External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G50940 223 / 2e-69
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT3G50930 221 / 2e-67
BCS1
cytochrome BC1 synthesis (.1)
AT5G17760 193 / 3e-57
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
AT2G18193 192 / 7e-57
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G17740 186 / 1e-54
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G17730 183 / 5e-54
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT3G28600 183 / 5e-54
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT3G28610 181 / 7e-53
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT3G28580 180 / 2e-52
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT2G18190 179 / 6e-52
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G020900
231 / 6e-72
AT3G50930 561 / 0.0
cytochrome BC1 synthesis (.1)
Potri.005G119900
223 / 8e-69
AT5G17760 512 / 3e-179
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
Potri.007G020800
222 / 3e-68
AT5G17760 506 / 1e-176
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
Potri.002G032700
207 / 3e-63
AT3G50930 437 / 1e-149
cytochrome BC1 synthesis (.1)
Potri.007G020500
205 / 2e-62
AT2G18193 511 / 1e-179
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.008G177200
203 / 2e-61
AT3G50940 401 / 4e-136
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.007G020600
201 / 3e-60
AT2G18193 553 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.007G012400
195 / 9e-59
AT3G50940 367 / 2e-123
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.011G111200
195 / 1e-58
AT3G50940 302 / 2e-98
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024275
207 / 1e-62
AT3G50940 434 / 2e-149
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10007391
202 / 8e-61
AT3G50940 424 / 2e-145
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10015802
195 / 2e-58
AT3G50940 462 / 9e-161
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10037004
193 / 9e-58
AT3G50930 457 / 9e-159
cytochrome BC1 synthesis (.1)
Lus10014284
193 / 2e-57
AT3G50930 535 / 0.0
cytochrome BC1 synthesis (.1)
Lus10032188
188 / 2e-55
AT5G40010 497 / 4e-173
ATPase-in-Seed-Development, AAA-ATPase 1 (.1)
Lus10014496
187 / 4e-55
AT5G40010 577 / 0.0
ATPase-in-Seed-Development, AAA-ATPase 1 (.1)
Lus10025989
175 / 1e-51
AT3G50930 445 / 1e-153
cytochrome BC1 synthesis (.1)
Lus10041918
175 / 5e-50
AT3G50930 383 / 2e-126
cytochrome BC1 synthesis (.1)
Lus10031318
167 / 2e-47
AT2G18193 415 / 1e-140
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00004
AAA
ATPase family associated with various cellular activities (AAA)
CL0023
PF14363
AAA_assoc
Domain associated at C-terminal with AAA
Representative CDS sequence
>Potri.001G392300.2 pacid=42791657 polypeptide=Potri.001G392300.2.p locus=Potri.001G392300 ID=Potri.001G392300.2.v4.1 annot-version=v4.1
ATGGTGTCTCTCCAGAACTTGCCTAATACAAAAACAGTTCTTTCTGTGGTAGCTTCTCTTGCAGCTTCAGCAGTGCTGATTCCAACAGCTGCCAATCTTC
GCATTTTTGCTCATCTTTTCCGTCCTCAATTCACTCTTGTTATAGAAGAATATGGACCGGATTACTTCTGTGATGAGTTGTTTTTGGCTGCTGAGACTTA
TCTGGGAACTAAATCGGCTCCATCAATTCGAAGAATAAAGGCATGCAAGAAGGAGAAAGAGAAAAAACCAGCGATTTCCCTGGACAGAGACCAAGAAATT
CTTGATGTTTTTGAAAATATTGAAGTAAAGTGGAGGATGGTTATTAGAGAAAACTCTGAGGTTAGAAATTATACCTTAGTAGCACGACTAAGATCCTATG
AGCTGGTTTTTCACAAGAAGCACAAAGAGAAGGTCCTGGGTTCTTACCTGCCGTTCATTTTACGGCAGGCGAAGGCCATTCAAGAGGAAAACAAGGTAAG
ACAGCTCAACAGTCTTGGAGGTTTATCTTGGTTAACATCAACCATAATAGATCATCCGATGACCTTTGAGACCATTGCAATGGATGAGAGGCTCAAGGAA
GAAATAATAGGTGATCTGAACACCTTCGTGAAGTCAAAGGAGTACTACAGAAAAATCGGGAAAGCTCGGAAACGTGGATACCTGATACACGGTCCTCCTG
GAACAGGAAAGTCGAGCTTGATCGCCGCCATGGCTAATCACTTGAACTACAGTATCCATGACTTGGATCTACAGGATGATAATTTCCTTACAAGTTACGA
TATTAGGTCAGTGTTGCTTCACCATTTAGACAAAACTATACTTGTCATCAAAGATATTGATTCCACTGTCACAGTATCATGCCGAAGGCGGAGATTTGAG
CCAAGGAAGGTGGTTGAGGTTCAAAAACAGGCTATGCTCTTAAGGCTACTGCAGTTTATTGATGGACTTTTGGCTGCCTCGCATCAATGA
AA sequence
>Potri.001G392300.2 pacid=42791657 polypeptide=Potri.001G392300.2.p locus=Potri.001G392300 ID=Potri.001G392300.2.v4.1 annot-version=v4.1
MVSLQNLPNTKTVLSVVASLAASAVLIPTAANLRIFAHLFRPQFTLVIEEYGPDYFCDELFLAAETYLGTKSAPSIRRIKACKKEKEKKPAISLDRDQEI
LDVFENIEVKWRMVIRENSEVRNYTLVARLRSYELVFHKKHKEKVLGSYLPFILRQAKAIQEENKVRQLNSLGGLSWLTSTIIDHPMTFETIAMDERLKE
EIIGDLNTFVKSKEYYRKIGKARKRGYLIHGPPGTGKSSLIAAMANHLNYSIHDLDLQDDNFLTSYDIRSVLLHHLDKTILVIKDIDSTVTVSCRRRRFE
PRKVVEVQKQAMLLRLLQFIDGLLAASHQ
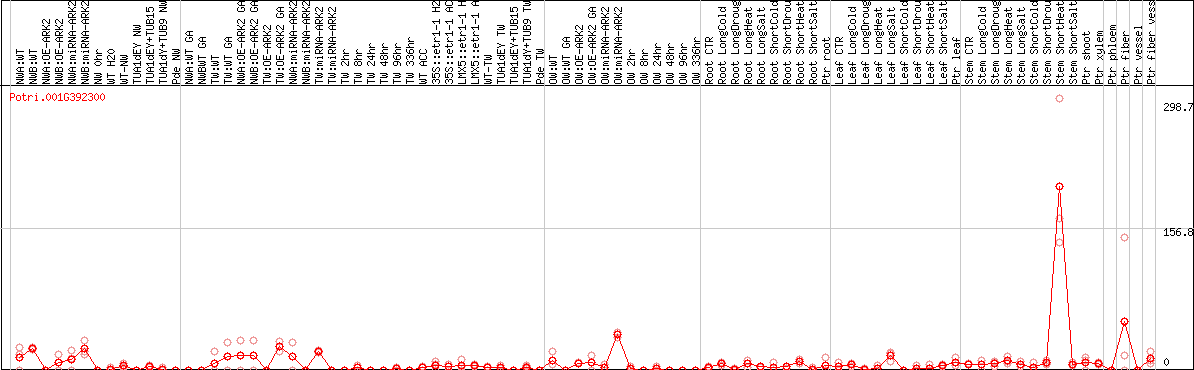
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G392300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.