Lus10039850 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10039850 pacid=23165029 polypeptide=Lus10039850 locus=Lus10039850.g ID=Lus10039850.BGIv1.0 annot-version=v1.0
ATGGCTTCCAACGACGACGGAGACGACGACGGCGATATTTCTCCCGCTGTGGAGCCGTGTCGCGCGCGGTGGAACTACGATGTCTTCCTCAGCTTCAGAG
GAGCCGACACGAGAACTGGGTTCACCGGCCACCTCTACTCGGCGCTCCTTCGCGACGGGATCCACACTTTCCTCGACGACAAGGAGATGGAGACCGGCGG
AGAGGTGGGCCCGGAGTGCCTGAGAGGGATCGAAGAGTCGAGATTCTCCTTCGTCGTCCTGTCCCAACACTACGCTTCCTCGATCTGGTGCTTGGAGGAG
CTCGTTCACATCCTCAAATGCAGGAGGGAAAAGGGTCACGGCGTCTGGCCGGTGTTCTACGGCGTAGATCCGTCGGACGTCGAGGAGTGCAAAGGGAGTT
TCGCCGATGCGTTCGCCAAGCACGAGGCCGATTTCAAATCGGAGATTGAGACAGGGAAGAGGACGATCGGAGACTGGGAAGGGTTCGCGGCTCGGATTGA
AGAGTGGAGATCTGCGCTACGTGAAGTCTCTTATCTGAAAGGATTGGATGTACGGAACCACGCTGATGGCAACGAAGCAACGAATATCGAGCACCTGGTG
AAGGAGATCTCCAAGAGACTTGACAGAAAAGTTCTGAGCGTTGCTAGAAATCCTGTAGGGGTTCGTCCTCGAGCACAGCCGGTGATTTCTTTCCTAGGTT
TCGACTCTCCAGGCGTCCGTGTTGTTGGGATACATGGCATGGGGGGAGTTGGCAAGACTACAGTGGCGAAGGAGGTTTACAACTCGGTCTTCAATAATTT
CGAAGGCTCTTGTTTCCTGTCAAATGTCAGGGAACAATTGGCGGTCAAAGGTGTACCATACTTGCAAAGGCAACTCCTTTCCGAGATCCTGAAGAGGAGA
TACGAGAAGATTGACAATGGTGATCGAGGGTTGAATCTCATACGAGACAGATTATGCCACAAACAGCTTCTTCTCGTCCTGGACGATGTTGACGAAATGA
GTCAAGTGAACAAAGTGATTGGAAGCCTTGCTTGGCTATCCGAAGGAAGCCGAGTGATCATCACTACTAGAACTAAGAATCTCCTGCAACCATCTGAATT
CTTTTGGCAGTATGAGGTTCAGGAATTAGACGAGAGAGATTCTCTCGAGCTCTTGAGTTTGCACGCCTTTGATAGGCGTGATCCACCGGAACACTACGAA
AATAGTGCAGGACAGATGCTGCAATATACCAGGGGAGTTCCGCTAGCTCTTGAAGTTTTGGGTAGTTCGTTACGTGGCCAAACTGTCGAAATATGGAATA
GCAGAATGGAGAAGCTTAAGCTGATGGCTAATGAAGATATTCAGCCTGTGTTAAAGACGAGCTACGATTCGCTTGACGATACTGAGAAGTTCATACTCCT
CGACATTGCTTGCTTCTTTGTTGGATATGATAAAGACTATGTGATGAGCATACTTGAAGGTTGTGGTTTCTTTCCAACTGATGGAATCAATACTCTGATG
AGGCGGAGCCTTTTGAAAGTCGATTATCGGAACAAGCTATGGATGCATGATTTGCTTCGGGGCATGGGGAGAGAAATCGTTCGTCAAGAGAGTCCTGTAG
