Potri.004G170232 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G170232.2 pacid=42795080 polypeptide=Potri.004G170232.2.p locus=Potri.004G170232 ID=Potri.004G170232.2.v4.1 annot-version=v4.1
ATGGAGGCTTTTCTTACGTTTCCCATGGAGGTGACTTTGACAAGGGTCAGTTCCATAGCTGCTGAAGGGATCAGACTTGCCTGGGGATTGGAGGGACAGC
TGCAAAAGCTCGAAGAGTCCTTGACCATGATCAAAGCTGTGCTCAAAGATGCAGCCAGAAGGCCCGTAACAGACGATTTTGCGAAGCTTTGGCTGGAGAA
GCTACAAGATGTAGCTTACGATGCTGAAGATGTTCTAGACGAGTTTGCTTATGAGAATCTCCGAAAAGACCAAAAGAAGAGAAACGTACGTAACTTTTTT
TCATCCCACAACCCTACCAAATTCCGTTGGAATATGGGTCAAAAAGTTCAGAAGATCAATGAAGCCCTGGATGATATTCAGAAACTTGCAACTCGCTTTG
GGTCTGGGCTGGGAGTTGCATCTCAACATGTAGAGAGTGGTCCTGAAGTTATCCGGGCTAAAGACCGAAAGACAGATTCCTTGCTTGGAAGCTCAGAAGT
TGTAGTAGGAAGGGAGGATGATGTCTCTAAAGTTATGAAATTGCTGATTGGCTCGATTGATCAACAAGTTCTGTCTGTTGTCCCTATAGTGGGGATGGGT
GGCCTCGGAAAGACTACTATAGCAAAAAAAGTTTATAAAGTAGCGAGGGAGAAAAAGCTTTTTGATGTAACAATATGGGTTTGTGTTTCTAATGAATTTA
GTAAACAGAGGATTTTAGGAGAGATGTTGCAAGGTGTTGGTGGACCTATGTTGAGCAATCTAAATGAAATTATGGAAAGACTTAAAGAAAAGCTGGAAAA
GAAGACATTTTTTCTTGTACTTGATGATGTGTGGGAAGGCCATGATAAGTGGAATGATTTGAAGGAGCAACTGTTAAAAATCAACAATAAGAAGGGGAAT
GCTGTTGTTGTAACAACGCGTATTAAGGAAGTAGCGGACACGATGGAGACTTCTCCAGGGATTCAGCACGAGCCAGGGAGACTTTCAGATGATCAATGTT
GGTCCATTATCAAGCAAGAGGTGAGCAGGGGTGGACGAGAAACAATAGCTTCAGACTTGGAGTCTATTGGAAAAGATATTGCAAAGAAATGTGGAGGGAT
TCCATTACTTGCTAAAGTTTTGGGAGGAACTCTTCACGGAAAGCAGGTACAGGAGTGGCAGTCAATTCTAAACAGTAGAATTTGGGATTCTCAGGATGGA
AATAAAGTTTTGCTCATATTGAGACTCAGTTTTGATCACCTGTCATCGCCATCACTGAAAACATGTTTTGCATACTGTTCCATTTTCCCTAAAGATTTTA
AAATTGAAAGGGAAGAGCTGGTTCAACTTTGGATGGCCGAAGGTTTTCTTGGGACATCAAATGGGAGGACGGAAGACGAAGGCAACAAGTGTTTCAATGA
CTTGCTTGCAAATTCCTTTTTCCAAGATGTTGAAAGGAATGAGTGTGAGATCGTCACAAGTTGCAAGATGCATGACCTTCTGCATGATCTTGCATTGCAG
GTCTCAAAATCAAAAGTGTTAAATCTAGAGGCAGATTCGGCTGTTGATGGCGCATCTCATATTCGTCATCTAAATCTCATATCTTGTGGGGATGTTGAGG
CAGCATTTCCAAGGGGTGATGCTAGAAAATTGCGAACTGTTTTCTCAATGGTTGATGTATTCAATGGGTCTTTGAAATTCAAAAGCTTGAGAACTCTCAA
