External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G45050 105 / 2e-26
WRKY
ATWRKY16, TTR1
TOLERANT TO TOBACCO RINGSPOT NEPOVIRUS 1, Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT5G45260 105 / 4e-26
WRKY
ATWRKY52, SLH1, RRS1
SENSITIVE TO LOW HUMIDITY 1, RESISTANT TO RALSTONIA SOLANACEARUM 1, ARABIDOPSIS THALIANA WRKY DOMAIN PROTEIN 52, Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT5G36930 102 / 2e-25
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT4G36140 100 / 1e-24
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT4G19520 99 / 5e-24
disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G17890 95 / 2e-22
CHS3, DAR4
CHILLING SENSITIVE 3, DA1-related protein 4 (.1)
AT3G51560 94 / 2e-22
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G45230 93 / 7e-22
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT4G16890 92 / 9e-22
BAL, SNC1
SUPPRESSOR OF NPR1-1, CONSTITUTIVE 1, BALL, disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT4G36150 87 / 7e-20
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013729
321 / 4e-107
AT5G36930 310 / 2e-92
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10000121
184 / 3e-56
AT5G17680 277 / 1e-82
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10026201
184 / 7e-54
AT5G17680 511 / 1e-160
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10005588
182 / 6e-53
AT5G17680 499 / 2e-156
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10041078
143 / 1e-39
AT5G17680 399 / 6e-126
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10030345
139 / 4e-38
AT5G17680 528 / 5e-168
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10029722
127 / 6e-34
AT5G17680 608 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10014829
122 / 3e-32
AT5G17680 441 / 1e-136
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10018021
108 / 2e-27
AT4G12010 434 / 5e-131
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G069500
127 / 5e-34
AT5G17680 711 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G070700
127 / 8e-34
AT5G17680 722 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.T002300
119 / 3e-31
AT5G36930 635 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.001G066500
119 / 3e-31
AT5G36930 462 / 2e-146
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G068300
119 / 6e-31
AT5G17680 580 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G070001
119 / 6e-31
AT5G17680 481 / 5e-148
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G070565
118 / 7e-31
AT5G17680 482 / 3e-148
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G021681
118 / 9e-31
AT5G36930 641 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T001500
118 / 1e-30
AT5G36930 641 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.007G143300
117 / 1e-30
AT5G36930 519 / 2e-162
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Lus10039211 pacid=23150619 polypeptide=Lus10039211 locus=Lus10039211.g ID=Lus10039211.BGIv1.0 annot-version=v1.0
ATGGTGACTAGTAAGGATCACCAGGTTCTGAAGATAGTATGCGACGAAATCTACGAAGTCAAGGGACTGGACGGTGACGAATCTCTAGAGCTCTTCGTTT
TGAGATGGTTTGGCAAATGGAAATCCGGTCGTGAAAATGCACAAGATTCAATTGTTGTAAACGATGATGATCGTCCGGAGGAGTACTACATCGAGCTGTC
GAAGAAGGTTGTGAACTACACGGGAGGGAATCCATTAGCGATTATTATTTTCGGGAAGGCTCTGAAAGGTAGGAGCAGAGAGTATTGGGATAGTACATTG
TTCAGACTACAGAGGATACCGAATGAAATGATACAGAGGATGTTGAAAGCGAGCTTCGAAAGGCTGGACTGCGACGAGAAGAACAGGTTCCTCAACATTG
CTGTTTCTTTTAAGGGAGAGAGTAAAGAGTATGTGAAGTCGAGATCGGATGATGCTTATGGGTATGGATCGTCGGAGTACATCATAACTACCCTTGCTGA
CAAATCCTTGGTGAGGGTTTCGCCCGATGATGATGAGAGGGTCGAGATGCACGGTTTGGTGCAGGAAATGGGGCGTGGGATTGTTAACGAGGATTGGAAC
TAG
AA sequence
>Lus10039211 pacid=23150619 polypeptide=Lus10039211 locus=Lus10039211.g ID=Lus10039211.BGIv1.0 annot-version=v1.0
MVTSKDHQVLKIVCDEIYEVKGLDGDESLELFVLRWFGKWKSGRENAQDSIVVNDDDRPEEYYIELSKKVVNYTGGNPLAIIIFGKALKGRSREYWDSTL
FRLQRIPNEMIQRMLKASFERLDCDEKNRFLNIAVSFKGESKEYVKSRSDDAYGYGSSEYIITTLADKSLVRVSPDDDERVEMHGLVQEMGRGIVNEDWN
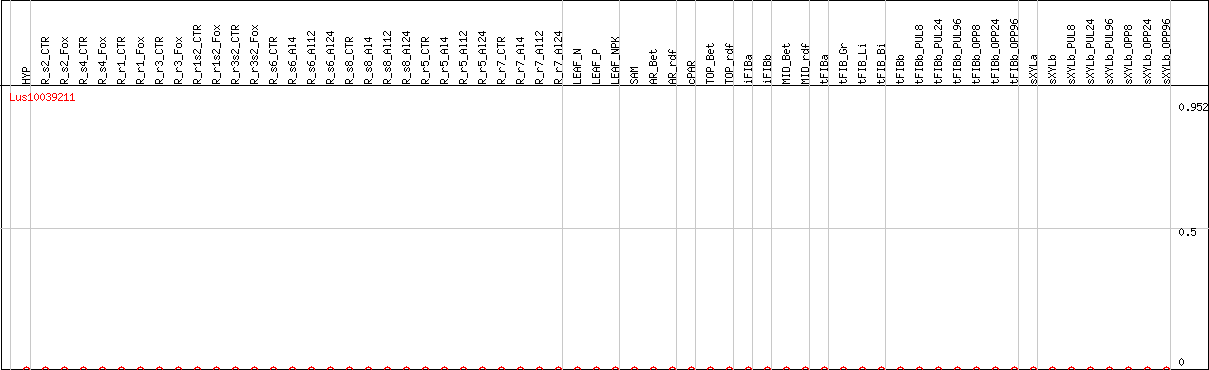
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039211 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.