Potri.001G444050 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G444050.1 pacid=42788263 polypeptide=Potri.001G444050.1.p locus=Potri.001G444050 ID=Potri.001G444050.1.v4.1 annot-version=v4.1
ATGAGCAGCTCGTATCATTCGTGCAGGTGTCAAAGTGATCATCATCAACCTTCCATACAGAGAGACTGGTGGAAAATGGCTATCGAAAGTGTTGGTGGGT
CCGTTGTATCTAAGATAGCAGAGCTCTTGGTGGAACCAGCAATCAAGCAGTTCCGTTACATGTTCTGTTTCAACAATTTTGTTCAAGAATTCGATGAACA
AATGATGAACCTGGCTTTGGCATTTTATCGTCTGCAAGATGCTGTCAACGTAGCTGAAAGGAATGCTGAAGAAATTGAGATTGATGTCAACACATGGCTG
GAAAATGCAAAAAACGAAATTGAAGGTGTGAATCGTTTGCAAAATGAAAAAGGAAAAATCATCAAATGCTTTACTTGGTGTCCAAATTCGATGCGACAAT
TCAAGTTAAGCAAGGCACTGGCAAAGAAGACGGAGACTTTGAGAAAACTTGAAGAAAATAGCAGAAAGTTTCCAAAAGTGTCCCACAAAGCACCTCTTCA
AGAGATAAAATTTCTTCCATCAAAGGAATTCACACTCTCAGGATCGTCAAAAGAAGCTTTCAAACAGATTATGAAAGCTCTCAAAGATGACAAGGTCAAT
ATGATCGGACTGTACGGCATGGGAGGGGTGGGTAAAACCACCCTGGTGAAAGAAGTAGGCAGGAGAGCCAAAGAGTTGCACCTTTTTGATGAAGTTTTGA
TAGCTACGGTGTCCCAGAATCCAAATGCCACAGGCATCCAGGATCAAATGGCAGATAGTTTAGATCTGCAATTTGACAAGAAGAGTAAAGAAGGGAGAGC
AAAAGAACTATGGCAGAGACTGCAGGGAAAGAAGATGCTTATAGTCCTGGATGATGTTTGGAAAGATATTGACTTCCAAGAGATAGGGATCCCATTTGGT
GATGATCACAGGGGTTGTAAAATTCTTCTAACAACACGTCTTGAAGACATGTGTTCTTATATGAAGTGCAAGGAAAAAGTTTTTTTATGTCTCTTTTCTA
AAGAGGAAGCATGGGCTTTATTCAGAATCAATGCAGCTTTACGTGACGAGGACTCTACCTTGAACAGGGTGGCAAAGAAGGTTGCGAGAGAATGTAATGG
ATTGCCTGTAGCACTTGTGACAGTGGGAAGGGCTCTAAGAGATAAATCTGTTGTTGAGTGGGAAGTAGCGTCTGAAGAGCTCAAAAACTCTCAATTTCGG
CATTTGGAACAAATTGACGGACAAAAAAATGCATATGCATGTCTTAAGTTGAGCTATGATTATTTGACGTCCGATGAAACCAAGTCATGTTTCTTGCTAT
GCTGTTTATTTCCAGAAGATTACGACATTCCGATTGAGGACTTGACGAGATACGCAGTTGGCTATGGGTTACATAAAGATGCGGAGTCCATTGAAGATGC
AAGGAAACGAGTTTATGTGGCAATCAAAAACCTCAAAGCTTGTTGTATGCTGTTAGGAACTTTTAGTGAAGAATATGTCAAAATGCATGACTTGGTTCGT
GATGTTGCTATTCGGATAGCATCATCAGAAAAATACGGATTCATGGTAAAGGCTGGCATTGGGTTGCTAGAGTGGCCAACGAGCAATAAAAGCTTTGAAG
