Potri.004G196100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G196100.1 pacid=42794470 polypeptide=Potri.004G196100.1.p locus=Potri.004G196100 ID=Potri.004G196100.1.v4.1 annot-version=v4.1
ATGGCAATTGGTGAGATTTTTCTTGCTGCATTCCTTGGAATGCTGTTTACAAGATTGACATCTCCTGAGTTCTTGAAGTTTGCACGTCGAGAGGGCATTT
GGAAGAAGGCAGAGAAGTGGAGAGGGATGTTATTGAAGGTCCAGGAGGTACTTGATGACGCGGAAGAAAAGCAGCTGACGGAAAAGGCTGTGAAGATATG
GCTGGATGATCTAAGAGACTTGGCATATGATGTGGAGGACTTGCTTGATGAATTTGCTACTGAATCTTTACGAAGGGAGTTGATGGCAGCAGAAGAGGCT
AGCACAAGCAAGGTACGGAGGATAGTGTCTACTACCTTATCATTTACCAAAATCAGTGCAAGTGCTATTAAGTTCAATCCCAAGATGAGGTCAAAGATGA
AAGAGGTGAGTAGCCGGTTAGATGGAATGGCGAAGCAGAGAATTGAACTTGGTTTAGAAAAGATGAGTGGAGGTAGAAGGACATCAACTGATGTATGGCA
GAAACCACCTAGTGCCTCTGTCCCAAATGAACCTGTCATCTATGGCAGGGATGGAGATAAGAAGAAGGTAATTGATTTGCTGTTGACAGAAGAGGCAAAT
CACGGTGATACTAATTTTCATGTCGTTCCAATTGTCGGTATGGGTGGTATCGGCAAAACAACTCTTGCACAGCATGTGTTTCAGGATGAATTGGTCAAGG
AATGGTTCAGTACAAAAGCATGGGCTTGTGTGTCGGATGACTTTGATGTTATGAGAATATCAAAGGCGATTCTCGAGTCAGTCACTCCGCATCCCTGTGA
TTTCAAGGAGTACAATCAAGTTCAGGTCAAATTGAGGGAGGCGCTAGCCGGAAAGAAGTTTCTTCTTGTTTTAGATGATGTATGGAACAAGAATTATGGT
CTCTGGGTAGCGTTGAAAACTCCCTTTGCAGCCGGAGCACCAGGAAGTAAGATAATTCTGACCACTCGTGATGCAGACGTTGCACTAATGGTTGGACCGA
CTGAATACCATTGCCTAAAGCCACTATCAGACCAAGATTGCTGGTCTGTGTTTGTGAAGCATGCATTCGAAAATAGAGATCTCGGCGCACAAACAAACCT
CCAATCAGTTTGTGAGAGAATCGTTACCAAGTGTAAGGGTTTGCCTTTGGCAGCCAGAACTCTTGGTGGCCTCTTACGCACTAAGCAAAGGGAGGATGAG
TGGGAAGACATACTGAATAGTAAGATATGGGACTTGTCAGACTCCCAAAGTGACATTCTCCCTGTTTTAAGATTGAGCTACTACCATCTCCCTTCACATT
TGAAGAGGTGTTTTACATATTCTGCTCTAATTCCGAAGGATTTTGAATTTGAAGAGAAGGATCTTGTCCTTCTATGGATGGCAGAAGGTTTGGTTCCACA
ACAAGTACAGAACAAGCAAATGGAAGATATGGGTGCGGAGTATTTTCGTGATCTAGTGTCGAGGTCAATTTTTCAAGTAGCAAACTGTGATGAATCTCGA
TTTGTGATGCATGATCTCGTGAGTGATCTGGCTCAGTGGGCTGCGGGAGACACATGTTTTCAATTGGGTAATGATTTGAATGCCATCAAGCAGTTCAAAG
TATCCAAGAGAGCTCGCCACTCCTCCTATATCCGTGGCTGGGATGGTATCAGAAAGTTTGAAGTATTCCATACTACTAAACGTTTACGAACTTTCCTACC
ACTGCCATCTCTTTTAGGTCATAATACAGGCTACTTAACTAGTCATGTCCCTTTTGATTTGTTGCCAGAATTAGAATTTCTGAGGGTGCTGTCTTTAAGT
GGTTACTGCATTGATACGCTACCAAATTCAATTGGAGATCTGAAACATTTAAGGTTTCTTAATCTTTCCTTCTCGGCAATCAGAAATTTACCTCAATCAG
