External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G03340 280 / 4e-90
ATPase, AAA-type, CDC48 protein (.1)
AT3G09840 275 / 5e-88
ATCDC48, CDC48A, CDC48
cell division cycle 48 (.1)
AT3G53230 265 / 4e-84
ATPase, AAA-type, CDC48 protein (.1)
AT3G01610 106 / 1e-26
CDC48C, EMB1354
embryo defective 1354, cell division cycle 48C (.1)
AT3G56690 91 / 4e-21
CIP111
Cam interacting protein 111 (.1)
AT5G53170 84 / 5e-19
FTSH11
FTSH protease 11 (.1)
AT1G45000 82 / 3e-18
AAA-type ATPase family protein (.1.2)
AT5G43010 81 / 5e-18
RPT4A
regulatory particle triple-A ATPase 4A (.1)
AT3G16290 81 / 8e-18
EMB2083
embryo defective 2083, AAA-type ATPase family protein (.1)
AT2G03670 79 / 3e-17
CDC48B
cell division cycle 48B (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037385
335 / 4e-111
AT5G03340 1501 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Lus10021442
282 / 9e-91
AT5G03340 1520 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Lus10016123
272 / 5e-87
AT5G03340 1526 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Lus10023018
268 / 9e-86
AT5G03340 1419 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Lus10021441
266 / 1e-84
AT5G03340 1529 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Lus10016124
266 / 1e-84
AT5G03340 1511 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Lus10001403
103 / 6e-29
AT3G09840 114 / 6e-31
cell division cycle 48 (.1)
Lus10001402
96 / 4e-23
AT3G53230 803 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Lus10020255
90 / 7e-21
AT3G56690 1066 / 0.0
Cam interacting protein 111 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G125500
281 / 1e-90
AT5G03340 1496 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Potri.016G091600
281 / 2e-90
AT3G53230 1484 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Potri.015G080600
279 / 1e-89
AT5G03340 1455 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Potri.012G088200
270 / 5e-86
AT5G03340 1459 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Potri.017G108200
241 / 2e-75
AT5G03340 1311 / 0.0
ATPase, AAA-type, CDC48 protein (.1)
Potri.006G035000
92 / 8e-22
AT3G56690 1080 / 0.0
Cam interacting protein 111 (.1)
Potri.013G151600
89 / 2e-20
AT3G01610 747 / 0.0
embryo defective 1354, cell division cycle 48C (.1)
Potri.001G128700
87 / 6e-20
AT2G03670 766 / 0.0
cell division cycle 48B (.1)
Potri.012G000700
83 / 1e-18
AT5G53170 1076 / 0.0
FTSH protease 11 (.1)
Potri.003G050600
83 / 2e-18
AT3G16290 1159 / 0.0
embryo defective 2083, AAA-type ATPase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00004
AAA
ATPase family associated with various cellular activities (AAA)
Representative CDS sequence
>Lus10041332 pacid=23179580 polypeptide=Lus10041332 locus=Lus10041332.g ID=Lus10041332.BGIv1.0 annot-version=v1.0
ATGGACGGCATGTCAGCAAAGAAGACCGATTTCATAATCGGAACCACAAACAGGCCCGACATCATCGATCCGGCACTCCTGAGGCCCGGCCGTCTTGACC
AGTTGATTTATATCCCTCTTCCGGATGAGGATTCGCGCCACAACATCTTCAAAGCCTGGTTAAGGAAATCGCCCGTGGCAAAGGATGTTGACTTGAGAGC
ATTGGCTAAATACACGCAAGGGTTCAGTGGCGCTGATATCACTGAGATATGCCAGCGGCCTTGCAAGTATGCCATAAGAGAGAACATAGAGACGGATATT
GAAAGGGAAAGGAGGAGGAGGGACACCCCGGAAGCGATGCAGGAGGACATGGACGACGAGGTTGCAGAGATCAAAGCTGCACATTTCGAAGAGTCGATGA
AGTATGCTCGAAGGAGTGTGAGTGACGCAGATATCCGCAAGTACCAGACGTTTGCTCAGACATTGCAGCAATCTAGGGGTGTAGGATCAGAGTTCAGGTT
TTTGGAATCTCGAACCGGGATGGCTGCTGCGGCCGATCCGTTCTCAACTTCGGCTGGTGGAGCTGATGAAGATGATTTGCATAGTTAG
AA sequence
>Lus10041332 pacid=23179580 polypeptide=Lus10041332 locus=Lus10041332.g ID=Lus10041332.BGIv1.0 annot-version=v1.0
MDGMSAKKTDFIIGTTNRPDIIDPALLRPGRLDQLIYIPLPDEDSRHNIFKAWLRKSPVAKDVDLRALAKYTQGFSGADITEICQRPCKYAIRENIETDI
ERERRRRDTPEAMQEDMDDEVAEIKAAHFEESMKYARRSVSDADIRKYQTFAQTLQQSRGVGSEFRFLESRTGMAAAADPFSTSAGGADEDDLHS
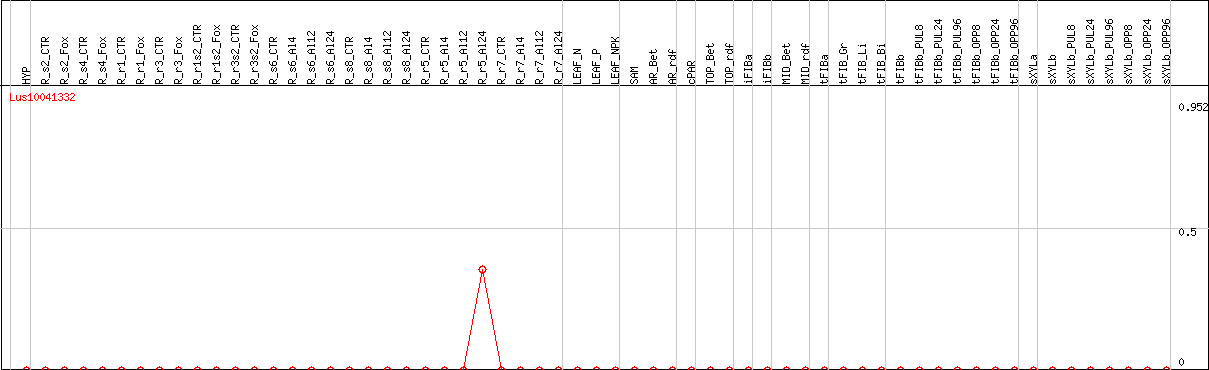
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041332 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.