Lus10043013 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10043013 pacid=23167996 polypeptide=Lus10043013 locus=Lus10043013.g ID=Lus10043013.BGIv1.0 annot-version=v1.0
ATGAAACTTGTACTTGCTTCCTCTCCTCTTTCTCTCCATTCCCGTTCCCCTCCTTTGACTTCCTTCCACCACCATCCTTCTTCCCCTTCTCAATCCCTTC
CTTCTTCTTCCTCTGCTCGTTGTTTCGGCGTTTCCATTTCTCATCATAATCACAACCACCTCTCTTCCAGAATCAGTTGCAGCCGTTCCAGAAAGAAAAG
CCGTCGTACAGTCTTCACCGTTTCCGCCGTCTTCGAGAGGTTCACCGAGCGAGCTATCAAAAGTGTGATTTTCTCTCAGAGAGAAGCTCGAGCCTTGGGG
AAGGATACTGTTGCCCCTCAGCACCTCTTGCTCGGTCTAATCGTTGAGGATCGTGACCTTGATGGATTCCTTGGGTCTGGAATCACTATCGACCGTGGTA
GACAAGTTGTTAGGGAGATTTGGGTTAACGATGGTGATGATGCTGCGGATCATCATCCGACTTCTTCTTCTTCTACTTCTACTACGGATATGCCTTTCTC
TGCTGCCACCAAACGTGTTTTCGAGGCTGCGGTTGAGTATTCCAGGAATATGAAGCATAAGTTCATTTCCCCTGAGCATATTGCTATTGGCTTGTTCACC
GTTGATGATGGAAGTGCAGCTCAAGTTTGTCAAAGATTAGGGGTAAATTCAAATCACTTGGCAGCTGTGGCAATCACCAGATTGCAAGGAGAGCTAGCCA
AGGATGGAAGAGGAGAACCATCTCTGTCCAAAGGGACAAGTGAGAAAGTTCTATCTAAAAGAGCTGTCGCTTTAAGATCCGCTGAAAAAGCTAAGGAGAA
AAGTGCCCTGGCCCAATTCTGTGTAGATATTACCTCTCGAGCTGTTGAAGGGCGCATCGATCCAGTTATCGGACGAGAGAGTGAGATTCAAAGAGTTATT
CAGATACTTTGCCGCAGAACCAAGAACAATCCGATTCTTCTTGGTGAAAGTGGAGTTGGGAAAACGGCCATTGCTGAAGGACTAGGAATGAAAATTGCTC
AGGCTGAAGTGCCTGCTTCCCTCTTAGGAAAACGTGTCATCTCTTTGGATATTGGACTGTTAATTGCTGGTGCAAAGGAAAGGGGAGAACTTGAGTCACG
TGTTACTTCATTAATCAAAGAGATACGAGAAGAAGGGAATGTAGTTCTCTTCATTGATGAAGTCCATACTCTTGTTGGGTCGGGTACAGTGGGACGAAAA
AGCAAGGGAAATAGCCTTGACATTGCTAATCTACTGAAGCCAGCTCTTGGAAGGGGTGAATTACAGTGTATTGCATCCACAACTATGGATGAACACAGGT
CACATATTGAAAGTGATAAAGCATTGGCAAGAAGATTTCAGCCTGTCATGATTAACGAACCGAGCCAGGAGGATGCTGTCAGGATATTGCTGGGTTTACG
TCAAAAGTACGAGACCCATCATAACTGCAGATATACATTGGAAGCAATAAATGCAGCAGTTCATTTGTCAGCAAGATATATTGCTGATAGATTTCTGCCA
GATAAAGCTATTGATCTCATTGACGAGGCAGGAAGCAGAGCTCGTATTGAAGCTTACAGGAAGAAGAAAGAACAGCAAACTGGTATACTGTCGAAGTCAC
CAGATGATTATTGGCAGGAAATCAGAACTGTTCAGGCCATGCATCAAGTGGTTATTTCAAGTAAGCCAATCGCTGATGATAGTGAATCAAGCATTGATGA
TGATTCTGGAGAACTCAAATCAGAGCCGCCTTCACCTTCTACATCCGATAATGCCGAACCCATGGTGGTGGGGCTTAATGACATTGCCTCTGTGGCTTCG
CTCTGGTCCGGGATCCCTGTTCAGCAACTCACTGCTGATGAAAGAATCGCTCTCGTGGGTCTCGAGGAACAGCTTAGGAAAAGAGTCGTTGGCCAAGATG
AGGCTGTTGGTGCTATATCTAGAGCAGTTAAGAGGTCTCGTTTCGGCCTCAAAGACCCCGATAGACCCATTGCAGCCTTACTTTTCTGTGGCCCTACTGG