ATCCCGGGGAGCGTAGCAGGTTATGGCGTGATGAAGATGTTTCAAACGTGCTAACTAGTAAAACAGGGACAAAAGTCATTGAAGGGCTGGCAGTGAACTC
ACTGGCGATTGAGTTTTCTGCGACTGCAAAGGCATTCAAGAAGATGAAAAGGCTGAGACTGCTAAAGCTTAATCATGTACGTGTCAAGGGAAGCTATGAG
TACATTTCTTGTAAGTTGAGGTGGCTGTGCTGGCATGGATTCTCATTGGAATCTATCCCGTCTGATTTTGAATTGGAGAATCTGATTGCTCTTGATATGC
GTCACAGCAGCCTGGAAATTTTCTCAGAGGAAGACGAGAAGTCCATGAAGAAGCTGAAATTCCTTGACCTGAGCCACTCTCTTAAGCTCATCAGGACTCC
TAATTTTGAAGACATCCCCAGGATCGAGAAACTGAAGCTCAAGGGCTGCATTAGCTTGGAAGAAGTTGATGACTCCATCAGATATCTTATTCATCTCCGT
TCTCTCAATTTGCAAGGATGCCACAAGCTTAAAACTCTGCCAGACAGCATTAGCGCCCTGGCCGGATCAATTGTAAACCTGAACATGTCTGGTTGCTCAG
CATTCGAGACGTTACCAGTGTCCGTTGGCTCGCTTGATCATCTTGCTCTCTTGAATTTGAAAGAATGCAGGAATCTTAAATGCCTCCCATCAAGCATCGG
CGGTTTGAAATCTCTCACGGACTTGGACTTGTCAGGCTGCTCGGAACTCGAAGAACTGCCTGAATCGGTCGGACTCTTGGGTCATCTCGAATCCTTAAAC
ATGTCTGGTTGTTCCAAACTGGAGCTACTTCCGGATTCCATTGGTTCCCTCACTCGCATACCGATTTTGAATTTGAGAGGCTGCACAAGTCTCAAGTATC
TCCCTCAAAGCATTGGTGCTCTAACGTCATTGAAGAAGATCAACCTGTCAGGATGCTCAAGTCTCGTCGAGTTGCCTGAGGGAATTGGAGCTTTAGAGCC
TTTACTTGTACTTATGGCCAACGAAACTGCATTGACTACTCTCCCGGAGACAACTTCAAACCTCAAGAAGCTTAAAGTATTATCTCTACGCGGTTGTGAC
CGGTTGTTCTTCGCTCCGAAGAACAACCCTTTGGTGCTCAACTTCCTTCCTGCTTCCTTGAGCAGGTTGGATCTTAGACACTGCAACATAACGGATGAGA
TAATGCCTGGAGATCTAGGAAGCTTGCGTCTCCTGAACATCTTGAAGTTATGTGGGAACAACTTTACTTGCTTGCCAGAGAGCATCAATTCCTTACCGGA
ACTTGCTGAGCTTTGGCTCAACAAGTGCAGGACACTTAGAAGAATACCGGAGCTTCATTCGGGTGTTAAAGTGTTGCAGGCGAATGACTGCCCGTCGCTG
GAGACGATCAACTTGAGAAACTACCATGGAGCTGCGGGCTTGGAAGTTAAGGGGTGTATGAACTTGAGAGAAATCGAAGGGTTTTACAGCTTGGAAACAC
TTGAACCTGAAACATTTGAGGTTCTTGAAAGTTGTGGGTTCTTCTTCTCGCAGGGTTCGGCATCAAACGTTCAAACTTACAAGGTTGACAGCCTTACTGA
TACGTACAGCGTTTGCCCCCTTCAGGCTCTGGCAGAAAGGGGGATATACAGCCTGTTCATCCCATCAACCAAAATTCCAACGTGGTTCACCCATCAAACA
GACGGCGACACCATATCCTTTCGTGTACCTCAACTCCCTAAAGGCTCCACCATCACCGGGATCGCTACTTGCGCCATTTACGCGTGGGATCGCCCCAGGG
AGTCGTGCTTCTTCATGCCGTACATTCGAATTGTCAACAGAACAAAGTTGTTCAACTGGCTGTATACTCCACACATCACCTTCTTCGTGAACGATGGCGA
