External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G09770 416 / 4e-141
MYB
ATCDC5, AtMYBCDC5
ARABIDOPSIS THALIANA MYB DOMAIN CELL DIVISION CYCLE 5, ARABIDOPSIS THALIANA CELL DIVISION CYCLE 5, cell division cycle 5 (.1)
AT5G02320 74 / 7e-15
MYB
MYB3R-5, ATMYB3R5
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3R5, myb domain protein 3r-5 (.1.2)
AT5G11510 73 / 2e-14
MYB
ATMYB3R-4, AtMYB3R4
myb domain protein 3R4, myb domain protein 3r-4 (.1.2)
AT3G18100 72 / 3e-14
MYB
ATMYB4R1
myb domain protein 4r1 (.1.2.3)
AT4G33450 69 / 7e-14
MYB
ATMYB69
myb domain protein 69 (.1)
AT5G40360 69 / 2e-13
MYB
ATMYB115
myb domain protein 115 (.1)
AT1G17950 68 / 2e-13
MYB
AtMYB52, BW52, MYB52
myb domain protein 52 (.1)
AT3G27920 67 / 3e-13
MYB
GL1, ATMYB0, AtGL1
GLABRA 1, myb domain protein 0 (.1)
AT3G09370 69 / 4e-13
MYB
ATMYB3R-3, ATMYB3R3
myb domain protein 3R3, myb domain protein 3r-3 (.1.2)
AT2G02820 67 / 9e-13
MYB
ATMYB88
myb domain protein 88 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034133
451 / 2e-153
AT1G09770 1207 / 0.0
ARABIDOPSIS THALIANA MYB DOMAIN CELL DIVISION CYCLE 5, ARABIDOPSIS THALIANA CELL DIVISION CYCLE 5, cell division cycle 5 (.1)
Lus10031900
72 / 3e-14
AT1G17950 220 / 2e-70
myb domain protein 52 (.1)
Lus10031326
72 / 3e-14
AT1G17950 219 / 3e-70
myb domain protein 52 (.1)
Lus10024394
71 / 1e-13
AT3G09370 456 / 8e-157
myb domain protein 3R3, myb domain protein 3r-3 (.1.2)
Lus10025351
71 / 1e-13
AT3G09370 456 / 8e-157
myb domain protein 3R3, myb domain protein 3r-3 (.1.2)
Lus10022256
69 / 2e-13
AT3G27785 175 / 2e-50
PLANT GROWTH ACTIVATOR 37, myb domain protein 118 (.1)
Lus10011687
69 / 3e-13
AT4G32730 502 / 3e-158
C-MYB-LIKE TRANSCRIPTION FACTOR 3R-1, ARABIDOOSIS THALIANA MYB DOMAIN PROTEIN 3R1, Homeodomain-like protein (.1.2)
Lus10038623
69 / 4e-13
AT4G32730 376 / 3e-118
C-MYB-LIKE TRANSCRIPTION FACTOR 3R-1, ARABIDOOSIS THALIANA MYB DOMAIN PROTEIN 3R1, Homeodomain-like protein (.1.2)
Lus10022136
69 / 5e-13
AT4G32730 682 / 0.0
C-MYB-LIKE TRANSCRIPTION FACTOR 3R-1, ARABIDOOSIS THALIANA MYB DOMAIN PROTEIN 3R1, Homeodomain-like protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10043451 pacid=23168344 polypeptide=Lus10043451 locus=Lus10043451.g ID=Lus10043451.BGIv1.0 annot-version=v1.0
ATGAGGATTATGATAAAAGGAGGTGTTTGGAAAAACACCGAGGATGAGATCCTTAAGGCCGCGGTTATGAAGTACGGCAAGAACCAGTGGGCTCGCATTT
CTTCGCTTCTCGTTCGCAAATCTGCCAAGCAGTGTAAGGCTCGCTGGTACGAGTGGCTCGACCCTTCAATCAAAAAGACTGAGTGGACCAGAGAAGAGGA
TGAGAAATTACTGCATCTTGCTAAGCTCATGCCTACACAATGGAGGACGATTGCACCGATTGTTGGGCGCACTCCTTCCCAGTGTCTTGAACGGTATGAG
AAGCTCCTTGATGCAGCTTGTGTGAAGGATGATAACTATGAACCTAGCGACGACCCTCGCAAGCTGCGGCCAGGAGAAATTGATCCTAATCCTGAATCCA
AGCCAGCGCGTCCTGATCCTGTTGATATGGATGAGGATGAGAAGGAAATGCTTTCTGAAGCACGAGCTAGGTTAGCTAACACTAGAGGTAAGAAGGCGAA
AAGGAAAGCTAGAGAAAAACAGCTTGAAGAGGCTAGGAGGCTTGCTTCCTTACAGAAGAGAAGAGAGCTTAAAGCTGCTGGTATTGATGCGAGGCAGAGG
AGAAGGAAAAGGAAAGGGATTGACTATAATTCAGAAATTCCCTTTGAAAAGAGGCCTCCTCCTGGCTTTTATGATGTTGCTGATGAAGATAGAACGGTAG
AACAGCCAACGTTCGCAACCAACATAAAAAAGGAAGAGGTCAAAGTTGATGCTTCCTGCACCACAGATTTCTAA
AA sequence
>Lus10043451 pacid=23168344 polypeptide=Lus10043451 locus=Lus10043451.g ID=Lus10043451.BGIv1.0 annot-version=v1.0
MRIMIKGGVWKNTEDEILKAAVMKYGKNQWARISSLLVRKSAKQCKARWYEWLDPSIKKTEWTREEDEKLLHLAKLMPTQWRTIAPIVGRTPSQCLERYE
KLLDAACVKDDNYEPSDDPRKLRPGEIDPNPESKPARPDPVDMDEDEKEMLSEARARLANTRGKKAKRKAREKQLEEARRLASLQKRRELKAAGIDARQR
RRKRKGIDYNSEIPFEKRPPPGFYDVADEDRTVEQPTFATNIKKEEVKVDASCTTDF
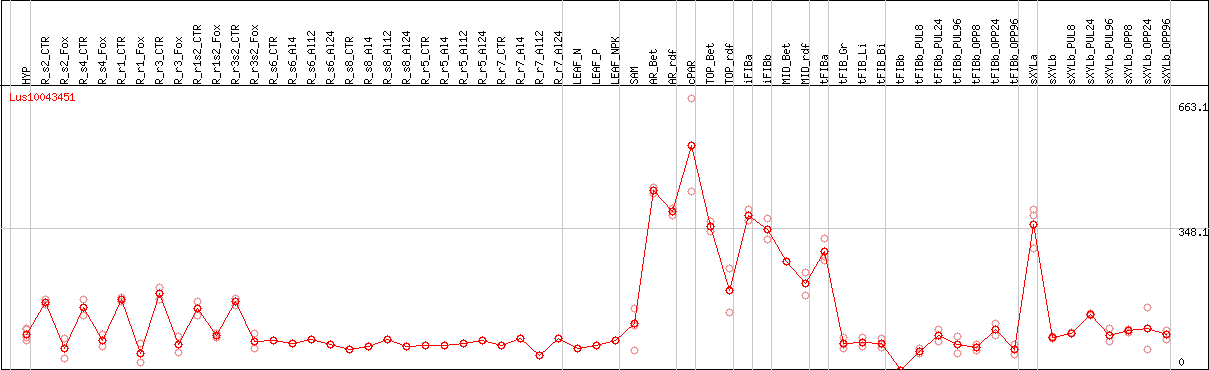
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043451 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.