TATGTTCTCTATACAACTTGCAGACTTTGTTACTGAAAGGATGTTGTCTTCTTGAAGGGCTGCCTTCAAAACTGGGGAGTCTCATCAACCTCCGGCATCT
TGATATCACAAGTGCAAGTTCAATCAAAGCGATGCCAATGGGAATTGAAAAGCTGACAAATCTGCAGACACTTTCCAATTTTGTCCTGGGCAAAGATAAG
GGCTCTAGGTTGAGTTCCTTGGTGAATTTGAAATCTCTTCGAGGAACGCTTTGTATTACAGGCTTGGAGAATGTGATTGATGCTCGGGAAGCAATGGAGG
CCAACATAAAAGATATAAATAACCTTGAAGTTTTGTTGCTGGAATGGAGTCCCAGAACAGATAATTCACGAAATGAGAAGGTTGACAAAGATGTGCTCGA
TGATCTTAGACCTCATGGAAAAGTCAAAGAACTCACCATCAACTGCTATGCTGGCCTAACATTTCCAACCTGGGTAGGAAATCCATCATTCTCCAGCATA
TTTCTCCTAAGGTTAGAGAACTGTACAAAATGCACTTCCTTGCCACCTCTTGGACTGCTGCCTTCTTTAAAAAATCTCAGCATAGTAAGTTTAACTGCAG
TAAAAAAAGTAGGTCCTGAATTTTATGGACAAGGTTGCTCAAAGCCTTTTCCAGTTTTAGAGACTTTACTTTTCAAGAACATGCAGGAATGGGAGGAATG
GATTCCTTGTGGGCTTGGTTCTGACGAATTCCCTCTCTTGAATAAGCTGTCTGTTAAAAGCTGTCCCAATCTGTGCAAAAAATTACCTCCAAGTGTTCCA
TCACTAGAAAAACTTGTGATTAAAAAATGCGAGAAGTTGGTGGTATTAATTCATAGTCTTCCAAAGCTGTGCAAAATGGTAATCAATGGCTGCAAAGAAG
TGGTGTACGAAGGTGGCGTTTATCTCAGATCTCTCAACTCCATGACAATTTCCAACATTTCAAAGCTCACTTACCTAGCAGAAGGATTCATCCAACCATT
GGCAGAAGTACAGGAATTGGAGATTGCTAATTGCATGGAATTGGCATCTTTGTACGAAAATGGAGTGGCATTGGCAAAACAGCTCACTTCCCTTCTTAAG
TTGGAGGTTAGAAACTGTCCCCAAGTTGTTTCTTTGATGGAAGGAGAAGTACCTGTGTATATGCAACAGCAGCTGGCTAACTGCAAGCTTGAATCTTTGA
CATTTAGTACATGTGAGAGCCTCAAGAAGTTACCACAATGGGTTCACAGCCTAGTGTCCCTTAAAGAGCTGAAAATTCAATACTGTCCGAGACTCCTTTC
GTTTCCCGAGGCTGGTTTGCCCTCCACACTGAGAATCATTGAGATTGTTGGCTGCAACGCTTTAACCCCCTTACCTGCTGCAGTGACTTACAATATGATG
TGTCTTGAGCAGTTGCGCATTGAAAATTGCGAGTCTTTAATATCCTTTGGAAGAATCCAGCTACCTCCAACTTTAAAAAAGTTGGAGATAAGATATTGCG
AGAATCTGTTATGCTTACTGGATGATGGGGAGGGATCCTCGTCAAAGAAATCTGATGAGAATACCAGTTGCAGCGGGAACAACTCATCTCTTCTTGAGTA
CTTGTATGTTGGAATATGCAACTCTCTCACTTCCATAGGTGAGTTACCTTCAGCTCTTAAGTACCTCCAGGTTTGTTCATGTTCAAAGCTCAAATCCTTA
TCATCCAGAGACAAACTACCGGCAGGGCTTAAACACCTCGCTATTGACTCGTGTGAAAATTTGGAATCAATGCCGGACAGGTTTCAGGATAACATGTCCC
TTGAAAATTTGAAAATATGGTTCTGCTTTAACCTTAGATCTTTGCCCGAAGGCCTACATAAATTATGCCATCTTCGTGAAATTTCAATATGGTACTGTCC
AGCTCTTGTTTCCTTTGCCGCTGAGGGGTTGCCCATAAACCTAAGGAGGCTCTTCATCATCAAATGTGATGGACTCAAGGCTATCCCTGATCATATGCAC
AATCTCATGTCTCTTGAAGAACTGTCAATTTACTACTGTCCAGACATTGTATCATTTCCAGAAGAGGGTTTTCCTACTAGCCTAACATATCTTGCTACCG
TAGATCTTAAGATCTGTGAGCTACTGTTTAACTGGGGCATGCACAAACTCTCCGCTCTTAGAACCCTCATCATTCAAGGTGGATTCTCTCATATATCCTT
