTTGGACCATATTCACTCTGGCATAGGAAAGTTAGACCCATCATTAAGTGAGGATGCAAATGAAGGTGGGAGTGACATGGAAGA
TGCCACTGTGGATGGCTCTGAAGAAATTCCTTCTAAGTTGGACCTGGACCCACTCCCTCGAGTTATGTGTACTGCTGGGATTTTAAAGCAGGATGTTTCT
GATGGTAGGGTTGGAATCAATCTTGAAACTGATTTGAATGTATCCGATGAGAAAGATGTTGCAGGAGCTATAGAAGAGGAGAATTTGATGGCTGTAGATA
CTGTGAAAGGTCACTTAGTCCAGTCAACTCATCTTGCTGATGGTGAAGTCATTAACGGTTGTGGTCGTGTTTCTCTTCGAATGAATGGTCAGACAGTTCA
AATAGAATCTGGTGTTACAGAAGGTGAAAGAGATTGTCAAACCAAAGAAGTCCAACATCTTCCTTTGGATATGGGCTCGTCTTCAGATGACAGTCTTGTG
GGAAATATTACTCAGAGACAAGTTGAAGTGAATTCTGTGGAGAAGAGTTCGTCAATGAAAGTGAATAATAATTTGGTTAGTGCTGTTTCTGTGACATTGA
ACTCTGCTGCGACACACTGTGATAAAGCGCTCGATTCGGGGGAGATCTCAGGCAGTGTACAAGAGAATTGTAGTAAGCTGGATGGAAGTGGCTATGTGCA
ACCTTCCGTATGTTGCCCTGTGGTGAACCATGGGGAGTATTCTGAGAGTGGCTGTAGTAAAGTTCAAACTCTCACGGAGAAAGTGACCATGGGGAATACT
TCTGATGCTAAGCTCCTTTTGAACGCTGAACAGTTTATGGGCAACCAGTTGGTAGCTGAGGAGGGCTCCTGTCTTGAGAATTCCAGTAATTGTGAGGGTC
AATGTTCGGTTCCTGAACTTCCCCTTCTGAGCCAGCCTGGAAGTGCAACGAACATGGAAAAACGAAGCAGGAATGGAGATGTTAAACTATTTGGGAAAAT
ACTGAGTAATCCTGTATCATCGTCATCATCACAAAAGCCAGTTGCTGCACATGGGAGTGGTGAAAAGAGCACGCAGCCTCAGAGAACCTTAAAATTCAAT
GGTGATCACAGCGCAGATGATACTTTAGTGAATTTTGATAAAAATAATAGTGCAGGAGTGAAAAATGTTCCGATTAGGAGTTACGGTTTCTGGGATGGAA
GCAGAATACAGACAGGGCTGACTTCGTTGCCGGATTCAGCCATTCTACTTGCCAAGTATCCTGCTGCATTCACCAATTTTCAGGGGTCATCCTCACCGAA
GCTTGATCCGGTAGCACTGGCGACAACTTCCAAGAACATTGAACGAAGTATGAATGGAGTAGTTAGTATCCATCAAAGGGGGGACATAAACAGCAGCAGC
AGCAGTAGTAATGCACTGCTAGATTTCCAGAGGTACATAAGTACAGACGGTAGTAAGAAAGTGGAGCAGCAGGAGGGAGGGAAAGGGTTGGAGTCGAGGA
CGGTTTCGGATCCAGTGGCAGCGATTAAGATCCATTTCGCAAAGGCTGCTGATCAAAATTGCGGAGATAGTGAGGCTCCTGCTCAAGGGGCAATGGTAAA
ATAG
|
||||||||||||||||||||
|
AA sequence
|
>Lus10036515 pacid=23174444 polypeptide=Lus10036515 locus=Lus10036515.g ID=Lus10036515.BGIv1.0 annot-version=v1.0
