Potri.004G170400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G170400.1 pacid=42795907 polypeptide=Potri.004G170400.1.p locus=Potri.004G170400 ID=Potri.004G170400.1.v4.1 annot-version=v4.1
ATGTTTATTTCACAACTTTCAGATCATACTTCTCTCTTTTCATCAAATACTTCTTTGATTTCCCAAAGAAAACAAGAGAATATGGCAGCAGAGCTTTTTC
TTACGTTTGCCATGCAGGAGACCTTGACAAGGGTGAGTTCCATAGCTACTGAAGGGATCAGACTTGCTTGGGGATTGAAGGGCCAGCTGCAAAGGCTCAA
CAAGCCGTTGACCATGATCCAAGCTGTGCTGCGGGACGCAGCCAAAAGGCCAGAAACAGACGATTCAGTGAAGCTTTGGCTGGAGAGGCTACAAGATGTA
GCTTATGATGCTGAAGATGTTCTAGACGAGTTTGCTTATGAGATTCTCCGAAAAGACCAAAAGAAGGGAAAGGTACGTGACTGCTTTTCACTCCACAACC
CTGTTGCATTCCATTTGAATATGGGTCAAAAAGTTAAGAAGATCAATGAAGCCCTGGATGAAATCTGGAAAGATGCAGCCGGATTTGGGCTTGGATTGAC
ATCTCTACCTGTAGATAGAGCTCAAGAAGTTAGCTGGGATCCAGATCGAGAGACAGATTCGTTCCTTGACAGCTCAGAAGTTGTAGGAAGGGAGGATGAT
GTCTCTAAAGTTATGGAATTGCTGACTAGCTTGACCAAACACCAACATGTCCTTCTGGTTGTCCCTATTATGGGGATGGCTGGCCTTGGAAAGACTACTG
TAGCAAAAAAAGTTTGTGAAGTAGTGAGGGAGAAAAAACATTTTGATGTGACTCTATGGGTTTGCGTTTCTAATGATTTTAATAATGTAAAGATTTTAGC
AGCAATGTTGCAAATGATTGACAAAACTACGGGTGGGTTGAACAGTCTTGATGCAATTCTTCAAAACCTTAAGAAAGAGCTGGAAAATAAGACATTTTTT
CTTGTTCTTGATGACGTGTGGAATGAAGACCAAGATAAGTGGGATGATTTGAAGGAGCAACTGTTAAAAATTAAAAGCAAGAATGGGAATGCTGTTGTTG
TTACAACCCGCAGTAAGAAAGTAGCAGGCATGATGGAGACTTCTCCTGGGATTCAGCACGAGCCAGGGAGACTATCAGCTGATCAATGTTGGTCCATTAT
CAAGCAAAAGGTGAGCAGGGGTGGACAAGAAACAATACCTTCAGACTTGGAGACTATTGGAAAAGAGATTGCGAAGAAATGTGGAGGGATTCCGTTGCTT
GCTAAAGTTTTGGGAGGAACATTGCGCCAAAAGGAGACGCAGGAGTGGCAGTCAATTCTAAACAGTAGAATTTGGGATTCTCAGGATGGAAATAAAGCTT
TGCGCATATTGAGATTAAGTTTTGATTACTTGTCATCGCCAACATTGAAAAAATGTTTTGCATACTGTTCCATTTTCCCTAAAGATTTTGAAATTGAAAA
GGAAGAGCTGGTTCAACTTTGGATGGCTGAAGGTTTTCTCAGGCCATCAAATAGGAGGATGGAAGACGAAAGCAACGAGTATTTCAATGACTTGCTTGCA
AATTCCTTTTTCCAAGATGTTGAAAGGAATGGGTATGAGATCGTAACAAGATGCAAGATGCATGATCTTGTGCATGATCTTGCATTGCAGGTCTCAAAAT
