Potri.005G041101 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G041101.3 pacid=42805167 polypeptide=Potri.005G041101.3.p locus=Potri.005G041101 ID=Potri.005G041101.3.v4.1 annot-version=v4.1
ATGGATATTGTGATTTCTGTTGTAGCCAAAGTAGCTGAGCTGTTAGTTGTTCCCATTAGGAGACAGATTGGTTATGTAGTTGATTGTAATACCAATATTC
AGAACTTAAAAAATGAAGTTGAGAAGCTGAGAGATGCAAAAACAAGGGTGATTCATTCCATAGAGGAGGCTAGAAGGAATGGGGAAGAGATTGAAGTTGA
TGTTGAGAACTGGCTGCGAAGTGTTGATGGGGTCATTGAAGGGGGAGGTGGTGTTGTTGGAGATGAATCAAGTAAGAAGTGCTTCATGGGTTTGTGTCCT
GATCTAAAGATACGTTATCGGTTGGGTAAAGCAGCGAAGAAGGAGTTGCCTGTTGTTGCTGTTTTCCAGGAAAAAGGGAAATTTGAGAGAGTTTCTTACC
GTGCTGATCCATCGGGTATAGGGCTTGTCAAGGATTATGAGGCCTTTGAATCAAGAAATTCTGTATTAAATGCTATTGTGGATGCATTGAAGGATGGTGG
CGTGAACATGGTTGGGGTGTATGGGATGGGTGGTGTGGGGAAGACCACACTTGTGAAGAAGGTTGCTGAACAAGTCAAGGAGGGTAGGCTGTTCGATAAG
GTGGTTCTGGCACTTGTGTCACATACTCCAGACATCAGAAGAATTCAAGGTGAAATATCAGATGGGCTGGGATTCAAATTGGATGCAGAGACTGATAAAG
GAAGGGCAAGTCAACTATGCGGGGGATTGAAGAAAGTGACTAAAGTACTTGTAATTCTTGATGATATTTGGAAAGAACTTAAATTGGAGGATGTTGGAAT
CCCTTCAGGGAGCGATCATGAAGGATGCAAGATACTCATGTCTTCAAGAAATGAATATGTACTGTCTCGCGAGATGGGTGCAAACAAAAATTTCCCGGTT
CAAGTTTTGCCTGTAAGAGAAGCATGGAATTTTTTCGTAAAGATGGTGGGGGTTACTGTTAAAAACCCTAGTGTACAACTAGTTGCTGCTGAAGTAGCCA
AAAGATGTGCAGGTTTGCCGATTTTACTTGCTACAGTTGCTAGGGCACTGAAAAACAAAGATTTATATGCTTGGAAGGAGGCCTTGAAACAGCTAACAAG
GTTTGATAAAGATGAAATTGACAATCAAGTTTACTCGTGTCTAGAACTGAGTTACATGGCTTTGAGAGGTGATGAAATCAAGCCATTGTTCTTGCTTTGT
GGGCAATTCCCAACTTATGATACTTCAATCTCAGACTTGTTAAAGTATGCTATAGGCTTGGATTTGTTCAAGGGACGTTCCACATTGGAAGAAGCAAGAA
ACAGACTGCATACATTGGTGGATGAACTTAAAGCCTCTTGTTTGTTGCTAGAAGGTGACTTTGGTGGAAGCGTGAAAATGCATGATGTTGTTCAAAGTTT
TGCCATCTCTGTTGCATCAAGGGACCACCATGTGCTTATCGTGGCAGATGAATTCAAGGAATGGCCAACCAATGATGTGCTTCAACATTACACTGCAATC
TCTTTGCCTTTCCGGAAAATTCCTGATCTTCCTGTAATATTAGAATGCCCAAATCTCAATTCGTTTCTACTGCTGAACAAGGATCCCTCACTGCAAATAC
CAGACAACTTTTTCAGAGAAATGAAGGAGCTTAAAGTCTTGGATCTGACAGGAGTTAATCTTTCACCACTGCCTTCGTCACTTCAATTTCTTGAGAACCT
TCAAACGCTGTGTTTGGATCATTGTGTTTTGGAAGATATATCTATAGTTGGAGAGCTAAAGAAGTTAAAAGTTCTCAGCTTAATCAGTTCTAATATTGTT
TGTTTGCCAAGAGAAATCGGAAAATTGACTCGTCTGCTACTTCTAGATTTGAGCAATTGTGAAAGGCTTGAAGTAATATCACCAAATGTCCTTTCTTCCT
TGACCCGGCTAGAAGAATTATATATGGGAAACAGCTTCGTGAAATGGGAGACTGAAGGACCAAGCAGTCAAAGAAACAATGCTTGCCTGTCAGAATTGAA
GCTTCTGTCAAACTTAATTACCTTACACATGCAAGTTACAGATGCAGATAATATGCCCAAGGACTTGTTTTTGTGCTTTCAAAAGTTGGAAAGGTTCAGA