CGAGAGCCAGAGAGAGACCATGTGGCTCTGCCACTGGACTTTCGACCCGAATGCAGTACACGTGGACGACATCGATGCTGGCTGGAGGCTTATGGGCGAG
ACCGAACAAGGCGACGAGGTTGAGTTCTGTGTTGACATGGGGTATGGGATTCGAGTCAAGAAATGCGGCGTCCATCCACTTTATAGTAACAACTTTGGTG
ATGGTGATGGTTTGGTGATGAGACCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10039850 pacid=23165029 polypeptide=Lus10039850 locus=Lus10039850.g ID=Lus10039850.BGIv1.0 annot-version=v1.0
MASNDDGDDDGDISPAVEPCRARWNYDVFLSFRGADTRTGFTGHLYSALLRDGIHTFLDDKEMETGGEVGPECLRGIEESRFSFVVLSQHYASSIWCLEE
LVHILKCRREKGHGVWPVFYGVDPSDVEECKGSFADAFAKHEADFKSEIETGKRTIGDWEGFAARIEEWRSALREVSYLKGLDVRNHADGNEATNIEHLV
KEISKRLDRKVLSVARNPVGVRPRAQPVISFLGFDSPGVRVVGIHGMGGVGKTTVAKEVYNSVFNNFEGSCFLSNVREQLAVKGVPYLQRQLLSEILKRR
YEKIDNGDRGLNLIRDRLCHKQLLLVLDDVDEMSQVNKVIGSLAWLSEGSRVIITTRTKNLLQPSEFFWQYEVQELDERDSLELLSLHAFDRRDPPEHYE
NSAGQMLQYTRGVPLALEVLGSSLRGQTVEIWNSRMEKLKLMANEDIQPVLKTSYDSLDDTEKFILLDIACFFVGYDKDYVMSILEGCGFFPTDGINTLM
RRSLLKVDYRNKLWMHDLLRGMGREIVRQESPVDPGERSRLWRDEDVSNVLTSKTGTKVIEGLAVNSLAIEFSATAKAFKKMKRLRLLKLNHVRVKGSYE
YISCKLRWLCWHGFSLESIPSDFELENLIALDMRHSSLEIFSEEDEKSMKKLKFLDLSHSLKLIRTPNFEDIPRIEKLKLKGCISLEEVDDSIRYLIHLR
SLNLQGCHKLKTLPDSISALAGSIVNLNMSGCSAFETLPVSVGSLDHLALLNLKECRNLKCLPSSIGGLKSLTDLDLSGCSELEELPESVGLLGHLESLN
MSGCSKLELLPDSIGSLTRIPILNLRGCTSLKYLPQSIGALTSLKKINLSGCSSLVELPEGIGALEPLLVLMANETALTTLPETTSNLKKLKVLSLRGCD
RLFFAPKNNPLVLNFLPASLSRLDLRHCNITDEIMPGDLGSLRLLNILKLCGNNFTCLPESINSLPELAELWLNKCRTLRRIPELHSGVKVLQANDCPSL
ETINLRNYHGAAGLEVKGCMNLREIEGFYSLETLEPETFEVLESCGFFFSQGSASNVQTYKVDSLTDTYSVCPLQALAERGIYSLFIPSTKIPTWFTHQT
DGDTISFRVPQLPKGSTITGIATCAIYAWDRPRESCFFMPYIRIVNRTKLFNWLYTPHITFFVNDGDESQRETMWLCHWTFDPNAVHVDDIDAGWRLMGE
TEQGDEVEFCVDMGYGIRVKKCGVHPLYSNNFGDGDGLVMRP
|
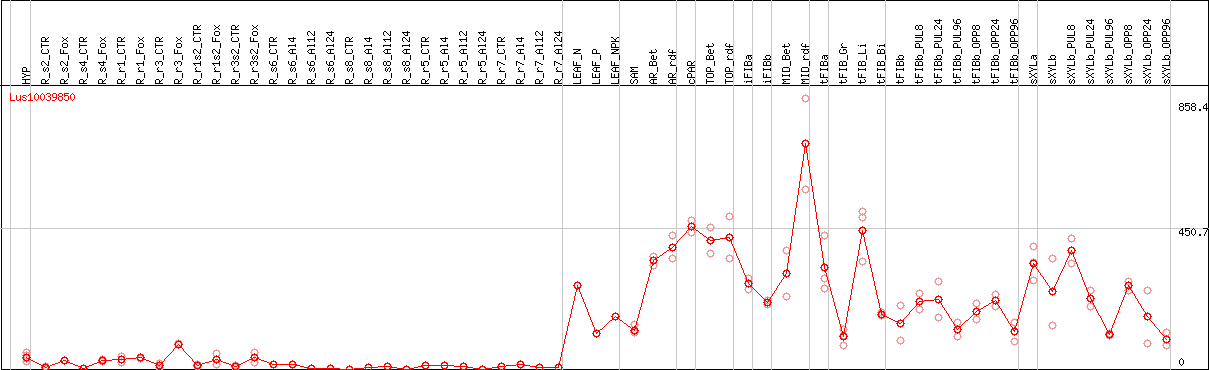
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039850 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
