Potri.001G426030 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G426030.1 pacid=42788769 polypeptide=Potri.001G426030.1.p locus=Potri.001G426030 ID=Potri.001G426030.1.v4.1 annot-version=v4.1
ATGGCTCTCGAAAGTGCTGGTGGATCCATTATAGCTATGCTAGCAGAACTCATGGTGGAACCAGTAGGAAGGCAGTTCCGTTACATGTTCTGTTTCAACA
ATTTTGCTCAAGAATTCAAAGAACAAAAGGAGAACCTGGTTTCGGCAAAAGAACGTCTGCAAGACGATGTCGAAGCTGCTGAAAGGAATGCTGAAAAAAC
TTACAGAGATGTCAAAAAGTGGCTGGAAGATGCAAACAACCAAATTGAAGGTGCGAAGCCCTTGGAAAATGAAATAGGAAAAAACGGCAAATGCTTTACT
TGGTGCCCAAACTGCATGCGACAATTCAAGTTAAGCAAGGCACTGGCCAAGAAGTCGGAGACTTTCAGAAAAATTCTAGAAAATAGCACAAAGTTTAAAA
CAGTGGCCCAAAAAGCTCCTCCTCAGCCCATAGAATTTCTAACATCAAAGGAATTCACGCCCTCAGAATCGTCAAAAGAAGCTTTGGAACAGATCATGAA
AGCTCTCAAAGATGACACCGTCAATATGATCGGACTGTACGGCATGGGAGGGGTGGGTAAAACAACCCTGGTGAAAGAAGTAGGCAGGAGAGCCAAAGAG
TCGCAGCTTTTTCCTGACGTTTTGATGGCTACGGTGTCCCAGAATCCAAATTTCATAGGCATCCAGGATCGAATGGCAGATAGTTTACATCTGAAATTTG
AAAAAACGAGTAAAGAAGGTAGAGCAAGTGAATTATGGCAGAGACTGCAGGGAAAGAAGATGCTTATAATCCTAGATGATGTTTGGAAACATATTGACTT
GGAAGAGATAGGGATCCCATTTGGTGATGATCACAGGGGTTGTAAAATTCTTCTAACAACACGTGTTCAAGGCATATGTTTTTCTATGGAGTGCCAGCAA
AAAGTGCTTTTAAAAACTATCTTCATAAAAGAGTGTGGTAAACTGGAATATGTCTTCCCTGTCTCCATGTCTCCAAGCCTTCCGAACCTGGAACAGATGA
CGATTTATTATGCTGACAATTTAAAGCAAATATTTTACAGTGGAGAAGGAGATGCACTCACCACAGATGGCATCATCAAGTTCCCTCAACTAAGAGAATT
GTCTCTTGGGCTCCGATCAAATTACGGCTTTATAGGTCCAAGGAATTTTGATTTTCAATTGCCTTCTTTGCAGAATCTAAACATTAAAGGCCACGAAGAA
GTGGGTAATTGGTTGGCACAGCTACAACAAAACGGCTTCTTACAAAGATTAGAAGCTGTACACATGAGGGATTGTGGGGATGTTCGCGCTCCATTTCCAG
CAAAATTGCTGCGAGCTGTGAAAAATCTAAAGAAAGTGGGTGTTTGGTGCTGCAAATCTTTGGAAGAGGTATTTGAATTAGGTGAGGCTGATGAAGGAAG
CAGTGAGGAGAAGGAGCTGCTGTCATCTTTAACAGAGTTACTGCTGTCAGGGTTACCTGAGCTCAAATGCATTTGGAAGGGGCCCACCAGACATGTCAGC
CTCCAAAGTCTTGCTTATCTGTATTTGAATTCTCTCGACAAACTGACATTTATCCTCACACCGTCTCTCGCTCGAAGTCTGCCAAAGCTAGAAACACTTG
AGATAAGTGAATGCGGTGAATTGAAGTATATTATCAGAGAAGAGGATGGTGAAAGGGAAATAATTCCAGAGTCTCCTTGCTTCCCAAGATTAAAAACTAT
CTTCATAAAAGACTGTGGTAAACTGGAATATGTCTTCCCTGTCTCTGTGTCTCCGAGCCTTCCGAACCTGGAATTGATGACGATTGATCGTACTGACAAT
TTAAAGCAAATATTTTACAGTGGAGAAGGAGACGCACTCACCACAGATGGCATCATCAAGTTCCCTCGATTAAGTGACTTGGTTCTTTCTTCCATATCAA
ATTACAGCTTTTTTGGTCCAACGAATCTTGCTGCCCAATTGCCTTCTTTGCGATTTCTAAAAATCAATGGCCACAAAGAATTGGGAAATTTGTTTGCACA
