External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G61310 69 / 4e-13
LRR and NB-ARC domains-containing disease resistance protein (.1)
AT5G05400 65 / 8e-12
LRR and NB-ARC domains-containing disease resistance protein (.1)
AT1G61190 63 / 2e-11
LRR and NB-ARC domains-containing disease resistance protein (.1)
AT1G61180 62 / 8e-11
LRR and NB-ARC domains-containing disease resistance protein (.1.2)
AT1G12210 60 / 3e-10
RFL1
RPS5-like 1 (.1)
AT1G63350 59 / 6e-10
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G61300 59 / 7e-10
LRR and NB-ARC domains-containing disease resistance protein (.1)
AT1G51480 57 / 2e-09
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G12290 57 / 3e-09
Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
AT5G43740 56 / 6e-09
Disease resistance protein (CC-NBS-LRR class) family (.1), Disease resistance protein (CC-NBS-LRR class) family (.2)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020533
45 / 3e-05
AT1G27170 405 / 1e-122
transmembrane receptors;ATP binding (.1.2)
Lus10008221
42 / 0.0004
AT4G12010 400 / 8e-120
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10006789
42 / 0.0004
AT4G12010 325 / 7e-96
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10002247
42 / 0.0005
AT5G36930 363 / 1e-108
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10007828
42 / 0.0005
AT5G36930 319 / 7e-92
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10007790
41 / 0.0005
AT5G36930 343 / 4e-99
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10004257
41 / 0.0006
AT5G36930 446 / 7e-136
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10023051
41 / 0.0007
AT4G12010 415 / 2e-125
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10042165
41 / 0.0008
AT5G36930 326 / 9e-94
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10016029
41 / 0.0008
AT4G12010 451 / 2e-138
Disease resistance protein (TIR-NBS-LRR class) family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Potri.001G427670.1 pacid=42792063 polypeptide=Potri.001G427670.1.p locus=Potri.001G427670 ID=Potri.001G427670.1.v4.1 annot-version=v4.1
ATGGCTCTCGAAAGTGCTGGTGGATCCATTATAGCTATGCTAGCAGAACTCATGGTGGAACCAGTAGGAAGGCAGTTCCGTTACATGTTCTGTTTCAACA
ATTTTGCTCAAGAATTCAAAGAACAAAAGGAGAACCTGGTTTCGGCAAAAGAACGTCTGCAAGACGATGTCGAAGCTGCTGAAAGGAATGCTGAAAAAAC
TTACAAAGATGTCAAAAAGTGGCTGGAAGATGCAAACAACCAAATTGAAGGTGCGAAGCCCTTGGAAAATGAAATAGGAAAAAACGGCAAATGCTTTACT
TGGTGCCCAAACTGCATGCGACAATTCAAGTTAAGCAAGGCACTGGCCAAGAAGTCGGAGACTTTCAGAAAAATTCTAGAAAATAGCACAAAGTTTAAAA
CAGTGGCCCAAAAAGCTCCTCCTCAGCCCATAGAATTTCTAACATCAAAGGAATTCACGCCCTCAGAATCGTCAAAAGAAGCTTTGGAACAGATCATGAA
AGCTCTCAAAGATGACACCGTCAATATGATCGGACTGTACGGCATGGGAGGGGTGGGTAAAACAACCCTGGTGAAAGAAGTAGGCAGGAGAGCCAAAGAG
TCGCAGCTTTTTCCTGACGTTTTGATGGCTACGGTGTCCCAGAATCCAAATTTCATAGGCATCCAGGATCGAATGGCAGATAGTTTACATCTGAAATTTG
AAAAAAACGAGTAA
AA sequence
>Potri.001G427670.1 pacid=42792063 polypeptide=Potri.001G427670.1.p locus=Potri.001G427670 ID=Potri.001G427670.1.v4.1 annot-version=v4.1
MALESAGGSIIAMLAELMVEPVGRQFRYMFCFNNFAQEFKEQKENLVSAKERLQDDVEAAERNAEKTYKDVKKWLEDANNQIEGAKPLENEIGKNGKCFT
WCPNCMRQFKLSKALAKKSETFRKILENSTKFKTVAQKAPPQPIEFLTSKEFTPSESSKEALEQIMKALKDDTVNMIGLYGMGGVGKTTLVKEVGRRAKE
SQLFPDVLMATVSQNPNFIGIQDRMADSLHLKFEKNE
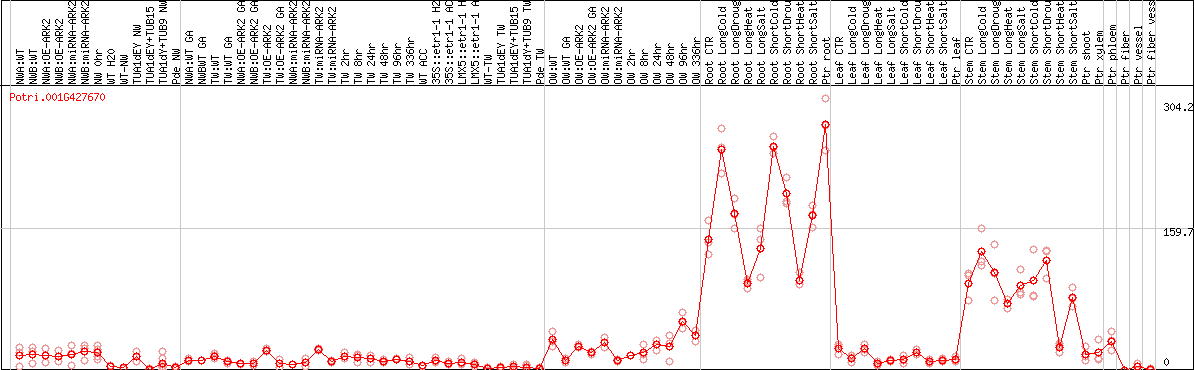
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G427670 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.