Pt-STI.4 (Potri.004G230300) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-STI.4 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G230300.1 pacid=42794274 polypeptide=Potri.004G230301.1.p locus=Potri.004G230300 ID=Potri.004G230300.1.v4.1 annot-version=v4.1
ATGTCAGAAATGAGATTTTCTGATCCAAGCAGACTTCATTTAAAGAAGGAGCTTACTCAAATCAGGAAGGCAGCTAGAGTTTTGAGAGATCCAGGAACAA
GTTCTTCATGGAAGTCACCTCTTAATTCAGCTAGATCTGCAGCGACTATGGCGGCGGCTGCGTCTTCTACTTCTGCTTCCGCTTGGAAGCATTTTGAGAC
TGAGAATGCAATCCAAAATGGTGGTGGTGGTGGTAGTCATAATAACAATAGCGCCCATTTGGATTCGCATTTCAAGAGTGGGAATAATCATGGGAAAGAC
AAAAGAGTGTTCCTTTACAATTGGAAGTCTCAGAAATCTTCCAGTGAGAAGAGTGCGTTGGCAAGGAATGATGCTGATGATGACTACGAGTCTTGTTCTA
TTCAAGGGAGCCTTGATGACAGCCTGAGTGATGCAAGAAATGCTGGGGATTCGAAGAGTGACACTTATTTGGGTGAGACAAGGTCTGCTGCAATGATTTT
CAGGTGTAGAGATGCAAATCTCGTGTCACCATCTATGAGAAGAGCTATGGGGATCAAGAAAAAGAGTAAGAAAACTAATGCCCGTTTCGATGTTTTGTCT
AGATATCAACAAAAAGAGATGAACTTGAGGAGATTACTCAAGGGCCATCCTTCAATGGGCTTGGGTTTAGGTTTAGGAAGGGATGATGTTGTAGAGCAAT
CTGATGATACTGAAGAATATAGTAATTCAGAGTACTTAAGAAAGATTTCTGGGGCATCCCCATTGCTGTTAAAACTTAAGCATAAGAATCGGTCACATTC
TCCATCTAAATTATTGAGAACTACTCGAAAAGAGGACTCTTCTTATTCTCACAGCACACCTGCACTATCAGCTAGTTCTTATGATAAGTATCGTAAGCGC
AATCCGAGCAATGTTGGATCTTGGGATGCCACCACAACCTCAGTGAATGATGGGGATGATGAGGATGATGATCATTTAGATTTACCTGGTCGCCAGGGAT
GTGGCATTCCTTGCTACTGGTCCAAACGGACGCCAAGATACAGAGGTGTATGTGGGAGTAGTTGTTGCTCACCTTCCCTTTCAGATACTTTAAGGAGAAA
AGGAAGTAGCATGTTCTGTGGTAGTCAGCCACTGTATCATAGACGCCGACGTTCATGGTCAATTTCAAATAAGCGCAGAATTGGTTCAAGAACTGGACAT
GCATTGCTCCCATTGCTTACAAATAGTGGTGATGGCATAGGAGGGTCATCAATAGGAACTGGGCTTAGTGATGATGAGCTTTCTACAAACTATGGGGAAC
TTGATTTAGAGGCCTTGAGTAGATTGGATGGGAGGAGGTGGTCAAGTTGCAGGAGTCAGGATGGGTTGGAGATTGTGGCTTTAAATGGAGACGGAGAAGA
GGAAGGCACGCCTGAAAATATTGGAAGCTTAAGCCAGAAATACAAGCCAGTTTTCTTCAGTGAATTGATTGGGCAAAACATTGTTGTTCAATCACTTACC
AATGCTATTTCTAGGGGAAGGATAGCACCGGTTTATCTTTTCCAAGGTCCACGTGGTATTGGTAAAACATCAGCAGCTAGGATTTTTGCCTCTGCCTTAA