ATTGCAACGGTCTAATATCACAGAGTTACCAGATTCAATTTGGAAGCTGAGACATTTGAGATACCTTGATGTCTCACGTACCAGTATCAGAGTATTGCCA
GAATCCATCACAAAGCTCTACCATTTGCAAACATTAAGATTCACTGATTGCAAGTCACTAGAAAAGCTTCCCAAGAAAATGAGAAATTTAGTCAGCTTGA
GACATCTCCATTTTGATGATCCAAAGCTAGTGCCAGCTGAGGTGAGACTCTTAACACGCCTTCAAACTCTACCTTTATTTGTTGTGGGTCCAGATCATAT
GGTTGAAGAGCTGGGATGTTTGAATGAACTAAGAGGAGCATTGGAGATATGCAAGCTTGAGCAAGTGAGAGACAAAGAAGAAGCTGAGAAGGCAAAACTG
CGTGGAAAAAGAATTAACAAATTGGTGTTTGAATGGAGTTATGATGAAGGTAACAACAGTGTCAATAGCGAGGATGTGCTGGAAGGCCTGCAGCCTCACC
CAGACTTAAGAAGCTTGACAATTGAGGGTTATGGAGGTGGATATTTCTCATCATGGATATTGCAACTCAATAATCTGACGGTGCTGAGATTGAATGGCTG
CAGCAAGTTGAGGCAACTCCCAACACTTGGATGTCTTCCTCGCCTTAAGATTCTAAAGATGAGTGGAATGCCTAATGTGAAATGTATAGGCAAGGAGTTC
TATAGTAGCAGCATTGGCAGTGCAGCAGAACTGTTTCCAGCACTAGAAGAACTCACTTTGCGGGGTATGGATGGTCTTGAAGAATGGATGGTACCAGGTG
GAGAAGGTGATCTAGTATTTCCTTGTCTTGAAGAGTTGTGCATTGAGGAGTGTCGCCAGTTGAGGCAACTCCCAACACTTGGATGTCTTCCTCGCCTTAA
GATTCTAAAGATGAGTGGAATGCCTAATGTGAAATGTATAGGCAAGGAGTTCTATAGTAGCAGCATTGGCAGTGCAGCAGAACTGTTTCCAGCACTAGAA
GAACTCACTTTGCGGGGTATGGATGGTCTTGAAGAATGGATGGTACCAGGTGGAGAAGTTGTTGCAGTATTTCCTCGTCTTGAAAAGTTGAGCATTTGGC
AGTGTGGAAAGTTGGAAAGCATTCCGAGATGTCGTCTTTCATCTCTTGTAGAATTTGAAATTCACGGATGTGATGAGCTGAGATATTTTTCTGGTGAGTT
TGATGGCTTCAAGTCTCTTCAGATTTTAAGGATATTGAAGTGTCCAATGCTGGCATCCATTCCAAGCGTACAACACTGCACAGCTCTGGTGCAATTGCGT
ATATACGACTGCCGTGAGTTGATCTCAATTCCTGGTGATTTTCGAGAATTGAAATATTCTTTGAAGACATTGAGTGTTAATGGGTGTAAATTGGGAGCTC
TTCCAAGCGGACTACAATGTTGCGCATCTCTAGAGGAATTAACAGTAATTGATTGTAGTGAGCTTATCCATTTCAGTGGTTTACAAGAATTGTCTTCGCT
ACGAAGTTTAGGGATTATACGTTGTGATAAGCTCATCAGTATTGATTGGCATGGTTTACGACAATTGTCTTCTCTTGTTTATTTACAAATCATCACGTGT
CCCAGTTTGAGAGATATCCCAGAGGATGATTGCTTAGGCGGCCTCATGCAACTGCATTATTTGAGCATCGGTGGTTTCTCAAAGGAGATGGAGGCTTTTC
CTGCAGGAGTTTTAAACTCATTCCAACACCTCAACTTGAGTGGGTCCCTCAAATACCTAATGATTGATGGATGGGATAAACTGAAGAGTGTACCACACCA
