External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G61220 285 / 7e-96
SDR1
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G24190 280 / 6e-94
SDR2
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G01800 260 / 3e-86
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT5G51030 189 / 2e-58
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G59710 166 / 2e-49
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G61830 166 / 2e-49
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G09750 69 / 3e-13
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G50600 67 / 2e-12
ATHSD1
hydroxysteroid dehydrogenase 1 (.1)
AT5G50700 67 / 2e-12
HSD1
hydroxysteroid dehydrogenase 1 (.1)
AT3G47350 58 / 1e-09
ATHSD2
hydroxysteroid dehydrogenase 2 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G235500
490 / 8e-177
AT3G61220 313 / 7e-107
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156800
302 / 2e-102
AT3G61220 363 / 1e-126
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156300
302 / 2e-102
AT3G61220 367 / 6e-128
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G157000
299 / 2e-101
AT3G61220 406 / 9e-144
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G156600
297 / 2e-100
AT3G61220 348 / 7e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.014G080000
296 / 3e-100
AT3G61220 332 / 2e-114
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.014G080100
291 / 5e-98
AT3G61220 350 / 3e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.009G158200
256 / 6e-84
AT2G24190 283 / 1e-94
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.015G108400
185 / 2e-56
AT5G51030 468 / 2e-167
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014693
282 / 2e-94
AT3G61220 347 / 3e-120
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014694
281 / 3e-94
AT3G61220 369 / 4e-129
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014688
279 / 2e-93
AT3G61220 348 / 8e-121
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10008413
274 / 3e-91
AT2G24190 273 / 3e-91
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10040933
272 / 8e-91
AT3G61220 342 / 1e-118
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014690
271 / 3e-90
AT3G61220 335 / 2e-115
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10019546
266 / 3e-88
AT2G24190 253 / 2e-83
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10006894
264 / 2e-87
AT3G61220 325 / 1e-111
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10014689
269 / 2e-86
AT3G61220 336 / 2e-112
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10040932
257 / 1e-84
AT3G61220 327 / 3e-112
short-chain dehydrogenase/reductase 1, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Potri.004G235600.1 pacid=42796116 polypeptide=Potri.004G235600.1.p locus=Potri.004G235600 ID=Potri.004G235600.1.v4.1 annot-version=v4.1
ATGGCAGAAGTAATTCCAACCAAAAGGATTGCAGTCGTTACGGGTGCAAATAAAGGGATTGGTTTAGAGATTTGCAGGCAGTTGGCATCTAAAGGGGTGT
TGGTGGTTTTAACAGCTAGAGATGAGGAGAGGGGTCTTGAAGCTGTCAAGAGCCTCAAAGTTTCAGGTTTTTCTGATGTAGTTTTTCATCAACTTGATGT
GGTGGACGATCTCAGTATTGCTTCCTTCGCCAATTTCATCAGAAACCAATTCGGAAGGCTTGATATATTGGTGAACAATGCAGGCATTACTGGAACAGAA
ATTAAAGAAGATGACTGGAAGAAGTTAAGATTTGGTGTTGAAGATATTATAGGTGTAAATGCTGCATCACAAAGAAAACTTATGAAGCAAACTTATGAAA
TGTCAATTAGTTGCCTGAGAACAAATTATTATGGAATCAAGCACCTAACTGAAGCATTAATTCCTATCCTTGAACGATCCAATTCTGCAAGAATAGTCAA
TGTCTCCTCCTCCTTTGGGAAGCTAAAGTTTTTTCCTAATGAAAAAACCAAAAAGATGCTAGGTGATGTTGATGGCCTCACAGAAGAAAAAGTTGAGGAG
CTGGTGGAAGAGTTCCTGGAGGATTTTAAGAATGATTTAGTGGAAACCAAACGTTGGCCTACCCTTTTTTCAGCTTATACTGTGTCCAAGGCTGCTCAAA
ATGCCTACACAAGGATTTTGGCAAAGAAGTACCCCAAAATCGCAATCAATGCAGTTTGTCCTGGTTTTACTTGCACGGACCTTAATTGTAACAATGGAAG
CGTAACTACTGAAGAAGGTGCACGAGGTCCTGTTATGCTGGCTCTGATGCCTGATCATCAGCGCCCTTCTGGCTGCTTCTTTTTTCAGACAGAAATGTCG
ACTTTTGAATGA
AA sequence
>Potri.004G235600.1 pacid=42796116 polypeptide=Potri.004G235600.1.p locus=Potri.004G235600 ID=Potri.004G235600.1.v4.1 annot-version=v4.1
MAEVIPTKRIAVVTGANKGIGLEICRQLASKGVLVVLTARDEERGLEAVKSLKVSGFSDVVFHQLDVVDDLSIASFANFIRNQFGRLDILVNNAGITGTE
IKEDDWKKLRFGVEDIIGVNAASQRKLMKQTYEMSISCLRTNYYGIKHLTEALIPILERSNSARIVNVSSSFGKLKFFPNEKTKKMLGDVDGLTEEKVEE
LVEEFLEDFKNDLVETKRWPTLFSAYTVSKAAQNAYTRILAKKYPKIAINAVCPGFTCTDLNCNNGSVTTEEGARGPVMLALMPDHQRPSGCFFFQTEMS
TFE
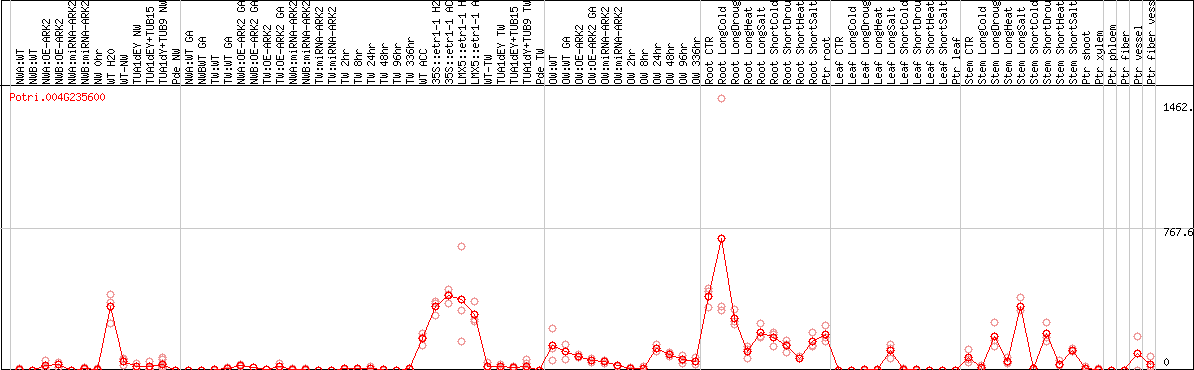
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G235600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.