TGTTGGCAAAACGGAGCTAACTAAAGCACTGGCTGCATGCTACTTCGGAACCGAGTCCTCAATGCTCAGACTGGACATGAGTGAATATATGGAGCGACAC
ACGGTGAGCAAACTAATTGGAGCACCTCCGGGTTATGTCGGTCATGGCGAGGGAGGCACGCTTACCGAAGCTATCAGAAGAAAACCTTTCACGTTGGTTC
TACTTGACGAGATAGAGAAGGCCCATCCAGATATCTTCAATATCCTCCTCCAAGTATTTGAAGATGGCCACCTAACAGATTCTCAGGGCCGAAGAGTATC
ATTCAAGAACGCTTTGATAGTTATGACCTCTAACATCGGATCGTCTGCCATCGCAAAAGGAGGGCGTACTTCCATTGGGTTTGTATCTGGAGCTGATCAA
TCAATTACTTCATACGTGGGAATGAAGGAAATGGTAATGGAGGAACTCAAGTCATATTTCCGCCCTGAGTTGCTGAACAGAATAGACGAGCTAGTCGTAT
TCCGCGCACTTGAGAAGACTCAGATGATGGAGATTGTGAACATGATGCTGGAAGAAGTGAAAGGAAGACTGATATCTCTTGGGATCGGGTTGGAGGTATC
CGATTCAATCAAGGATTTGGTCTGCAAGCAAGGTTATGACCCTGCGTATGGTGCGCGGCCGCTCAGGCGAGCAGTCACCCAGGTAATAGAGAACCCATTG
AGTGAAGCTTTACTTGGTGGAGGGTTCAAACCAGGCGACACAGCTCTGATCGATGTTGATGCAACCGGGAACCCTGTCGTTATAAATCGAGAGATGGAGC
ATAGTATCCACTTAACAGACACTACACTCTCTAACTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10043013 pacid=23167996 polypeptide=Lus10043013 locus=Lus10043013.g ID=Lus10043013.BGIv1.0 annot-version=v1.0
MKLVLASSPLSLHSRSPPLTSFHHHPSSPSQSLPSSSSARCFGVSISHHNHNHLSSRISCSRSRKKSRRTVFTVSAVFERFTERAIKSVIFSQREARALG
KDTVAPQHLLLGLIVEDRDLDGFLGSGITIDRGRQVVREIWVNDGDDAADHHPTSSSSTSTTDMPFSAATKRVFEAAVEYSRNMKHKFISPEHIAIGLFT
VDDGSAAQVCQRLGVNSNHLAAVAITRLQGELAKDGRGEPSLSKGTSEKVLSKRAVALRSAEKAKEKSALAQFCVDITSRAVEGRIDPVIGRESEIQRVI
QILCRRTKNNPILLGESGVGKTAIAEGLGMKIAQAEVPASLLGKRVISLDIGLLIAGAKERGELESRVTSLIKEIREEGNVVLFIDEVHTLVGSGTVGRK
SKGNSLDIANLLKPALGRGELQCIASTTMDEHRSHIESDKALARRFQPVMINEPSQEDAVRILLGLRQKYETHHNCRYTLEAINAAVHLSARYIADRFLP
DKAIDLIDEAGSRARIEAYRKKKEQQTGILSKSPDDYWQEIRTVQAMHQVVISSKPIADDSESSIDDDSGELKSEPPSPSTSDNAEPMVVGLNDIASVAS
LWSGIPVQQLTADERIALVGLEEQLRKRVVGQDEAVGAISRAVKRSRFGLKDPDRPIAALLFCGPTGVGKTELTKALAACYFGTESSMLRLDMSEYMERH
TVSKLIGAPPGYVGHGEGGTLTEAIRRKPFTLVLLDEIEKAHPDIFNILLQVFEDGHLTDSQGRRVSFKNALIVMTSNIGSSAIAKGGRTSIGFVSGADQ
SITSYVGMKEMVMEELKSYFRPELLNRIDELVVFRALEKTQMMEIVNMMLEEVKGRLISLGIGLEVSDSIKDLVCKQGYDPAYGARPLRRAVTQVIENPL
SEALLGGGFKPGDTALIDVDATGNPVVINREMEHSIHLTDTTLSN
|
DESeq2's median of ratios [FLAX]
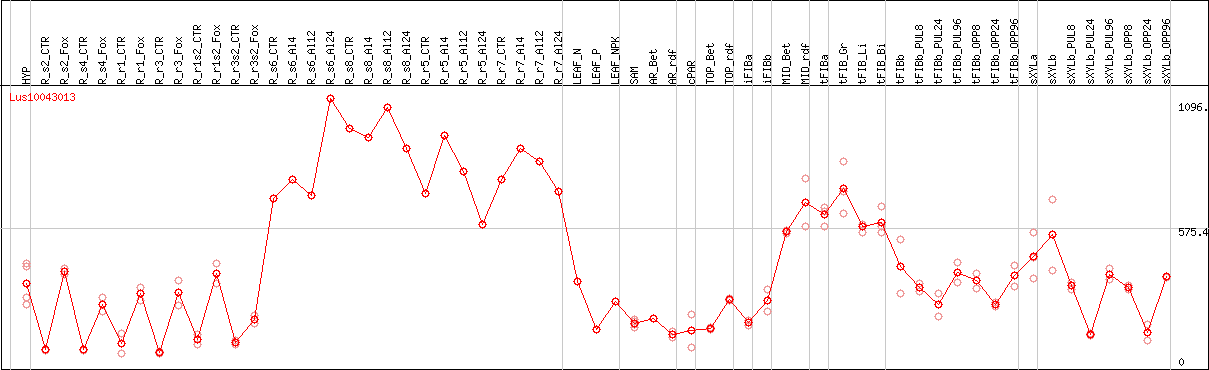
Coexpressed genes
Lus10043013 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