ACTGCACGTCTGCTGAAGAAATAAAACCATGTGGGTACTGTAGGGAATGCAGTGATTCCATTTCTGGGAAGACCAGGGATCTATGGGAAGTTGATGGCAC
TGACAAGAAGGGAATTGATAAAGTTAGATACCTTCTGAAGAAAATTTCGCATAGGCCCCCATTGGGCTCTTCACATTACAAGGTTTTTCTTATCGATGAG
TGTCACTTGTTACCCTCTAAGATGTGGTTAGCATTTCTCAAGTTTCTTGAAGAACCACCACAGCGAGTTGTGTTCATATTTGTAACAACTGATCCTGATA
ATGTGCCCCGTACTGTACAATCACGATGTCAGAAGTACCTCTTCAACAAAATTAAAGATGGAGATATTGTGGCAAGACTGAGGAAAATTTCTAAAGAAGA
GAATCTTGATGTTGAATTAGGTGCTCTGGATTTGATTTCTTTGAATGCAGATGGTTCACTTCGAGATGCTGAAACAATGTTAGACCAGTTGAGTTTGCTG
GGGAAGAAGATAACAACGTCTCTTGTAAATGAATTGGTGGGGGTTGTTTCAGATGAGAAATTACTGGAACTTTTGGAACTAGCGATGTCTTCAGACACTG
CAGAAACAGTAAAAAGGGCCAGGGATTTGATGGACTCTGGCGTTGATCCAATGGTTTTGATGTCTCAACTGGCTAGCCTCATTATGGACATTATTGCTGG
GACATACAATGTTGTTGATGCCAAACATGGTGATTCACTCTTTGGTACACACAACTTGACTGAAGCAGAACTGGAGAGACTAAAGCATGCTTTGAGGCTT
CTTTCAGAGGCTGAGAAACAGCTAAGGATCTCAAGTGACCGCTCAACATGGTTCACAGCAACTCTTCTACAGCTTGGTTCTACACCTTCAATGGATCTCA
CTCAGTCAAGCAGCAGTAGGAGGCAGAGCTCCAGGACAACAGAGGAAGATCCTTCAAGTGCTTCAAAGGAATCCAAGGTTTACAAGACAAAGTCCAATGC
TCAATATTTGACTCAGAGATCAAGTTCACCTCCATCCCTATACAGGGAAATAAATGGATGTTCCAGCCAACAAGGGGAATTTGGTTTTAATGCCAAGGCC
CCTCGTAGTCGATTGGTGAATAGCCGTACTTCATCTACTTCATTGGATGATGAAATAACTGGAAATATGATATTTAGATACAAAAACTCAGAGAAGTTGG
ATGATATATGGGAGAAATGTATTGAAAAGTGTCACTCACAGACATTGAGGCAGCTGCTGCATGCTCACGGAAAGCTTCTATCTATATCTGAAGTGGATGG
AGCCCTTGCTGTTTATGTTGCATTTGAGGACCAAGATATTAAAGCCAGAGCTGAAAGGTTTTTGAGCAGTATCACAAATTCAATTGAAATAGTCCTTAGA
CGCAATGTAGAGGTTAGAATTATTCTTATCACAGATGGTCTGGATTCTGTGATTTATGCGAATCAATCTGAGTTACAAGAAGGTCATAGGCAAGCAGACA
CTACGTTGGCAATTGAGCTGGGAAAGAAGGCTAATTGCTCTGATGCAGTAGCTGGTTATTCCAATTTGGACTTGCAAGAAGAGTTCCCAAAACTATCCAA
AGGAAGCTTTAATGATGCCAACGCTGAAAACAATGGAGAAGGGAAGCGGGAAATGCCAATGCAGAGAATAGAATCCATTATCCGTGAACAGAGGTTAGAA
ACTGCCTGGTTACAGGCGGCAGAGAAGGGCACTCCAGGATCATTGAGTTGTTTGAAACCCGAGAAGAATCAGGTGTTGCCTCAAGACGACACCTATCAAC
AAAGTCAAATAGATTCCATAGGTTCAGTGGCACCGTCCTCCCAGACATGGGGAGATGAATTAAATCATGAACTCAAAGTATTGAAGATGCAGAATCGGAG