GCTCCAAGGGTTCACAAATTTGAAAGAATTAAGCTTGGAATCCGTGCCTGACTTGAGGGGTCTTCTGCTGAGCAAATTGACTACTTTGGAGATGGCCGCC
CATGGGGAGCAAAACGGCTCCTTACATAGATTAGAACGTGTACGAGTGGACGATTGTGGGGATGTTCGCGCTCCGTTTCCAGCAAAATTGCTGCGAGCTT
TGAAAAATCTAAGCAGTGTGAACATTAACGGCTGCAAATCATTGGAAGAGGTATTTGAATTAGGTGAGCCTGATGAAGGAAGTAGGGAGGAGAAGGAGCT
GCCGCTTCTGTCATCTTTAACAGGGTTACGGCTGTCAGGTTTACCTGAGCTCAAATGCATGTGGAAGGGGCCCACCAGACATGTCAGCCTCCAAAGTCTT
GCTTATCTGGATTTGTGGTCTCTGGACAAACTGACATTTATATTCACACCGTCCCTCGCTCGAAGTCTGCCAAAGCTAGAAAGACTTTACATAGGTAAAT
GCGGTCAATTGAAGCATATTATCAGAGAAGAGGATGGTGAAAAGGAAATAATTCCAGAGCCTCCTGGGCAGGATGGTCAAGCTTCACCTATCAACGTTGA
GAAGGAGATAGTGCTTCCTAATCTAAAGGAGTTGTCGATACAACAATTATCAAGTATTGTCTGTTTTAGTTTCGGATGGTGTGATTATTTGTTATTCCCT
CGTTTGGAGAAGTTGGAGGTCCATCTATGTCCAAAGCTGACCACAAAATTTGCTAGTACACCAGATGGTTCAATGAGTGCTCAATCAGAGGTATCTGAAG
TAGCTGAAGATTCAAGCATTTATAGAGAGTGGACCCGAGATAAGGGGTGGAAAGAACGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G426030.1 pacid=42788769 polypeptide=Potri.001G426030.1.p locus=Potri.001G426030 ID=Potri.001G426030.1.v4.1 annot-version=v4.1
MALESAGGSIIAMLAELMVEPVGRQFRYMFCFNNFAQEFKEQKENLVSAKERLQDDVEAAERNAEKTYRDVKKWLEDANNQIEGAKPLENEIGKNGKCFT
WCPNCMRQFKLSKALAKKSETFRKILENSTKFKTVAQKAPPQPIEFLTSKEFTPSESSKEALEQIMKALKDDTVNMIGLYGMGGVGKTTLVKEVGRRAKE
SQLFPDVLMATVSQNPNFIGIQDRMADSLHLKFEKTSKEGRASELWQRLQGKKMLIILDDVWKHIDLEEIGIPFGDDHRGCKILLTTRVQGICFSMECQQ
KVLLKTIFIKECGKLEYVFPVSMSPSLPNLEQMTIYYADNLKQIFYSGEGDALTTDGIIKFPQLRELSLGLRSNYGFIGPRNFDFQLPSLQNLNIKGHEE
VGNWLAQLQQNGFLQRLEAVHMRDCGDVRAPFPAKLLRAVKNLKKVGVWCCKSLEEVFELGEADEGSSEEKELLSSLTELLLSGLPELKCIWKGPTRHVS
LQSLAYLYLNSLDKLTFILTPSLARSLPKLETLEISECGELKYIIREEDGEREIIPESPCFPRLKTIFIKDCGKLEYVFPVSVSPSLPNLELMTIDRTDN
LKQIFYSGEGDALTTDGIIKFPRLSDLVLSSISNYSFFGPTNLAAQLPSLRFLKINGHKELGNLFAQLQGFTNLKELSLESVPDLRGLLLSKLTTLEMAA
HGEQNGSLHRLERVRVDDCGDVRAPFPAKLLRALKNLSSVNINGCKSLEEVFELGEPDEGSREEKELPLLSSLTGLRLSGLPELKCMWKGPTRHVSLQSL
AYLDLWSLDKLTFIFTPSLARSLPKLERLYIGKCGQLKHIIREEDGEKEIIPEPPGQDGQASPINVEKEIVLPNLKELSIQQLSSIVCFSFGWCDYLLFP
RLEKLEVHLCPKLTTKFASTPDGSMSAQSEVSEVAEDSSIYREWTRDKGWKER
|
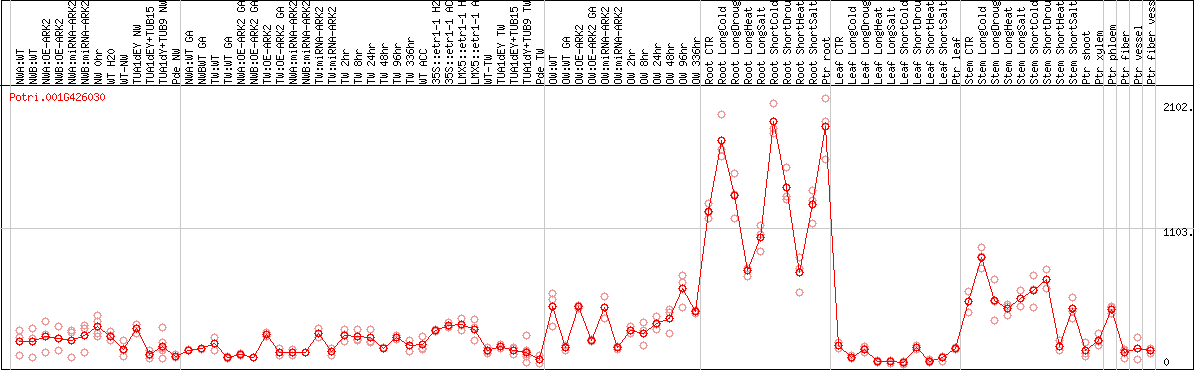
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G426030 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
