External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G01980 323 / 6e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
AT1G62610 116 / 4e-31
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
AT3G55290 115 / 2e-30
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G63380 115 / 2e-30
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G17845 114 / 4e-30
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G46170 113 / 9e-30
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G55310 110 / 1e-28
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29150 65 / 4e-12
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47130 62 / 4e-11
AtSDR3
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47120 59 / 5e-10
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G067332
456 / 2e-164
AT3G01980 323 / 4e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
Potri.016G035200
115 / 1e-30
AT3G55290 385 / 8e-136
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.016G073900
66 / 1e-12
AT3G51680 252 / 2e-83
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G206400
64 / 1e-11
AT3G51680 245 / 2e-80
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G073700
63 / 2e-11
AT1G52340 242 / 1e-79
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G039500
62 / 3e-11
AT2G29150 351 / 5e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.012G125900
62 / 4e-11
AT3G12800 421 / 9e-150
short-chain dehydrogenase-reductase B (.1)
Potri.001G024300
62 / 5e-11
AT1G52340 408 / 6e-145
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G200100
62 / 5e-11
AT3G26770 257 / 2e-85
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012548
381 / 6e-135
AT3G01980 337 / 2e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
Lus10041545
381 / 9e-135
AT3G01980 346 / 6e-121
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
Lus10006699
119 / 6e-32
AT3G46170 404 / 2e-143
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10007038
109 / 3e-28
AT1G63380 357 / 7e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10007039
106 / 5e-27
AT3G55290 369 / 3e-129
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10006698
90 / 5e-21
AT1G63380 385 / 5e-136
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10018323
71 / 4e-14
AT3G12800 395 / 4e-139
short-chain dehydrogenase-reductase B (.1)
Lus10014227
70 / 1e-13
AT3G51680 231 / 3e-75
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10017129
68 / 5e-13
AT3G12800 391 / 7e-138
short-chain dehydrogenase-reductase B (.1)
Lus10006696
64 / 1e-12
AT3G46170 78 / 1e-32
NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Potri.001G327800.2 pacid=42790558 polypeptide=Potri.001G327800.2.p locus=Potri.001G327800 ID=Potri.001G327800.2.v4.1 annot-version=v4.1
ATGGGGAGTGGCGTGAAGAAAGTGTTGCTTACTTCAAATGGAGATGAGATTTCGCAGAATATCGCGTTTCATTTGGTTAAACAAGGATGCAGGTTGGTTT
TGATGGGAAATGAAAAGGGTTTAAGGAGTATCGGTGAGAAGATAACAGGTGCAATCAAAGGTGCTTTGCCAGTGGAGGTAGTGGAAATGGATATGGAAGA
GGAAAGGGAGGCAAGTTTTGAAGAGGCAGTGGACAAGGCATGTGGGATTTTGGGGAATTTAGATGCTTTTGTGCATTGCTATACATATGAAGGAAAGATT
CAAGAACCTCTGCAAGTAACAGAACATGAGTTCAAAAAATTAGTGAAAATAAATTTTATGGCTCCATGGTTTCTGCTGAAGGCTGTTGGAAAAAAAATGC
GGGACTATAATTCAGGAGGCTCCATTATATTCTTGACCTCAATAATTGGTGCTGAGAGAGGACTGTATCCAGGGGCTGCTGCATATGGTTCATGTTCAGC
AGGAATACAGCAATTAGTTAGACATTCAGCTATGGAGATAGGAACATACAAGATCAGGGTTAATGGAATCGCTCGTGGTTTGCACCTAGAAGATGCATAT
CCTGTTTCAGAAGGGAAGGAGAGGGCAGAGAAGTTGGTGAAAGAGGTAGTGCCGATGCAGAGATGGCTTGATGCCAAGAAGGATGTAGCATCAACTGTCA
TCTACTTAATCAGTGATGGTTCAAGATACATGACTGGGACAACCATATTTCTTGACGGGGGGCAGTCCTTGACAAGGCCTAGGATGCGCTCATACATGTA
G
AA sequence
>Potri.001G327800.2 pacid=42790558 polypeptide=Potri.001G327800.2.p locus=Potri.001G327800 ID=Potri.001G327800.2.v4.1 annot-version=v4.1
MGSGVKKVLLTSNGDEISQNIAFHLVKQGCRLVLMGNEKGLRSIGEKITGAIKGALPVEVVEMDMEEEREASFEEAVDKACGILGNLDAFVHCYTYEGKI
QEPLQVTEHEFKKLVKINFMAPWFLLKAVGKKMRDYNSGGSIIFLTSIIGAERGLYPGAAAYGSCSAGIQQLVRHSAMEIGTYKIRVNGIARGLHLEDAY
PVSEGKERAEKLVKEVVPMQRWLDAKKDVASTVIYLISDGSRYMTGTTIFLDGGQSLTRPRMRSYM
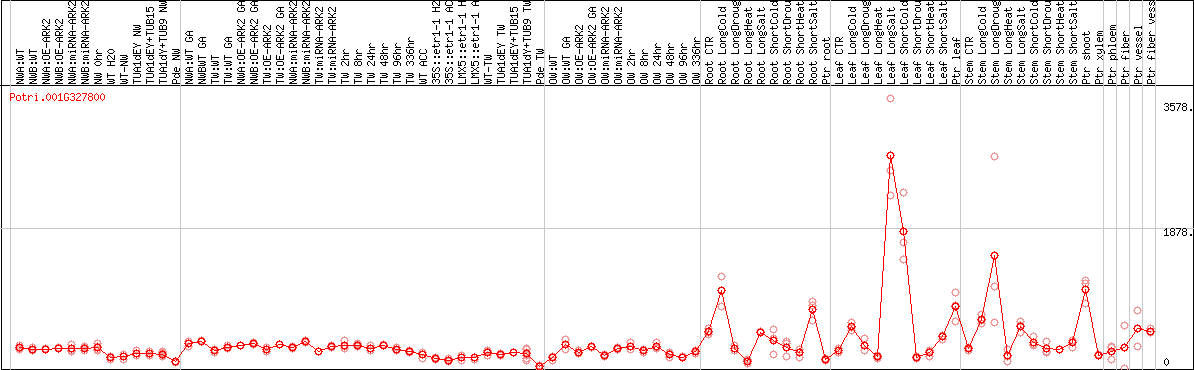
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G327800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.