ACTTCAACACCTCACTGCCCTCGAGAGATTGAAAATACGTTATTTCAATGGAGAGGAGTTTGAGGAAGCATTGCCAGAATGGTTGGCCAACCTTTCTTCT
CTTCAATGTCTTTCGATCGAGGATTGCAAGAATCTCAAGTATATGCCAAGTTCAACAGCCGCCATTCAACGCCTCTCCAAATTAGAGCTATTGTATATCT
GGTACTGTCCACATCTATCAGAAAATTGCAGAGAGGAGAATGGCTCTGAGTGGCCCAAGATCTCTCATATCCCAAAAATCTATATAAGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G170232.2 pacid=42795080 polypeptide=Potri.004G170232.2.p locus=Potri.004G170232 ID=Potri.004G170232.2.v4.1 annot-version=v4.1
MEAFLTFPMEVTLTRVSSIAAEGIRLAWGLEGQLQKLEESLTMIKAVLKDAARRPVTDDFAKLWLEKLQDVAYDAEDVLDEFAYENLRKDQKKRNVRNFF
SSHNPTKFRWNMGQKVQKINEALDDIQKLATRFGSGLGVASQHVESGPEVIRAKDRKTDSLLGSSEVVVGREDDVSKVMKLLIGSIDQQVLSVVPIVGMG
GLGKTTIAKKVYKVAREKKLFDVTIWVCVSNEFSKQRILGEMLQGVGGPMLSNLNEIMERLKEKLEKKTFFLVLDDVWEGHDKWNDLKEQLLKINNKKGN
AVVVTTRIKEVADTMETSPGIQHEPGRLSDDQCWSIIKQEVSRGGRETIASDLESIGKDIAKKCGGIPLLAKVLGGTLHGKQVQEWQSILNSRIWDSQDG
NKVLLILRLSFDHLSSPSLKTCFAYCSIFPKDFKIEREELVQLWMAEGFLGTSNGRTEDEGNKCFNDLLANSFFQDVERNECEIVTSCKMHDLLHDLALQ
VSKSKVLNLEADSAVDGASHIRHLNLISCGDVEAAFPRGDARKLRTVFSMVDVFNGSLKFKSLRTLKLQRSNITELPDSIWKLRHLRYLDVSRTSIRVLP
ESITKLYHLQTLRFTDCKSLEKLPKKMRNLVSLRHLHFDDPKLVPAEVRLLTRLQTLPLFVVGPDHMVEELGCLNELRGALEICKLEQVRDKEEAEKAKL
RGKRINKLVFEWSYDEGNNSVNSEDVLEGLQPHPDLRSLTIEGYGGGYFSSWILQLNNLTVLRLNGCSKLRQLPTLGCLPRLKILKMSGMPNVKCIGKEF
YSSSIGSAAELFPALEELTLRGMDGLEEWMVPGGEGDLVFPCLEELCIEECRQLRQLPTLGCLPRLKILKMSGMPNVKCIGKEFYSSSIGSAAELFPALE
ELTLRGMDGLEEWMVPGGEVVAVFPRLEKLSIWQCGKLESIPRCRLSSLVEFEIHGCDELRYFSGEFDGFKSLQILRILKCPMLASIPSVQHCTALVQLR
IYDCRELISIPGDFRELKYSLKTLSVNGCKLGALPSGLQCCASLEELTVIDCSELIHFSGLQELSSLRSLGIIRCDKLISIDWHGLRQLSSLVYLQIITC
PSLRDIPEDDCLGGLMQLHYLSIGGFSKEMEAFPAGVLNSFQHLNLSGSLKYLMIDGWDKLKSVPHQLQHLTALERLKIRYFNGEEFEEALPEWLANLSS
LQCLSIEDCKNLKYMPSSTAAIQRLSKLELLYIWYCPHLSENCREENGSEWPKISHIPKIYIR
|
DESeq2's median of ratios [POPLAR]
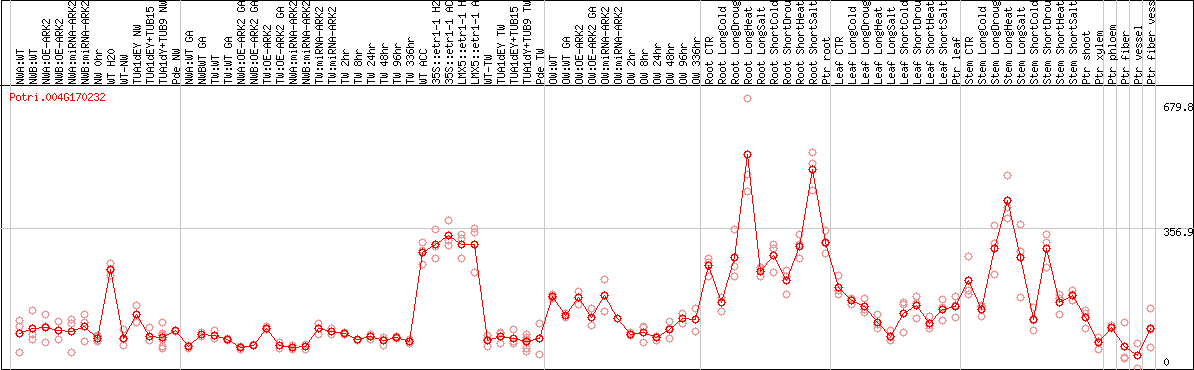
Coexpressed genes
Potri.004G170232 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
