External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G46410 67 / 2e-16
MYB
CPC
CAPRICE, Homeodomain-like superfamily protein (.1)
AT1G01380 67 / 2e-16
MYB
ETC1
ENHANCER OF TRY AND CPC 1, Homeodomain-like superfamily protein (.1)
AT5G53200 66 / 1e-15
MYB
TRY
TRIPTYCHON, Homeodomain-like superfamily protein (.1)
AT4G01060 62 / 2e-14
MYB
CPL3, ETC3
ENHANCER OF TRY AND CPC 3, CAPRICE-like MYB3 (.1.2.3)
AT5G14750 64 / 4e-14
MYB
WER1, WER, AtMYB66
WEREWOLF 1, WEREWOLF, myb domain protein 66 (.1)
AT5G35550 64 / 4e-14
MYB
ATTT2, TT2, AtMYB123
TRANSPARENT TESTA 2, MYB DOMAIN PROTEIN 123, Duplicated homeodomain-like superfamily protein (.1)
AT4G09460 64 / 6e-14
MYB
ATMYB6, ATMYB8
myb domain protein 6 (.1)
AT3G13540 64 / 8e-14
MYB
ATMYB5, ATM2
myb domain protein 5 (.1)
AT5G40330 63 / 1e-13
MYB
ATMYBRTF, ATMYB23
myb domain protein 23 (.1)
AT5G52600 62 / 1e-13
MYB
AtMYB82
myb domain protein 82 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018366
88 / 3e-24
AT2G46410 67 / 2e-15
CAPRICE, Homeodomain-like superfamily protein (.1)
Lus10007643
85 / 2e-23
AT2G46410 65 / 3e-15
CAPRICE, Homeodomain-like superfamily protein (.1)
Lus10018414
88 / 6e-22
AT4G05120 548 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Lus10001043
70 / 9e-17
AT1G22640 119 / 1e-33
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
Lus10005949
67 / 3e-16
AT4G01060 91 / 2e-25
ENHANCER OF TRY AND CPC 3, CAPRICE-like MYB3 (.1.2.3)
Lus10028513
68 / 5e-15
AT1G66370 205 / 3e-65
myb domain protein 113 (.1)
Lus10038822
64 / 5e-15
AT5G53200 96 / 1e-27
TRIPTYCHON, Homeodomain-like superfamily protein (.1)
Lus10039173
67 / 7e-15
AT3G13540 244 / 3e-81
myb domain protein 5 (.1)
Lus10033438
67 / 1e-14
AT1G22640 194 / 2e-61
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
Lus10014933
63 / 1e-14
AT2G46410 96 / 1e-27
CAPRICE, Homeodomain-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Potri.004G021300.1 pacid=42796429 polypeptide=Potri.004G021300.1.p locus=Potri.004G021300 ID=Potri.004G021300.1.v4.1 annot-version=v4.1
ATGGCTGACACTGAACATTCTTCTTCTGATGAAACTTCTGTGGATTCTAGAGAGGAAACAAGTCAGGAATCAAAGCTTGAATTCTCTGAGGATGAGGAGA
CGCTTATAATTAGGATGTTTAATCTAGTTGGAGAGAGGTGGTCTCTAATTGCTGGAAGGATTCCAGGAAGAACAGCTGAGGAAATAGAGAAGTATTGGAA
CACTAGATGCTCTACGAGTGAATGA
AA sequence
>Potri.004G021300.1 pacid=42796429 polypeptide=Potri.004G021300.1.p locus=Potri.004G021300 ID=Potri.004G021300.1.v4.1 annot-version=v4.1
MADTEHSSSDETSVDSREETSQESKLEFSEDEETLIIRMFNLVGERWSLIAGRIPGRTAEEIEKYWNTRCSTSE
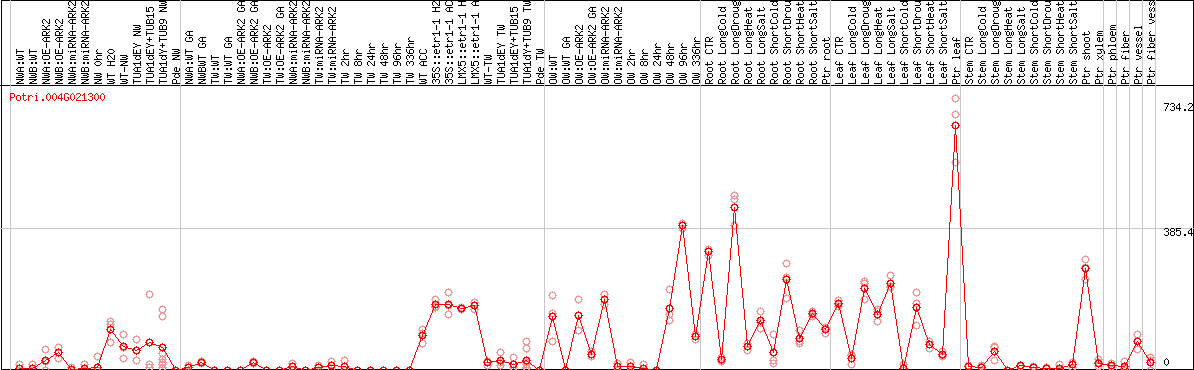
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G021300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.