GTTGTACAACAATTTCGTTAATGGGCAATAAACTAGCAGAACTTCCTGAAGGATTGGTGTGTCCACAGCTCAAAGTTCTATTGTTAGGACTGGATCGTGG
TTTGAATGTTCCAGAGAGGTTTTTTGAAGGGATGAAAGCAATAGAAGTTTTGTCTCTAAAGGGAGGGTGTTTGTCATTGCAATCACTTCAATTCTCAACG
AACCTTCAATCGTTGCTGTTGATCGAGTGTGAATGCAAGGACCTCATTTGGTTGAGAAAGCTGCAAAGACTTAATATTCTTGTTTTCAGGCGGTGCGGCT
CCATTGAAGAATTACCTGATGAAATTGGGGAGCTCAAGGAGTTGAGGTTGTTGGATTTGACAGGTTGTGAAAATCTAAGAAGGATTCCTGTGAATTTGAT
AGGAAGGTTGAAGAAGTTAGAAGAACTGCTGATCGGGGATCGGAGCTTCAAGGGATGGGATGTTGTTGGATGTGACAGCACAGAAGGAATGAATGCAAGC
CTAACAGAACTAAACTCGTTGTCTCATTTAGCTGTATTATCATTGAAGATACCGAAGGTTGAATGCATTCCCAGAGATTTTGTTTTTCCCAGGTTGCTCA
AATATGATATAGTGTTAGGGGATTGGTATTCAGGACCCCATAAAGAATACCCAACCTCGACAAGATTATATTTGGGTGATATAAGCGCCACATCCTTAAA
TGCAAAGACATTTGAGCAGTTGTTTCCTACCGTGTCTCATATTTGGTTTTGGAGAGTGGAGGGTTTAAGAAATATAGTATTGTCCTCTGATCAGATGACC
TCCCATGGCCATGGGTCGCAAAAGGACTTCTTTCAAAGATTAGAATATGTAGCAGTGAGAGAATGTGATGATATTCGCACTCTGTTTCCAGCAAAATGGC
GGCAAGCTTTGAAAAATCTAAGGAGAGTGGAAATTGAAGACTGCCAATCATTGGATGAAGGAATTAACGAGGAGAAGGAGCTGCCATTTTTAACGGAGTT
ACAGCTGTCCTGGTTACCTGAGCTCAAATGTATATGGAAGGGGCCCACTAGACATGTCAGCCTCCAAAGTCTTACTCGTCTGGAGCTCAGATGTCTTAAC
AAACTGACATTTATCTTTACTCCGTCCCTCGCTCAAAGTTTTATTCATCTGGAAACGCTACGGATAGAAAATTGCCATGGATTGAAGCGTCTTATCAGAG
AACAGGATGACGAAATCAGGACAGAGTCTCTTGGCTTCCCAAAATTAAAAAATCTCTCTATAATATTCTGTGATAAACTTGAATATGTCTTCCCTGTCTC
CGTGTCTCCAAGTCTTCAGAACCTGGAAGAGATGCAGATTGTTTTTGCTGACAATTTAAAGTCTCTTGTATTTTACAGTGGAGAAGGAGATGACATCATC
GTCAAGTCTAAAATTAAAGATGGCATCATCGACTTCCCTCAGCTAAGAGAATTGTCTCTTTCAAAGTGCAGCTTTTTTGGTCCAAAGGATTTTGCTGCCC
TATTGCCTTCTTTGCAATGTCTAAGGATCTCTGGCCTTGAAGAATGGGGTAATTTGTTGGCACAGCTTCGAGGGTTTACAAGTTTGGAAACATTGAAGTT
GTCCTCCCTGCTTGTGCCTGACTTGAGGTGTATATGGAAGGGTCTCGTGCCATGCAATTTGACTACTTTGGAGGTGAAAGAGTGTAAGAGGCTGACACAT
