External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G06060 337 / 3e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29350 323 / 1e-111
SAG13
senescence-associated gene 13 (.1.2.3)
AT1G07440 320 / 2e-110
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G29150 319 / 3e-110
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29340 312 / 1e-107
NAD-dependent epimerase/dehydratase family protein (.1.2.3)
AT2G29360 308 / 5e-106
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29290 306 / 2e-105
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G29330 303 / 4e-104
TRI
tropinone reductase (.1)
AT1G07450 303 / 6e-104
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29310 296 / 2e-101
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016497
489 / 2e-177
AT5G06060 345 / 1e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016498
481 / 4e-174
AT5G06060 351 / 7e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040735
448 / 4e-161
AT5G06060 341 / 4e-119
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016499
390 / 3e-138
AT5G06060 337 / 2e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040732
369 / 6e-130
AT5G06060 269 / 7e-91
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040731
358 / 1e-125
AT5G06060 308 / 5e-106
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040736
343 / 2e-119
AT5G06060 382 / 3e-135
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10040271
332 / 5e-115
AT5G06060 388 / 2e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10004704
322 / 2e-111
AT5G06060 387 / 3e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G142600
330 / 9e-115
AT2G29290 354 / 3e-124
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.013G026000
326 / 3e-113
AT2G29290 362 / 2e-127
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.008G059100
322 / 3e-111
AT5G06060 416 / 8e-149
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G039500
311 / 3e-107
AT2G29150 351 / 5e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G039300
311 / 4e-107
AT2G29150 356 / 7e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G245000
305 / 7e-105
AT2G29150 343 / 9e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G244900
307 / 2e-104
AT2G29260 387 / 4e-135
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G089800
284 / 2e-96
AT2G29150 307 / 2e-105
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G089700
276 / 3e-93
AT5G06060 320 / 6e-111
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.013G026100
274 / 5e-93
AT2G29150 284 / 5e-97
NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10040733 pacid=23157598 polypeptide=Lus10040733 locus=Lus10040733.g ID=Lus10040733.BGIv1.0 annot-version=v1.0
ATGGAGGCTGTAGTAGAAAGTAGCAGTTTCAAGAGCAAATTGAGTAGCAGATGGTCTCTCCATGGCAAGACTGCACTTGTCACTGGCGGCACTAAAGGAC
TTGGGTATGCAATCGTTGAAGAATTGGCAGGACTAGGGGCTATTGTGCACACATGCTCCAGAAACCAGGAAGAAATTGGCAAGTGTTTGAATGAATGGAA
GGAGAAGGGTCTTGAGGTAACTGGTTCTGTCTGTGATTTGACCTCTGCAGCAGAAAGAAAGAATCTAATGGAGACCGTGTCATGTCAGTTCAACGGAAAG
CTAAACATCCTGATTAATAATGCAGGGACTAACATATATAAGCCGACACTGGAGTTCACTGACGAAGATTACTCGTACCTGATGCGAACCAATCTCGAAT
CGGCCTATAGTCTGAGTCTGCTTGCTCACCCTCTGCTGAAAACCTCAGGGGAAGGCAACATAGTCTTCATGTCGTCTGCTGCTGGTGTGATTTCCATGAA
TGTTGGATCCATATATGGGATGACCAAAGCTGCTATGGTTCAACTGACAAAGGACCTAGCCTGCGAGTGGGCAAAAGACAAAATAAGGATCAACTGTGTT
GCACCCTGGTTCATCATAACTCCTATCAATGCAGAGTTATTCGAGAGTGAAAAGTTCACAAGTGCAGTGATAGCTCAAACCCCGATGGGACGAACGGGGA
AGCCGGAGGAGGCAGCTGGGCTGGTGGCATTCCTCTGCCTTCCCGCTTCCTCTTATGTCACTGGCCAGACCGTAGCCGTCGACGGAGGAGCTACTGTGAA
TTGCTTCTCCCCTCCATGA
AA sequence
>Lus10040733 pacid=23157598 polypeptide=Lus10040733 locus=Lus10040733.g ID=Lus10040733.BGIv1.0 annot-version=v1.0
MEAVVESSSFKSKLSSRWSLHGKTALVTGGTKGLGYAIVEELAGLGAIVHTCSRNQEEIGKCLNEWKEKGLEVTGSVCDLTSAAERKNLMETVSCQFNGK
LNILINNAGTNIYKPTLEFTDEDYSYLMRTNLESAYSLSLLAHPLLKTSGEGNIVFMSSAAGVISMNVGSIYGMTKAAMVQLTKDLACEWAKDKIRINCV
APWFIITPINAELFESEKFTSAVIAQTPMGRTGKPEEAAGLVAFLCLPASSYVTGQTVAVDGGATVNCFSPP
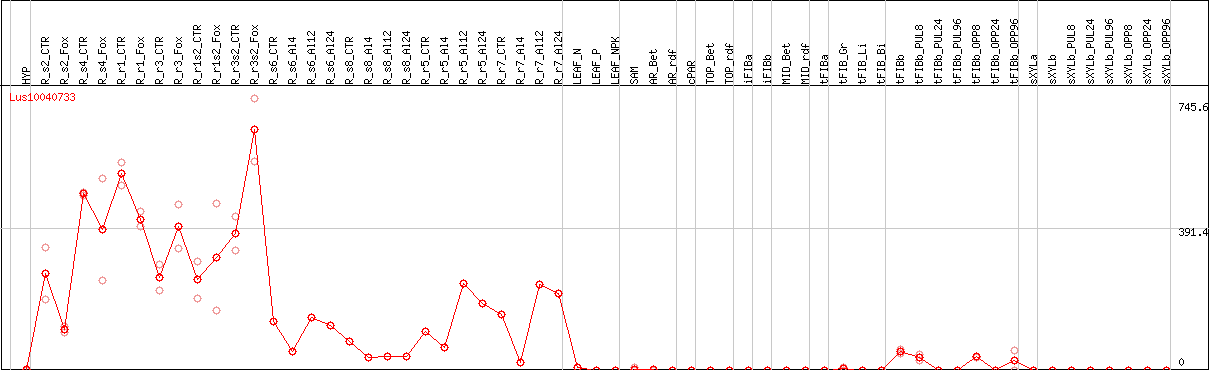
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040733 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.