ATATTTATTGGAGATGGGTGGGACTGGTCTGTTGAGGATGCAACGTCAAGAACTTTGAAACTCAAGCTCTATACAGTTATTCAGTTGGAGGAGTGGGTGA
AAACGCTGCTGAAGATCACTGAAGAACTTCATTTGCAAGAACTGAATGGTGTTAAGAGTATCCTCAATGACTTGGACGGGGAAGGTTTCCCTCAGCTGAA
GCATCTTCACGTCCAAAATTGTCCTGGAATTCAATATATCATCAACTCGATTAGGATGGGTCCTCGAGCTGCTTTTCTGAACTTGGACTCACTGTTTCTT
GAAAATTTGGATAACTTGGAGAAGATCTGTCATGGCCAACTTATGGCAGAGTCGCTTGGAAACTTGAGAATTTTGAAGGTAGAAAGTTGTCATAGGTTGA
AGAATTTATTTTCATTCTCCATGGCAAGAAGACTTGTGCGCCTTGAAGAAATCACAATAATTGATTGCAAGATCATGGAAGAGGTTGTTGCTGAGGAAAG
TGAGAATGATGCTGCTGACGGTGAACCAATAGAGTTCACTCAATTACGTAGACTGACACTTCAATGTCTGCCTCAGTTCACAAGCTTTCACTCCAATGTC
GAGGAATCTTCTGACTCGCAGAGAAGGCAGAAGCTAGTAGCAAGTGATGCGAGGTCAAAAGAAATTGTTGCGGGGAATAAGCTTGGGACTTCCATGTCAC
TTTTCAATACAAGGATTTTATTCCCCAATTTGGAGGACCTGAAGCTCTCCTCTATCAAAGTGGAAAAGATATGGCATGACCAACCTGCTGTGCAGTCTCC
CTGTGTTAAAAACTTAGCAAGCATTGTTGTGGAGAATTGTAGTAATTTAAACTATTTATTGACATCTTCTATGGTCGAAAGGCTTGCTCAACTCAAAAGG
CTTGAAATATGTAATTGCAAGTCCATGGAAGAAATAGTTGTTCCAGATGGAATCGGAGAAGGCAAAATGATGAGCAAGATGCTATTCCCTAAACTACTCA
TCCTAGAGCTCAATGGTCTGCCGAAACTCACAAGATTCTGCACCAGTAACTTGCTTGAATGCCACTTCCTGAAAGTGCTGACGTTAGGGAATTGCCCTGA
ACTGAAGGAATTTATTTCCATCCCATCAAGTGCAGATGTCCCTGTCATGAGCAAACCTGACAACACAAAATCAGCTCTCTTCGATGACAAGGTTGCATTC
CCTGACTTGGAGGTATTTCGAATTTTCGACGTGGATAATTTGAAGGTGATATGGCACAATGAGCTCCATTCTGATTCCTTTTGCAAACTAAAGACACTGG
AGGTTAGGCGTGGAAAGAATTTACTGAATATTTTCCCTTCCAGTATGCTGAGAAGATTCCATAATCTAGAGAATCTGAGTATAGCTGATTGTGATTCAAT
AGAAGAGATTTTTGATCTCCAAGTGCTCATAAATGTGAAACAAAGACTTGCTGTTACAGCCACTCAATTGAGAGTTGTGCGTCTAATGAATCTTCCATAT
CTGAAGCATGTTTGGAATACGGATCCACAAGGAATTCTTTCCTTCCATAACCTATGTACCGTGCATGTCCGGAGATGTCCTGGTCTAAGAAGTCTCTTTC
CTGCCTCTATAGCCCAAACCCTTCTACAACTTGAAGAGCTTCAGATAGATACTTGTGGAGTGGAGGAAATTGTTGCAATGGATGAAGGACTAGGAGAAGG
TCCTTCATCCTTTCGGTTTTGGTTTCCTAAAGTTACCCATCTGCATCTTGTGCAATTACCAGAACTACAGAGATTCTATCCAGGAATACATACATCAGAA
TGGCCGAGGTTGAAAAACTTTCGGGTGTATGACTGTGAAAAAATTGAAATATTTCCTTCAGAAATCAAGTGCACTCACGAACCATGCAGGGAGGACCACA
TGGACATCCAGGGTCAGCAACCACTTCTCTCATTTAGGAAGATCATCCCCAACTTGGAGCAATTAATTTTGGAAAGCAAGGATGCATCGGCATTATTGAA
AGGCCTGTGTCCTCAAGACTTATTTCACAAACTCAAAGTTCTCGAGCTAATTTTCTTTCGTGATGCACATGCAACTTTTCCATTTGATCTCCTTCCAAGG
TTCCCTAATATGGAAAAACTTATTGTGGGACGTAGTACGTTCAAAGAGTTGCTCCCGTCAAGACTTGATGGTATGGAGAAGCATGCAAGAGTTCTTTCAC
CGATACGACACTTAGAGTTGGATTCTCTTCCTTGCCTAGAGCATTTATGGAAGAGCAATTCCCAACTGGACCAGGCTCTTCAAACTCTTCAAACTCTAAG
