PtpRR11 (Potri.001G146200) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | PtpRR11 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G146200.35 pacid=42791504 polypeptide=Potri.001G146200.35.p locus=Potri.001G146200 ID=Potri.001G146200.35.v4.1 annot-version=v4.1
ATGGTTTGCACCACTAATGATTTATCAGCATGGAAGGATTTTCCAAAGGGCCTCAGAGTCCTTCTCCTTGATGAGGACAGCATGTCTGCTGCCGAGATAA
AATCGAAACTGGAAGCAATGGACTATATTGTTTATACATTCTGCAATGAGACTGAAGCGCTCTCTGCAATTTCAAATGAACCTGGAAGCTTCCATGTTGC
TATAGTGGAGGTGAGCATGAGCAATAGCAGCAGGAGTTTCAAGTTTCTAGAAACTTCGAAGGACTTGCCAACCATTATGACTTCAAGCATTGATTGCCTG
AACACCATGATGAAGTGCATAGCGCTTGGTGCAGTTGAATTCCTTCGGAAACCACTCTCTGAAGATAAACTGAGGAATATATGGCAGCACGTCGTTCATA
AGGCATTTAATGCAGGGGGGAGTGTTCAGTCCAAATCACTGAAGCCTGTCAAAGATTCTGTAGTTTCCATGCTGGAGCTCAAAGATGGACTCGAAGAAAA
TGAGAATAAGAACATGGAGAAAACAGAAAATGTGTCTCAGGCTCAGGAAAGTAATGAGCAGCAGTCACCAGCTAGTGACAAGTATCCGGCTCCTTCAACC
CCTCAATTGAAACAAGGAGAGAGACTGCTGGATGATGGAGATTGCCAAGACCATATTAACTGTTTAATAGAAAAGGAGAGTGTGGAGCAAGAGGGAGATT
CTAAATCCGTTGAAACTACTTGTGCCATGTCTGAAGAAACTCTACAGGCAGGGGATCCTCAGAGCTTTACAGAGACTGTGATCAAAGAGGAGGATGACTC
AACTGATGGTGTCAAGAGTGAGAATAACATGTGTCCCAATTCACAAAATAAAGACACTCTCAACCATTCAAATGGTTGTGCTGAAAAAGCTTCTAGCCTT
CATAATTCCCATGGGACTAGAGCTAATAGAAAGAAGATGAAGGTAGACTGGACACCAGAGCTTCACAGAAAGTTTGTGCAGGCAGTGGAAAAACTAGGTG
TAGATCAAGCGATTCCTTCAAGAATATTAGAGGTAATGAAAGTGGAAGGCTTGACGAGGCATAATGTAGCAAGTCATCTGCAGAAATATCGAATGCACAG
GAGACATATTTTGCCCAAGGAAGATGAACGGCAGTGGACGCAGCATAGAGATCAAGTACAAAGGAGTTACTATCCACATAAACCTATCATGGCGTATCCT
CCATATCATTCTAACCATGCTCTCCCACCTGGTCCGGTCTATCCTATGTGGGGAGCAACAGGTAGTCACACAGCTGGGGTGCATATGTGGGGCCCCCCCG
GCTACTCCCCATGGCCCCCGACAGAAAGTTGTCATTGGAAGCCTTATCCAGGGATGCATGCAGATGCGTGGGGTTGCCCTGTGATGCCACCACATAATCC
TTATTCTACCATTCCTCATTATGCATCTGGGTTTCAGAGTGCCAGCATGGTCAATAACAGCTGTGGTATGCCACAAAAACTGTTTGACCTACAGCCGGCC
GAGGAGGTTATCGACAAGGTAGTGAAAGAGGCAATCAACAAGCCATGGCTACCTTTGCCCTTGGGCCTCAAGCCCCCGTCTGCCGATAGCGTCCTAGCAG
AGCTCTCCAGGCAAGGCGTCTCATCAATTCCTCCCCGCATTAATGGCTCTAATTCCTTCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G146200.35 pacid=42791504 polypeptide=Potri.001G146200.35.p locus=Potri.001G146200 ID=Potri.001G146200.35.v4.1 annot-version=v4.1
MVCTTNDLSAWKDFPKGLRVLLLDEDSMSAAEIKSKLEAMDYIVYTFCNETEALSAISNEPGSFHVAIVEVSMSNSSRSFKFLETSKDLPTIMTSSIDCL
NTMMKCIALGAVEFLRKPLSEDKLRNIWQHVVHKAFNAGGSVQSKSLKPVKDSVVSMLELKDGLEENENKNMEKTENVSQAQESNEQQSPASDKYPAPST
PQLKQGERLLDDGDCQDHINCLIEKESVEQEGDSKSVETTCAMSEETLQAGDPQSFTETVIKEEDDSTDGVKSENNMCPNSQNKDTLNHSNGCAEKASSL
HNSHGTRANRKKMKVDWTPELHRKFVQAVEKLGVDQAIPSRILEVMKVEGLTRHNVASHLQKYRMHRRHILPKEDERQWTQHRDQVQRSYYPHKPIMAYP
PYHSNHALPPGPVYPMWGATGSHTAGVHMWGPPGYSPWPPTESCHWKPYPGMHADAWGCPVMPPHNPYSTIPHYASGFQSASMVNNSCGMPQKLFDLQPA
EEVIDKVVKEAINKPWLPLPLGLKPPSADSVLAELSRQGVSSIPPRINGSNSF
|
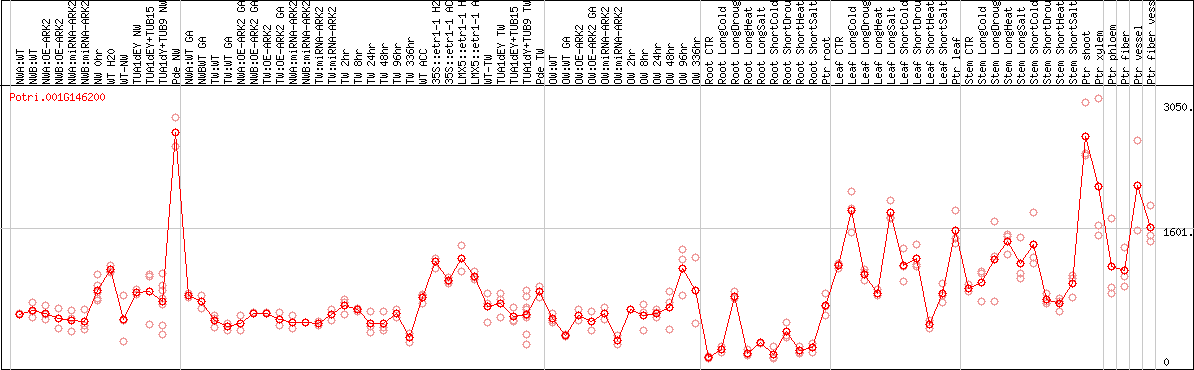
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G146200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
