External link
Symbol
Pt-APE2.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G46110 540 / 0
TPT, APE2
triose-phosphate ⁄ phosphate translocator, ACCLIMATION OF PHOTOSYNTHESIS TO ENVIRONMENT 2, Glucose-6-phosphate/phosphate translocator-related (.1.2.3.4)
AT5G54800 227 / 5e-72
ATGPT1, GPT1
ARABIDOPSIS GLUCOSE 6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 1, glucose 6-phosphate/phosphate translocator 1 (.1)
AT1G61800 221 / 7e-70
ATGPT2, GPT2
ARABIDOPSIS GLUCOSE-6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 2, glucose-6-phosphate/phosphate translocator 2 (.1)
AT5G17630 219 / 2e-68
Nucleotide/sugar transporter family protein (.1)
AT3G01550 177 / 5e-53
ATPPT2
phosphoenolpyruvate (pep)/phosphate translocator 2 (.1)
AT5G33320 166 / 4e-48
ARAPPT, CUE1
PHOSPHOENOLPYRUVATE/PHOSPHATE TRANSLOCATOR, CAB UNDEREXPRESSED 1, ARABIDOPSIS THALIANA PHOSPHATE/PHOSPHOENOLPYRUVATE TRANSLOCATOR, Glucose-6-phosphate/phosphate translocator-related (.1)
AT4G03950 123 / 3e-33
Nucleotide/sugar transporter family protein (.1)
AT1G77610 73 / 1e-14
EamA-like transporter family protein (.1)
AT1G21870 72 / 3e-14
GONST5
golgi nucleotide sugar transporter 5 (.1)
AT1G12500 66 / 4e-12
Nucleotide-sugar transporter family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G057800
550 / 0
AT5G46110 627 / 0.0
triose-phosphate ⁄ phosphate translocator, ACCLIMATION OF PHOTOSYNTHESIS TO ENVIRONMENT 2, Glucose-6-phosphate/phosphate translocator-related (.1.2.3.4)
Potri.008G095200
501 / 1e-179
AT5G46110 454 / 5e-159
triose-phosphate ⁄ phosphate translocator, ACCLIMATION OF PHOTOSYNTHESIS TO ENVIRONMENT 2, Glucose-6-phosphate/phosphate translocator-related (.1.2.3.4)
Potri.013G071900
233 / 1e-73
AT5G17630 526 / 0.0
Nucleotide/sugar transporter family protein (.1)
Potri.004G019900
224 / 1e-70
AT1G61800 592 / 0.0
ARABIDOPSIS GLUCOSE-6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 2, glucose-6-phosphate/phosphate translocator 2 (.1)
Potri.011G135900
220 / 2e-69
AT5G54800 583 / 0.0
ARABIDOPSIS GLUCOSE 6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 1, glucose 6-phosphate/phosphate translocator 1 (.1)
Potri.001G420200
218 / 1e-68
AT5G54800 563 / 0.0
ARABIDOPSIS GLUCOSE 6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 1, glucose 6-phosphate/phosphate translocator 1 (.1)
Potri.012G082100
187 / 2e-56
AT5G33320 458 / 1e-160
PHOSPHOENOLPYRUVATE/PHOSPHATE TRANSLOCATOR, CAB UNDEREXPRESSED 1, ARABIDOPSIS THALIANA PHOSPHATE/PHOSPHOENOLPYRUVATE TRANSLOCATOR, Glucose-6-phosphate/phosphate translocator-related (.1)
Potri.001G347300
183 / 9e-55
AT3G01550 411 / 1e-142
phosphoenolpyruvate (pep)/phosphate translocator 2 (.1)
Potri.015G077900
183 / 1e-54
AT5G33320 517 / 0.0
PHOSPHOENOLPYRUVATE/PHOSPHATE TRANSLOCATOR, CAB UNDEREXPRESSED 1, ARABIDOPSIS THALIANA PHOSPHATE/PHOSPHOENOLPYRUVATE TRANSLOCATOR, Glucose-6-phosphate/phosphate translocator-related (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013978
528 / 0
AT5G46110 641 / 0.0
triose-phosphate ⁄ phosphate translocator, ACCLIMATION OF PHOTOSYNTHESIS TO ENVIRONMENT 2, Glucose-6-phosphate/phosphate translocator-related (.1.2.3.4)