GGTTCACCATAAGGACCAGATTGGTCATATGGTGGACCATTATCCAATATCACCAAGCTTATTACATGGAAGCAGCTATGTGGTCAATGGAAGCAAAGAA
AGCCTGGGATACGAGTCAAGTTCGGCTGGTGGAGGTTGCAGTGGGCTTTTATGTTGGAACACCAGCAGATCTCATAGAGCAAAGGTGAAAGAGACTCCAG
TTCAACCTCGAGGCAGAAGTGGGCGATTTTCATTGTTTGGTGAGTGTGCAAAGCAAAAGAAACCGGATAGCAGAATTACAAGATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G230300.1 pacid=42794274 polypeptide=Potri.004G230301.1.p locus=Potri.004G230300 ID=Potri.004G230300.1.v4.1 annot-version=v4.1
MSEMRFSDPSRLHLKKELTQIRKAARVLRDPGTSSSWKSPLNSARSAATMAAAASSTSASAWKHFETENAIQNGGGGGSHNNNSAHLDSHFKSGNNHGKD
KRVFLYNWKSQKSSSEKSALARNDADDDYESCSIQGSLDDSLSDARNAGDSKSDTYLGETRSAAMIFRCRDANLVSPSMRRAMGIKKKSKKTNARFDVLS
RYQQKEMNLRRLLKGHPSMGLGLGLGRDDVVEQSDDTEEYSNSEYLRKISGASPLLLKLKHKNRSHSPSKLLRTTRKEDSSYSHSTPALSASSYDKYRKR
NPSNVGSWDATTTSVNDGDDEDDDHLDLPGRQGCGIPCYWSKRTPRYRGVCGSSCCSPSLSDTLRRKGSSMFCGSQPLYHRRRRSWSISNKRRIGSRTGH
ALLPLLTNSGDGIGGSSIGTGLSDDELSTNYGELDLEALSRLDGRRWSSCRSQDGLEIVALNGDGEEEGTPENIGSLSQKYKPVFFSELIGQNIVVQSLT
NAISRGRIAPVYLFQGPRGIGKTSAARIFASALNCTSAEEIKPCGYCRECSDSISGKTRDLWEVDGTDKKGIDKVRYLLKKISHRPPLGSSHYKVFLIDE
CHLLPSKMWLAFLKFLEEPPQRVVFIFVTTDPDNVPRTVQSRCQKYLFNKIKDGDIVARLRKISKEENLDVELGALDLISLNADGSLRDAETMLDQLSLL
GKKITTSLVNELVGVVSDEKLLELLELAMSSDTAETVKRARDLMDSGVDPMVLMSQLASLIMDIIAGTYNVVDAKHGDSLFGTHNLTEAELERLKHALRL
LSEAEKQLRISSDRSTWFTATLLQLGSTPSMDLTQSSSSRRQSSRTTEEDPSSASKESKVYKTKSNAQYLTQRSSSPPSLYREINGCSSQQGEFGFNAKA
PRSRLVNSRTSSTSLDDEITGNMIFRYKNSEKLDDIWEKCIEKCHSQTLRQLLHAHGKLLSISEVDGALAVYVAFEDQDIKARAERFLSSITNSIEIVLR
RNVEVRIILITDGLDSVIYANQSELQEGHRQADTTLAIELGKKANCSDAVAGYSNLDLQEEFPKLSKGSFNDANAENNGEGKREMPMQRIESIIREQRLE
TAWLQAAEKGTPGSLSCLKPEKNQVLPQDDTYQQSQIDSIGSVAPSSQTWGDELNHELKVLKMQNRRVHHKDQIGHMVDHYPISPSLLHGSSYVVNGSKE
SLGYESSSAGGGCSGLLCWNTSRSHRAKVKETPVQPRGRSGRFSLFGECAKQKKPDSRITR
|
DESeq2's median of ratios [POPLAR]
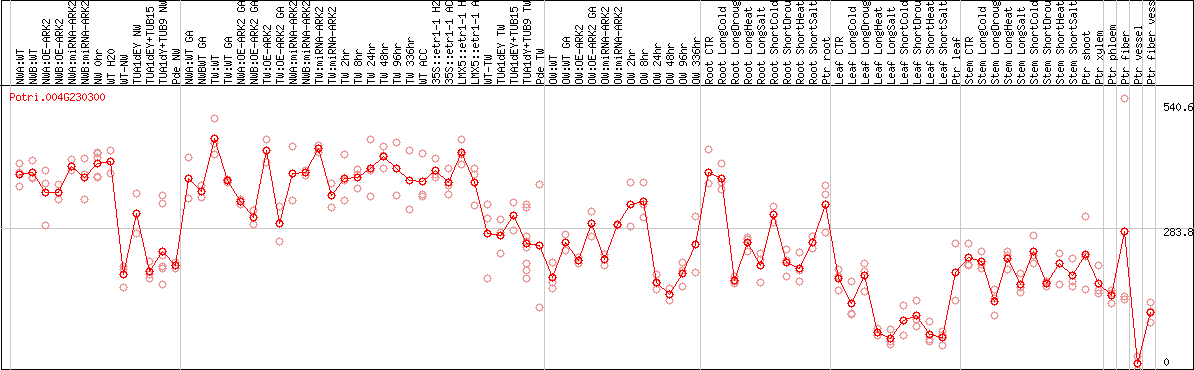
Coexpressed genes
Potri.004G230300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