CAGAAACGTTAAATCTGGAGGCGGGTTCGGCTGTTGATGGCGCATCTCATATTCGTCATCTAAATATCGTATCTTGTGGGGATGTTGAGGCAGCATTAAC
AGTGATTGATGCTAGAAAATTGCGAACTGTTTTCTCAATGGTTGATGTATTCAATGGGTCTTGGAAATTCAAAAGCTTAAGAACTCTCAAATTGCGACGG
TCTAATATCACAAAGTTACCAGATTCAATTTGCAAGCTGAGACAATTGAGATATCTTGATGTCTCGGATACTGCTATAAGAGTATTGCCAGAATCCATCA
CCAAGCTCTACCATTTGGAAACATTAAGATTCACTGATTGCAAGTCACTAGAAAAGCTTCCCAAGAAAATGAGGAAATTAGTCAGCTTGAGACATCTTCA
TTTTGATGATCCAAAGCTAGTGCCAGCTGAGGTGAGACTCTTAACACGCCTTCAAACTCTGCCTTTTTTTGTTGTGGGTCCAAATCATATGGTTGAAGAG
CTGGGATGTTTGAATGAACTAAGAGGAGCATTGAAGATATGCAAGCTTGAGCAAGTTAGAGACAGGGAAGAAGCTGAGAAGGCAAAACTGCATGAAAAAA
GAATGAGCAAATTGGTGCTTGAATGGAGCCTTAACAGCAATGTTAACAACGAGTATGTGTTAGAAGGCTTGCAGCCTCACCCAGACATAAGAAGCTTGAC
AATTGAGGGTTATGGAGGTGAAGATTTCTCATCATGGATGTCTACATTCCTACTCAATAATCTGATGGAGCTGAGTTTGAAAGACTGCAGCAAGTGCAGA
CAACTCCCAACACTTGGATGTCTTCCTCGCCTTAGGATTCTAGAGATGAGTGGAATGCCTAATGTGAAATGTATAGGCAATGAGTTCTATAGTAGCAGTG
GCCGTGCAGCAGTACTGTTTCCAGCACTAAAAGAACTCACTCTATCCAGTATGGAAGGTCTTGAAGAATGGATGGTACCAGGTGGAGAAGGTGATCAAGT
ATTTCCTTGTCTTGAAAAGTTGAGCATTGAGAGGTGTGGCAAGTTGAAAAGCATTCCGATATGCCGTCTTTCATCTCTTGTACAATTTAAAATTGAAAGA
TGTGAGGAACTGGGATATTTATGTGGTGAGTTTCATGGCTTCGTGTCTCTTCAATTTTTTAGCGTAACATATTGTCCAAAGATGGCATCCATTCCAAGCG
TACAACATTGCACAGCTTTGGTGGAATTGAGTATATGTTGGTGCCCTGAGTTGATCTCAATTCCTGGTGATTTCCGAGAATTGAAATATTCTTTGAAGAA
ATTGGGTATTTGGGGATGTAAATTGGGAGCTCTTCCAAGCGGATTAGAATGTTGCGCATCTCTAGAGGAATTGCGGATATGGAAATGCAGTGAGCTTATC
CATATCAGTGATTTACTAGAATTGTCTTCGCTTCGAAGTTTAGAGATTAGAGGTTGTGATAAGCTCATCAGTATTGATTGGCATGGTTTACGACAATTGC
CTTCTCTTGTTTATTTAGGAATCATCGGGTGTCCCAGTTTGAGTGATATCCCAGAGGATGATTGGTTAGGCGGCCTCACCCAACTGAAGGTATTGAGCAT
CGGTGGTTTCACAGAGGAGCTGGAGGCTTTTCCTTCAGGAGTTTTAAATTCATTCCAACACCTCAACTTGAGCGGGTCCCTTGAATCCTTAAGGATATGT
GGATGGGATAAACTGAAGAGTGTACCGCACCAACTTCAACACCTCACTGCCCTCAAGTCATTGTGGATATATGATTTCAAGGGCGAGGGATTTGAGGAAG
