External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G59620 45 / 2e-05
CW9
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT3G46710 43 / 9e-05
NB-ARC domain-containing disease resistance protein (.1)
AT1G58602 43 / 9e-05
LRR and NB-ARC domains-containing disease resistance protein (.1.2)
AT1G58410 40 / 0.0006
Disease resistance protein (CC-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G134450
183 / 2e-53
AT3G07040 488 / 1e-157
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134300
183 / 2e-53
AT3G07040 487 / 2e-157
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134701
181 / 2e-52
AT3G07040 464 / 6e-149
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.003G099100
172 / 2e-49
AT3G07040 482 / 8e-156
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.003G099000
125 / 4e-33
AT3G07040 479 / 2e-154
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134400
112 / 1e-28
AT3G07040 427 / 3e-134
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134425
91 / 5e-21
AT3G07040 347 / 6e-107
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134500
91 / 5e-21
AT3G07040 422 / 1e-132
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134475
61 / 1e-10
AT3G07040 413 / 1e-129
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014922
53 / 5e-08
AT3G07040 523 / 1e-171
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10038983
47 / 4e-06
AT3G07040 468 / 5e-150
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10027281
42 / 0.0003
AT3G07040 459 / 5e-147
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Potri.003G099200.1 pacid=42786923 polypeptide=Potri.003G099200.1.p locus=Potri.003G099200 ID=Potri.003G099200.1.v4.1 annot-version=v4.1
ATGTCTGAAGGGATAGTGACCTTCTTGCTCACGAAGCTTGGAGACTTCCTCCTAGAGAGGGGGAAACAATTGGAAACAGTGAAGAATGAAGCTGAGCATG
TTAGTGATGAACTTGCGTTCAATATGAAAGCCTTCCTGAGACTTGCAGATGCAATGGAGGAAAGTGATTCTGCACTCAAATTGCTGGTCAAGAAAGTAAG
AGATGTAGCTTATGACACAGAGGACGCACTTGATGAACTGATCAGGCTATGCCTTGCCAATGATAATGGGCATGGAGTTATTTCCTGCTTCAGGAAAATT
GTCTCTCTGTTAAGGATGCTAGAGCTCGGCGTCGAGTTGCTTCAAAAATTCAAGGCATCAAATCCAGAGTCATCAGTATATCAGAGTCACATCGGAGATA
CTGCAATAAAAATAACAAAATGGTTCATGGTTCAAGTTCCAACAGCATCAACTACGATGGAATGTCAAAGGGATGCCCTTCTACTTGAAGAAGCTGTTCA
AAAAGCAAAAAGCAGAGCGATTGAGCGGCTTCTTGGGACCAAATCTGGACGTGAAGTGGTTTCAGTGGTGGGCATGGGAGGATCGGGGAATACGACCTTG
GTTAAAAGGGTCTACGAGGATAAAAGTTAA
AA sequence
>Potri.003G099200.1 pacid=42786923 polypeptide=Potri.003G099200.1.p locus=Potri.003G099200 ID=Potri.003G099200.1.v4.1 annot-version=v4.1
MSEGIVTFLLTKLGDFLLERGKQLETVKNEAEHVSDELAFNMKAFLRLADAMEESDSALKLLVKKVRDVAYDTEDALDELIRLCLANDNGHGVISCFRKI
VSLLRMLELGVELLQKFKASNPESSVYQSHIGDTAIKITKWFMVQVPTASTTMECQRDALLLEEAVQKAKSRAIERLLGTKSGREVVSVVGMGGSGNTTL
VKRVYEDKS
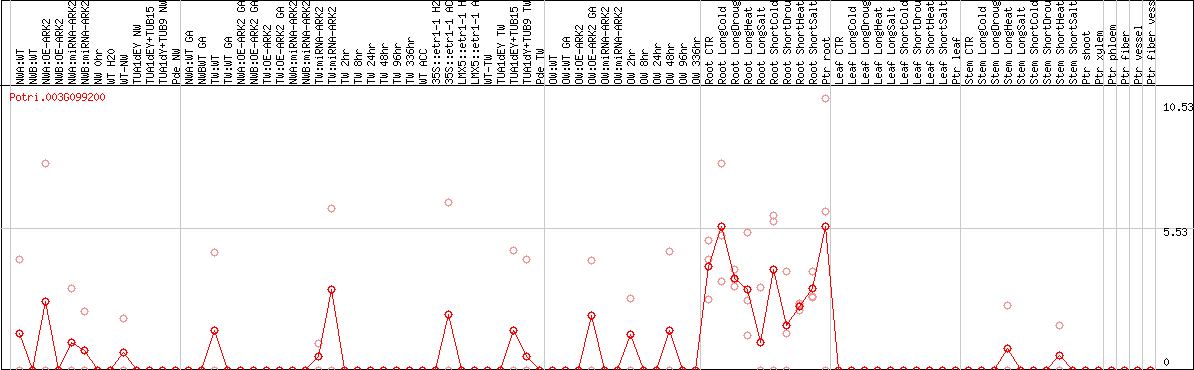
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G099200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.