MPPEPLPWDRKKHESSSSSLGSTPRWRDCPAGHYGSHRDFSPRWAGGSTEFRRPSGHGKQGGWHLYADESGYGYGPSRSNDKMKEATNGRPLFSRGDGKC
GRSGRDNRGPYGQRDWGAYSWDMSNGSTNGPRRPYDVSNDQMSVDDMLTCPPSHPPNSDFGSTWDQLQLRQQHGNNEDGGNALGSGQRGDKDNSLDWRPL
KWTRTGSLSSRGSGYSHSSSSMNLGAVKANGGKNEMIPKNGTPIHSTSGDVTVVRADSVTLSDETMSRKKPRLKWGEGLAKYEKKRVEGPETSVTRDDTT
LFTNSLEPVHSRSSSLPDKSPRQMVYPECVSPATPSSIACSSSPGVEDKVFCPVVSVDNDASNLCGSPSVGSQGREEFSFSLEKLDMGSIPNLGSSLIEM
IQSEDLCIVDSCFEKSTGMNKLLVWKGNALKALETTETEIDILENELKSLKSDSKSKLLGSSPSDSLTISGHAIACSEQGTLCRDDLCRPVSSMDEAVDG
IPNPDDAFEDGCAMDDDIDSPGTATSKFVEPLISVSDMVRCTQHSGVAADESSNLMVEKGSALAACNVSSIHCNNNALLSASKSSVEDGYSNLCTAILAS
NKESASTVGEEFYRLLPRDRCSVNHSGVANILSGQTDALVMKKFLARRRLMKIKEKIVALKFKSLQHLWREDMHLLSVNKFKAKSQKKFESSLRPIYSIH
KKHRSSSRMRICSPGSMGLVLSPEMLNFSRKLLADCSVKVYRDKLKMPIQILDDSERRVSMFISGNGLVEDPFAVERERAMINCWSLEEREIFIDKLAIC
GKDFKKISFYLEHKTTADCIEFYYKNHKSDSFGKVKKNKQYKSLTNYLMSSGKKWNREMGTAPLDTLGAASVVAAHPDRKMGSRKVSSSKIFIRGYDKCK
ILQGDDGNSERLNNYDVLVNERETAAADVLAGISGSMSSCITTSADRAEGFYDYNARRTDTTDRRPSISDVTQNVDEETCSDESCGEMDPADWTDEEKTA
FVRAVSLYGKDFAFISRCVRTRSQEQCKVFFSKANKCLGLDHIHSGIGKLDPSLSEDANEGGSDMEDATVDGSEEIPSKLDLDPLPRVMCTAGILKQDVS
DGRVGINLETDLNVSDEKDVAGAIEEENLMAVDTVKGHLVQSTHLADGEVINGCGRVSLRMNGQTVQIESGVTEGERDCQTKEVQHLPLDMGSSSDDSLV
GNITQRQVEVNSVEKSSSMKVNNNLVSAVSVTLNSAATHCDKALDSGEISGSVQENCSKLDGSGYVQPSVCCPVVNHGEYSESGCSKVQTLTEKVTMGNT
SDAKLLLNAEQFMGNQLVAEEGSCLENSSNCEGQCSVPELPLLSQPGSATNMEKRSRNGDVKLFGKILSNPVSSSSSQKPVAAHGSGEKSTQPQRTLKFN
GDHSADDTLVNFDKNNSAGVKNVPIRSYGFWDGSRIQTGLTSLPDSAILLAKYPAAFTNFQGSSSPKLDPVALATTSKNIERSMNGVVSIHQRGDINSSS
SSSNALLDFQRYISTDGSKKVEQQEGGKGLESRTVSDPVAAIKIHFAKAADQNCGDSEAPAQGAMVK
|
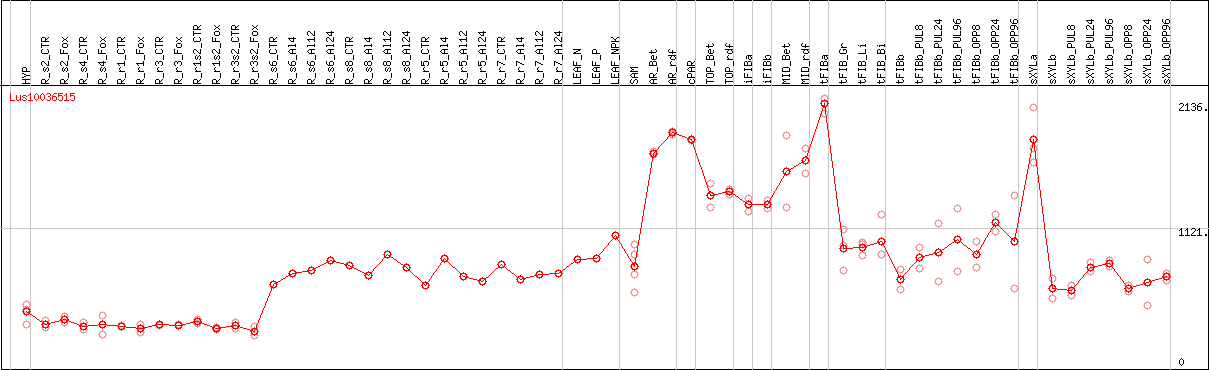
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036515 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