CTTTACCAGATTGGTTGGCCAACCTTTCTTCTCTTCAATCTCTCACTATTTGGAATTGCTACAATCTCAAGTATCTGCCAAGTTCAACAGCCATTCAAGG
CCTCTCCAAATTAAACGAATTGGAGATTTATGGATGTTCATTTCTATCAGAAAATTGTAGAAAGGAGAATGGCTCTGAGTGGCCCAAGATTTCTCATATC
CCATCAATCATAATAAGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G170400.1 pacid=42795907 polypeptide=Potri.004G170400.1.p locus=Potri.004G170400 ID=Potri.004G170400.1.v4.1 annot-version=v4.1
MFISQLSDHTSLFSSNTSLISQRKQENMAAELFLTFAMQETLTRVSSIATEGIRLAWGLKGQLQRLNKPLTMIQAVLRDAAKRPETDDSVKLWLERLQDV
AYDAEDVLDEFAYEILRKDQKKGKVRDCFSLHNPVAFHLNMGQKVKKINEALDEIWKDAAGFGLGLTSLPVDRAQEVSWDPDRETDSFLDSSEVVGREDD
VSKVMELLTSLTKHQHVLLVVPIMGMAGLGKTTVAKKVCEVVREKKHFDVTLWVCVSNDFNNVKILAAMLQMIDKTTGGLNSLDAILQNLKKELENKTFF
LVLDDVWNEDQDKWDDLKEQLLKIKSKNGNAVVVTTRSKKVAGMMETSPGIQHEPGRLSADQCWSIIKQKVSRGGQETIPSDLETIGKEIAKKCGGIPLL
AKVLGGTLRQKETQEWQSILNSRIWDSQDGNKALRILRLSFDYLSSPTLKKCFAYCSIFPKDFEIEKEELVQLWMAEGFLRPSNRRMEDESNEYFNDLLA
NSFFQDVERNGYEIVTRCKMHDLVHDLALQVSKSETLNLEAGSAVDGASHIRHLNIVSCGDVEAALTVIDARKLRTVFSMVDVFNGSWKFKSLRTLKLRR
SNITKLPDSICKLRQLRYLDVSDTAIRVLPESITKLYHLETLRFTDCKSLEKLPKKMRKLVSLRHLHFDDPKLVPAEVRLLTRLQTLPFFVVGPNHMVEE
LGCLNELRGALKICKLEQVRDREEAEKAKLHEKRMSKLVLEWSLNSNVNNEYVLEGLQPHPDIRSLTIEGYGGEDFSSWMSTFLLNNLMELSLKDCSKCR
QLPTLGCLPRLRILEMSGMPNVKCIGNEFYSSSGRAAVLFPALKELTLSSMEGLEEWMVPGGEGDQVFPCLEKLSIERCGKLKSIPICRLSSLVQFKIER
CEELGYLCGEFHGFVSLQFFSVTYCPKMASIPSVQHCTALVELSICWCPELISIPGDFRELKYSLKKLGIWGCKLGALPSGLECCASLEELRIWKCSELI
HISDLLELSSLRSLEIRGCDKLISIDWHGLRQLPSLVYLGIIGCPSLSDIPEDDWLGGLTQLKVLSIGGFTEELEAFPSGVLNSFQHLNLSGSLESLRIC
GWDKLKSVPHQLQHLTALKSLWIYDFKGEGFEEALPDWLANLSSLQSLTIWNCYNLKYLPSSTAIQGLSKLNELEIYGCSFLSENCRKENGSEWPKISHI
PSIIIR
|
DESeq2's median of ratios [POPLAR]
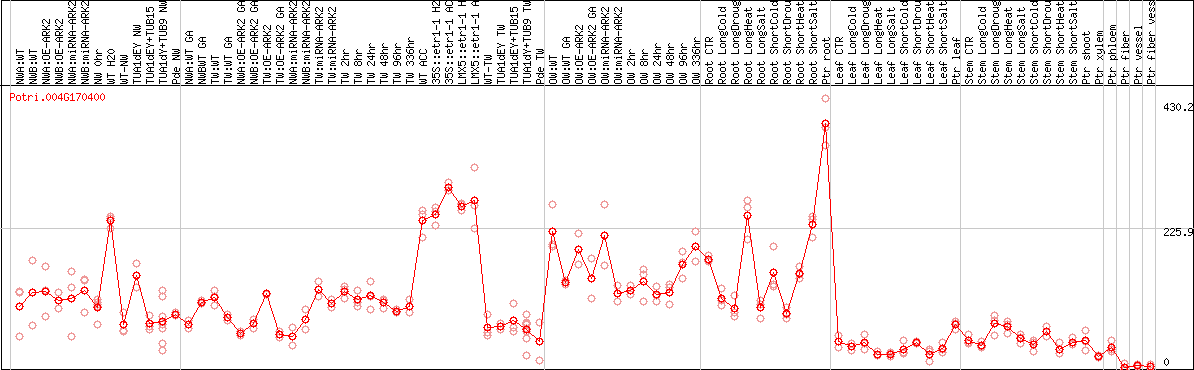
Coexpressed genes
Potri.004G170400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