GTATTCACAGACAGCATGATTGCTAGTCTTGTTCAACTGAAAGTTCTAGAGATATCAAATTGTGAGGAATTGGAGCAAATCATTGCTAAGGATAATGATG
ATGAAAAGGATCAGATATTTTCAGGAAGTGATCTCCAATCTGCATGCTTTCCTAATTTGTGTCGACTTGAGATTAGAGGATGCAACAAGTTGAAGAAGTT
GGAGGTGGATGGATGCCCAAAGCTGACCATAGAATCTGCTACTACATCAAATGACTCAATGAGTGGTCAATCAGAGGGGTTTATGAATTTGAAAGAAATA
TCTATTGGAAACTTGGAAGGAGTACAAGATTTAATGCAATTTGAACGTTTGGTAACTAATAGAAGAGGTGGGCATGAACTTTCACTCGTGAGTTTGGAAA
CATTACAGTTGAACTTATTGCCTGACTTGAGGTGTATATGGAAGGGCCTAGTGCCGAGCAATTTGACTACTTTGAAGGTGAAAAGGTGTAACAGACTGAC
ACATGTATTCACAGACAGCATGATTGCTAGTCTAGTTCAACTGAAAGTTCTAGAGATATCAAACTGTGAGGAATTGGAGCAAATCATTGCTAAGGATAAT
GATGATGAAAAGGATCAGATATTTTCAGGAAGTGATCTCCAATCTGCATGCTTTCCTAATTTGTGTCGACTTGAGATCAGAGGATGCAACAAGTTGAAGA
GTCTCTTCCCAGAAGCCATGGCTTCAGGTCTCAAAAAGCTCCAAATACTTAAAGTAAGAGAATCCTCTCAATTATTGGGAGTATTTGGGCAGGATGATCA
TGCTTCACCTGTCAATGTTGAGAAGGAGATGATGCTCCCTGATCTGCAGGAGTTGTTGCTTGTACAATTACCAAGTATTTCCTGCTTTAGTCTCGGATGT
TATGATTTCTTGTTCCCTCATTTGAAGAAGTTGGAGGTGGATGGATGCCCAAAGCTGACCACAGAATCTGCTACTACATCAAATGACTCAATGAGTGCTC
AATCAGAGGGGTTTATGAATTTGAAAGAAATATCTATTGGAAACTTGGAAGGAGTACAAGATTTAATGCAAGTTGGACGTTTGGTAACTAATAGAAGAGG
TGGGCGTGAACTTTCACTCGTTAGTTTGGAAACATTACGCTTGAACTTATTGCCAGACTTGAGGTGTATATGGAAGGGTATCCTGCCGAGCAATTTGACT
ACTTTGAAGGTGAACGACTGTAAGAGACTGACACATGTATTCACAGACAGCATGATTGCTAGTCTAGTTCAACTGAAAGTTCTAGAGATATCAGCTTGTG
AGGAATTGGAGCAAATCGTTGCTAAGGATAATGATGATGAAAAGGATCAGATATTTTCAGGAAGTGATCTCCAATCTGCATGCTTTCCTAATTTGTGTCG
ACTTGAGATTAGAGGATGCAACAAGTTGAAGAAGTTGGAGGTGGATGGATGCCCAAAGCTGACCATAGAATCTGCTACTACATCAAATGACTCAATGAGT
GGTCAATCAGAGGGGTTTATGAATTTGAAAGAAATATCTATTGGAAACTTGGAAGGAGTACAAGATGTTAGCGTGAGCTGTACAGTTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G444050.1 pacid=42788263 polypeptide=Potri.001G444050.1.p locus=Potri.001G444050 ID=Potri.001G444050.1.v4.1 annot-version=v4.1
