External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G27220 194 / 2e-55
NB-ARC domain-containing disease resistance protein (.1)
AT4G27190 191 / 2e-54
NB-ARC domain-containing disease resistance protein (.1)
AT4G26090 172 / 1e-47
RPS2
RESISTANT TO P. SYRINGAE 2, NB-ARC domain-containing disease resistance protein (.1)
AT5G63020 169 / 9e-47
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G61180 164 / 5e-45
LRR and NB-ARC domains-containing disease resistance protein (.1.2)
AT1G12220 162 / 2e-44
RPS5
RESISTANT TO P. SYRINGAE 5, Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
AT1G61300 161 / 3e-44
LRR and NB-ARC domains-containing disease resistance protein (.1)
AT1G61190 157 / 2e-42
LRR and NB-ARC domains-containing disease resistance protein (.1)
AT1G12280 152 / 4e-41
LRR and NB-ARC domains-containing disease resistance protein (.1)
AT1G61310 152 / 7e-41
LRR and NB-ARC domains-containing disease resistance protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020016
92 / 5e-20
AT3G07040 563 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10022464
89 / 4e-19
AT5G66900 531 / 2e-177
Disease resistance protein (CC-NBS-LRR class) family (.1)
Lus10002945
88 / 7e-19
AT3G07040 580 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10024173
86 / 4e-18
AT3G14460 681 / 0.0
LRR and NB-ARC domains-containing disease resistance protein (.1)
Lus10005218
86 / 4e-18
AT3G14470 379 / 2e-113
NB-ARC domain-containing disease resistance protein (.1)
Lus10022351
85 / 6e-18
AT3G14470 369 / 1e-108
NB-ARC domain-containing disease resistance protein (.1)
Lus10041243
84 / 2e-17
AT3G07040 526 / 1e-172
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10042117
82 / 6e-17
AT3G14470 354 / 2e-105
NB-ARC domain-containing disease resistance protein (.1)
Lus10018937
81 / 2e-16
AT3G07040 587 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10014712
81 / 3e-16
AT3G07040 492 / 3e-159
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Potri.001G427440.1 pacid=42791713 polypeptide=Potri.001G427440.1.p locus=Potri.001G427440 ID=Potri.001G427440.1.v4.1 annot-version=v4.1
ATGCTTATAATCCTGGATGATGTTTGGAAACATATTGACTTGGAAGAGATAGGGATCCCATTTGGTGATGATCACAGGGGTTGTAAAATTCTTCTAACAA
CACGTGTTCAAGGCATATGTTTTTCTATGGAGTGCCAGCAAAAAGTGCTTTTAAGAGTCTTACCTGAAGATGAAGCATGGGATTTATTCAGAATCAATGC
AGGTTTACGTGATGGGGACTCTACCTTGAACACAGTGGCAAGGGAGGTTGCGAGAGAATGCCAAGGCTTGCCTATAGCACTTGTGACAGTGGGAAGGGCT
CTAAGAGGTAAATCACGAGTTCAGTGGGAAGTAGCGTCTAAACAGCTCAAAGAATCTCACTTTGTGCGCATGGAACAAATTGATGAACAAAATAATGCTT
ACACATGTCTTAAGTTGAGCTATGATTATTTGAAGTACGAGGAAACCAAGTCATGTTTCGTGCTATGCTGTTTATTTCCAGAAGATTATGACATTCCAAT
CGAGGACTTGACGAGATATGCAGTTGGCTATGGGTTACATCAAGATGCGGAGCCCATTGAAGATGCAAGGAAAGGAGTTTCTGTGGCCATCGAAAACCTC
AAAGATTGTTGCATGCTGTTGGGCACTGAAACTGAAGAACATGTGAGAATGCATGACTTGGTTCGTGACTTTGCTATTCAGATAGCATCATCAGAAGAAT
ATGGATTCATGGTAAAGGCTGGCATTGGGTTGGAGAAGTGGGCAATGAGAAACAAAAGCTTTGAAGGTTGTACAACAATTTCGTTAATGGGCAATAAACT
AGCAGAACTTCCTGAAGGATTGGTATGTCCACAGCTCAAAGTTCTATTATTAGAACTGGAAGATGGTATGAATGTTCCAGAGAGGTTTTTTGAAGGGATG
AAAGAAATTGAAGTTTTGTCTCTGGAGGAGGGCGTTTGTCGCTGCAATCACTTGAACTCTCAACGAAACTTCAATCGTTGGTGTTGA
AA sequence
>Potri.001G427440.1 pacid=42791713 polypeptide=Potri.001G427440.1.p locus=Potri.001G427440 ID=Potri.001G427440.1.v4.1 annot-version=v4.1
MLIILDDVWKHIDLEEIGIPFGDDHRGCKILLTTRVQGICFSMECQQKVLLRVLPEDEAWDLFRINAGLRDGDSTLNTVAREVARECQGLPIALVTVGRA
LRGKSRVQWEVASKQLKESHFVRMEQIDEQNNAYTCLKLSYDYLKYEETKSCFVLCCLFPEDYDIPIEDLTRYAVGYGLHQDAEPIEDARKGVSVAIENL
KDCCMLLGTETEEHVRMHDLVRDFAIQIASSEEYGFMVKAGIGLEKWAMRNKSFEGCTTISLMGNKLAELPEGLVCPQLKVLLLELEDGMNVPERFFEGM
KEIEVLSLEEGVCRCNHLNSQRNFNRWC
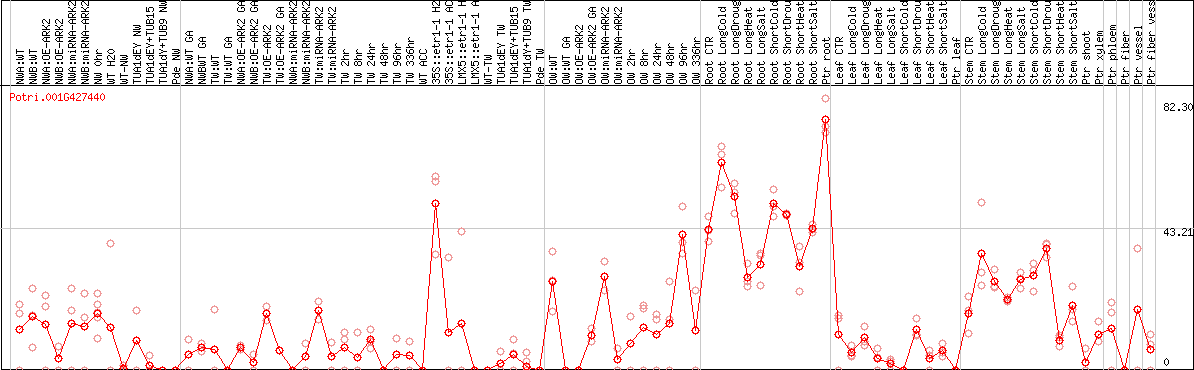
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G427440 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.