TCCATCTGTGGACATGGGAGTGAGGTTGCCTTCTGCTCTAAACAGACTAAGCATTGAAGATTTTCCGAATCTTGAGTACCTATCCTACAGCGGCTTTCAA
AATCTCTCCTCTCTTGAACGCCTAAGCATCAGTGACTGCCCCAAGCTCACATCATTTCCAGGGAAGGGCTTGCCTTCCTCACTTTTAGAACTACGTATTC
GAGCATGTCCCCTTTTAGTACAACAGATAAAGGGAAGAGTAAAAGAATGGCTCAAGATAAGGCACATTCCATACATTAACATAGACGGAAAAGTAGTCTC
TGACCCAGCCACCCAGTTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G196100.1 pacid=42794470 polypeptide=Potri.004G196100.1.p locus=Potri.004G196100 ID=Potri.004G196100.1.v4.1 annot-version=v4.1
MAIGEIFLAAFLGMLFTRLTSPEFLKFARREGIWKKAEKWRGMLLKVQEVLDDAEEKQLTEKAVKIWLDDLRDLAYDVEDLLDEFATESLRRELMAAEEA
STSKVRRIVSTTLSFTKISASAIKFNPKMRSKMKEVSSRLDGMAKQRIELGLEKMSGGRRTSTDVWQKPPSASVPNEPVIYGRDGDKKKVIDLLLTEEAN
HGDTNFHVVPIVGMGGIGKTTLAQHVFQDELVKEWFSTKAWACVSDDFDVMRISKAILESVTPHPCDFKEYNQVQVKLREALAGKKFLLVLDDVWNKNYG
LWVALKTPFAAGAPGSKIILTTRDADVALMVGPTEYHCLKPLSDQDCWSVFVKHAFENRDLGAQTNLQSVCERIVTKCKGLPLAARTLGGLLRTKQREDE
WEDILNSKIWDLSDSQSDILPVLRLSYYHLPSHLKRCFTYSALIPKDFEFEEKDLVLLWMAEGLVPQQVQNKQMEDMGAEYFRDLVSRSIFQVANCDESR
FVMHDLVSDLAQWAAGDTCFQLGNDLNAIKQFKVSKRARHSSYIRGWDGIRKFEVFHTTKRLRTFLPLPSLLGHNTGYLTSHVPFDLLPELEFLRVLSLS
GYCIDTLPNSIGDLKHLRFLNLSFSAIRNLPQSVCSLYNLQTLLLKGCCLLEGLPSKLGSLINLRHLDITSASSIKAMPMGIEKLTNLQTLSNFVLGKDK
GSRLSSLVNLKSLRGTLCITGLENVIDAREAMEANIKDINNLEVLLLEWSPRTDNSRNEKVDKDVLDDLRPHGKVKELTINCYAGLTFPTWVGNPSFSSI
FLLRLENCTKCTSLPPLGLLPSLKNLSIVSLTAVKKVGPEFYGQGCSKPFPVLETLLFKNMQEWEEWIPCGLGSDEFPLLNKLSVKSCPNLCKKLPPSVP
SLEKLVIKKCEKLVVLIHSLPKLCKMVINGCKEVVYEGGVYLRSLNSMTISNISKLTYLAEGFIQPLAEVQELEIANCMELASLYENGVALAKQLTSLLK
LEVRNCPQVVSLMEGEVPVYMQQQLANCKLESLTFSTCESLKKLPQWVHSLVSLKELKIQYCPRLLSFPEAGLPSTLRIIEIVGCNALTPLPAAVTYNMM
CLEQLRIENCESLISFGRIQLPPTLKKLEIRYCENLLCLLDDGEGSSSKKSDENTSCSGNNSSLLEYLYVGICNSLTSIGELPSALKYLQVCSCSKLKSL
SSRDKLPAGLKHLAIDSCENLESMPDRFQDNMSLENLKIWFCFNLRSLPEGLHKLCHLREISIWYCPALVSFAAEGLPINLRRLFIIKCDGLKAIPDHMH
NLMSLEELSIYYCPDIVSFPEEGFPTSLTYLATVDLKICELLFNWGMHKLSALRTLIIQGGFSHISFPSVDMGVRLPSALNRLSIEDFPNLEYLSYSGFQ
NLSSLERLSISDCPKLTSFPGKGLPSSLLELRIRACPLLVQQIKGRVKEWLKIRHIPYINIDGKVVSDPATQL
|
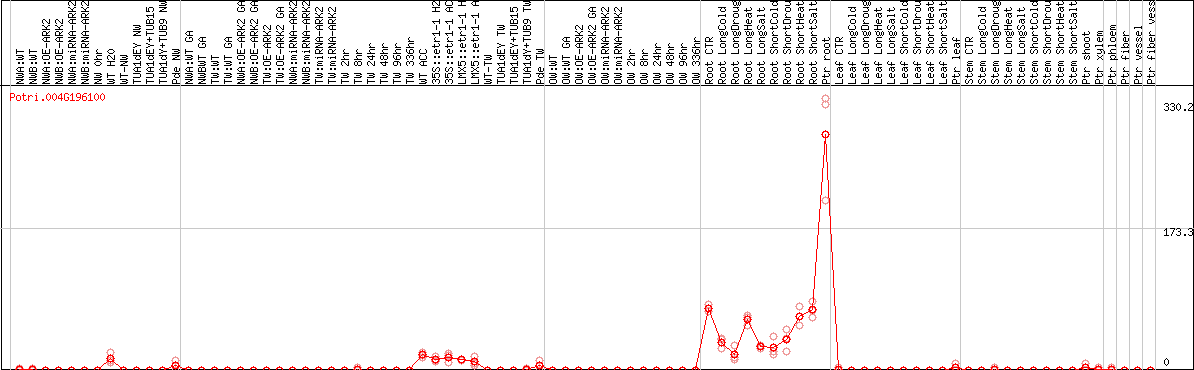
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G196100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