MSSSYHSCRCQSDHHQPSIQRDWWKMAIESVGGSVVSKIAELLVEPAIKQFRYMFCFNNFVQEFDEQMMNLALAFYRLQDAVNVAERNAEEIEIDVNTWL
ENAKNEIEGVNRLQNEKGKIIKCFTWCPNSMRQFKLSKALAKKTETLRKLEENSRKFPKVSHKAPLQEIKFLPSKEFTLSGSSKEAFKQIMKALKDDKVN
MIGLYGMGGVGKTTLVKEVGRRAKELHLFDEVLIATVSQNPNATGIQDQMADSLDLQFDKKSKEGRAKELWQRLQGKKMLIVLDDVWKDIDFQEIGIPFG
DDHRGCKILLTTRLEDMCSYMKCKEKVFLCLFSKEEAWALFRINAALRDEDSTLNRVAKKVARECNGLPVALVTVGRALRDKSVVEWEVASEELKNSQFR
HLEQIDGQKNAYACLKLSYDYLTSDETKSCFLLCCLFPEDYDIPIEDLTRYAVGYGLHKDAESIEDARKRVYVAIKNLKACCMLLGTFSEEYVKMHDLVR
DVAIRIASSEKYGFMVKAGIGLLEWPTSNKSFEGCTTISLMGNKLAELPEGLVCPQLKVLLLGLDRGLNVPERFFEGMKAIEVLSLKGGCLSLQSLQFST
NLQSLLLIECECKDLIWLRKLQRLNILVFRRCGSIEELPDEIGELKELRLLDLTGCENLRRIPVNLIGRLKKLEELLIGDRSFKGWDVVGCDSTEGMNAS
LTELNSLSHLAVLSLKIPKVECIPRDFVFPRLLKYDIVLGDWYSGPHKEYPTSTRLYLGDISATSLNAKTFEQLFPTVSHIWFWRVEGLRNIVLSSDQMT
SHGHGSQKDFFQRLEYVAVRECDDIRTLFPAKWRQALKNLRRVEIEDCQSLDEGINEEKELPFLTELQLSWLPELKCIWKGPTRHVSLQSLTRLELRCLN
KLTFIFTPSLAQSFIHLETLRIENCHGLKRLIREQDDEIRTESLGFPKLKNLSIIFCDKLEYVFPVSVSPSLQNLEEMQIVFADNLKSLVFYSGEGDDII
VKSKIKDGIIDFPQLRELSLSKCSFFGPKDFAALLPSLQCLRISGLEEWGNLLAQLRGFTSLETLKLSSLLVPDLRCIWKGLVPCNLTTLEVKECKRLTH
VFTDSMIASLVQLKVLEISNCEELEQIIAKDNDDEKDQIFSGSDLQSACFPNLCRLEIRGCNKLKKLEVDGCPKLTIESATTSNDSMSGQSEGFMNLKEI
SIGNLEGVQDLMQFERLVTNRRGGHELSLVSLETLQLNLLPDLRCIWKGLVPSNLTTLKVKRCNRLTHVFTDSMIASLVQLKVLEISNCEELEQIIAKDN
DDEKDQIFSGSDLQSACFPNLCRLEIRGCNKLKSLFPEAMASGLKKLQILKVRESSQLLGVFGQDDHASPVNVEKEMMLPDLQELLLVQLPSISCFSLGC
YDFLFPHLKKLEVDGCPKLTTESATTSNDSMSAQSEGFMNLKEISIGNLEGVQDLMQVGRLVTNRRGGRELSLVSLETLRLNLLPDLRCIWKGILPSNLT
TLKVNDCKRLTHVFTDSMIASLVQLKVLEISACEELEQIVAKDNDDEKDQIFSGSDLQSACFPNLCRLEIRGCNKLKKLEVDGCPKLTIESATTSNDSMS
GQSEGFMNLKEISIGNLEGVQDVSVSCTV
|
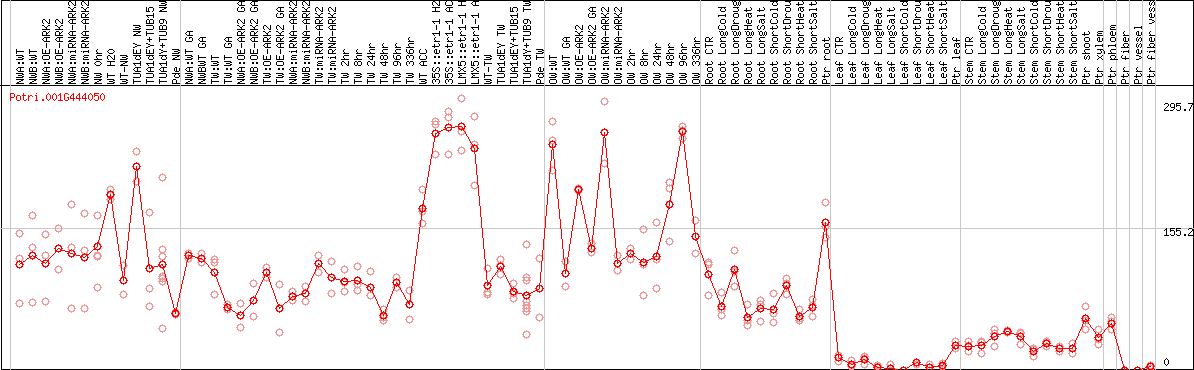
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G444050 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