Lus10015399
528 / 0
AT5G46110 642 / 0.0
triose-phosphate ⁄ phosphate translocator, ACCLIMATION OF PHOTOSYNTHESIS TO ENVIRONMENT 2, Glucose-6-phosphate/phosphate translocator-related (.1.2.3.4)
Lus10037085
489 / 7e-175
AT5G46110 530 / 0.0
triose-phosphate ⁄ phosphate translocator, ACCLIMATION OF PHOTOSYNTHESIS TO ENVIRONMENT 2, Glucose-6-phosphate/phosphate translocator-related (.1.2.3.4)
Lus10007653
223 / 1e-70
AT1G61800 559 / 0.0
ARABIDOPSIS GLUCOSE-6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 2, glucose-6-phosphate/phosphate translocator 2 (.1)
Lus10004312
223 / 3e-70
AT5G17630 484 / 4e-171
Nucleotide/sugar transporter family protein (.1)
Lus10018356
222 / 4e-70
AT1G61800 557 / 0.0
ARABIDOPSIS GLUCOSE-6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 2, glucose-6-phosphate/phosphate translocator 2 (.1)
Lus10019209
221 / 1e-69
AT5G17630 480 / 2e-169
Nucleotide/sugar transporter family protein (.1)
Lus10011155
220 / 3e-69
AT5G54800 581 / 0.0
ARABIDOPSIS GLUCOSE 6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 1, glucose 6-phosphate/phosphate translocator 1 (.1)
Lus10043060
220 / 3e-69
AT5G54800 580 / 0.0
ARABIDOPSIS GLUCOSE 6-PHOSPHATE/PHOSPHATE TRANSLOCATOR 1, glucose 6-phosphate/phosphate translocator 1 (.1)
Lus10013083
185 / 1e-55
AT3G01550 429 / 1e-149
phosphoenolpyruvate (pep)/phosphate translocator 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0184
DMT
PF03151
TPT
Triose-phosphate Transporter family
Representative CDS sequence
>Potri.004G048900.4 pacid=42796132 polypeptide=Potri.004G048900.4.p locus=Potri.004G048900 ID=Potri.004G048900.4.v4.1 annot-version=v4.1
ATGTGGTACTTTTTGAACGTGATATTTAACATACTTAACAAGAAGATTTACAATTACTTCCCTTATCCATATTTTGTTTCGGTGATTCATTTGTTTGTTG
GAGTGGTGTACTGTTTGGTGAGCTGGACTGTGGGCCTTCCAAAGCGCGCTCCTATTGACTCAAATCTCCTGAAGCTGTTGATTCCCGTTGCTGTCTGTCA
TGCATTAGGCCATGTGACCAGTAATGTCTCCTTTGCGGCGGTTGCTGTCTCCTTTACACACACAATCAAAGCACTTGAGCCCTTCTTCAATGCTGCTGCT
TCTCAATTCGTATTGGGACAGTCAATACCGATAACTCTGTGGCTATCACTTCTGCCTGTCGTTCTTGGTGTGTCCATGGCATCATTGACTGAGCTCTCAT
TCAATTGGACGGGCTTTATTAGTGCTATGATTTCCAACATCTCCTTCACTTACAGGAGTCTCTACTCGAAGAAAGCAATGACTGATATGGACAGCACTAA
CATTTATGCTTACATTTCCATCATTGCACTCTTTGTCTGCATTCCACCTGCCATTCTTGTCGAGGGACCTCAACTGATCAAGCATGGCTTTAATGATGCA
ATTGCTAAAGTGGGCCTAACCAAGTTCATCTCAGACCTTTTTTGGGTTGGAATGTTTTATCACCTCTACAATCAGTTGGCTACCAATACCTTGGAGAGGG
TTGCACCTCTTACACATGCAGTGGGCAATGTGCTGAAACGTGTATTCGTGATTGGCTTTTCCATCCTGATCTTTGGTAACAAAATCTCAACACAAACTGG
TATTGGAACAGGAATTGCAATTGCTGGAGTGGCAACCTACTCTTACATTAAGGCCAAGATGGAAGAGGAGAAACGACGAGGGAAAGCAGCATGA
AA sequence
>Potri.004G048900.4 pacid=42796132 polypeptide=Potri.004G048900.4.p locus=Potri.004G048900 ID=Potri.004G048900.4.v4.1 annot-version=v4.1
MWYFLNVIFNILNKKIYNYFPYPYFVSVIHLFVGVVYCLVSWTVGLPKRAPIDSNLLKLLIPVAVCHALGHVTSNVSFAAVAVSFTHTIKALEPFFNAAA
SQFVLGQSIPITLWLSLLPVVLGVSMASLTELSFNWTGFISAMISNISFTYRSLYSKKAMTDMDSTNIYAYISIIALFVCIPPAILVEGPQLIKHGFNDA
IAKVGLTKFISDLFWVGMFYHLYNQLATNTLERVAPLTHAVGNVLKRVFVIGFSILIFGNKISTQTGIGTGIAIAGVATYSYIKAKMEEEKRRGKAA
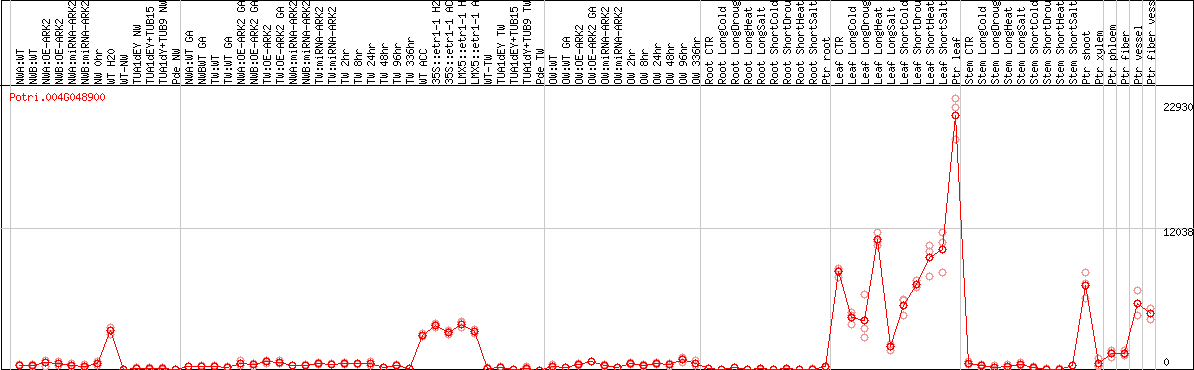
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G048900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.