GGTACGGAATGGCGGCAGTTTGATTTACTTAGCTCCATCGCGTGCTTCTTTCCAAAATCTAACAAATTTGTATGTGTGGAGATGCGAAAGATTAGTAATA
TTGGTGACATCCACAACAGCAAAAAGTCTGGCGCAACTCACAAGAATGAGTATAAACTACTGCAGTATGGTTACCGAAATAGTAGCAAACGAGGGAGACG
GAATAAAAGATGAGATTGTCTTCAGCAAATTAGAAAGATTAGAGCTTTATCGTTTACCAACCCTAACTAGCTTCTGCTCAGAAAAACACAGCTTTGATTT
CCCATCTCTGGTAGAAGTAACTGTCGAAAAATGTCCTGAGATGAAGTTTTTCTCCAACGGAGCACTGAGCACTCCCAAGCTGCGGAGAGTTAATTTGACT
AAAGTAGAGAAGAAAAATCTCAACACCACAATACAACAGTTGTCTACACAAACGAAAGCTCAAATTGCTGAAAGTTCATCATCGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G041101.3 pacid=42805167 polypeptide=Potri.005G041101.3.p locus=Potri.005G041101 ID=Potri.005G041101.3.v4.1 annot-version=v4.1
MDIVISVVAKVAELLVVPIRRQIGYVVDCNTNIQNLKNEVEKLRDAKTRVIHSIEEARRNGEEIEVDVENWLRSVDGVIEGGGGVVGDESSKKCFMGLCP
DLKIRYRLGKAAKKELPVVAVFQEKGKFERVSYRADPSGIGLVKDYEAFESRNSVLNAIVDALKDGGVNMVGVYGMGGVGKTTLVKKVAEQVKEGRLFDK
VVLALVSHTPDIRRIQGEISDGLGFKLDAETDKGRASQLCGGLKKVTKVLVILDDIWKELKLEDVGIPSGSDHEGCKILMSSRNEYVLSREMGANKNFPV
QVLPVREAWNFFVKMVGVTVKNPSVQLVAAEVAKRCAGLPILLATVARALKNKDLYAWKEALKQLTRFDKDEIDNQVYSCLELSYMALRGDEIKPLFLLC
GQFPTYDTSISDLLKYAIGLDLFKGRSTLEEARNRLHTLVDELKASCLLLEGDFGGSVKMHDVVQSFAISVASRDHHVLIVADEFKEWPTNDVLQHYTAI
SLPFRKIPDLPVILECPNLNSFLLLNKDPSLQIPDNFFREMKELKVLDLTGVNLSPLPSSLQFLENLQTLCLDHCVLEDISIVGELKKLKVLSLISSNIV
CLPREIGKLTRLLLLDLSNCERLEVISPNVLSSLTRLEELYMGNSFVKWETEGPSSQRNNACLSELKLLSNLITLHMQVTDADNMPKDLFLCFQKLERFR
IFIGDGWDWSVEDATSRTLKLKLYTVIQLEEWVKTLLKITEELHLQELNGVKSILNDLDGEGFPQLKHLHVQNCPGIQYIINSIRMGPRAAFLNLDSLFL
ENLDNLEKICHGQLMAESLGNLRILKVESCHRLKNLFSFSMARRLVRLEEITIIDCKIMEEVVAEESENDAADGEPIEFTQLRRLTLQCLPQFTSFHSNV
EESSDSQRRQKLVASDARSKEIVAGNKLGTSMSLFNTRILFPNLEDLKLSSIKVEKIWHDQPAVQSPCVKNLASIVVENCSNLNYLLTSSMVERLAQLKR
LEICNCKSMEEIVVPDGIGEGKMMSKMLFPKLLILELNGLPKLTRFCTSNLLECHFLKVLTLGNCPELKEFISIPSSADVPVMSKPDNTKSALFDDKVAF
PDLEVFRIFDVDNLKVIWHNELHSDSFCKLKTLEVRRGKNLLNIFPSSMLRRFHNLENLSIADCDSIEEIFDLQVLINVKQRLAVTATQLRVVRLMNLPY
LKHVWNTDPQGILSFHNLCTVHVRRCPGLRSLFPASIAQTLLQLEELQIDTCGVEEIVAMDEGLGEGPSSFRFWFPKVTHLHLVQLPELQRFYPGIHTSE
WPRLKNFRVYDCEKIEIFPSEIKCTHEPCREDHMDIQGQQPLLSFRKIIPNLEQLILESKDASALLKGLCPQDLFHKLKVLELIFFRDAHATFPFDLLPR
FPNMEKLIVGRSTFKELLPSRLDGMEKHARVLSPIRHLELDSLPCLEHLWKSNSQLDQALQTLQTLRVRNGGSLIYLAPSRASFQNLTNLYVWRCERLVI
LVTSTTAKSLAQLTRMSINYCSMVTEIVANEGDGIKDEIVFSKLERLELYRLPTLTSFCSEKHSFDFPSLVEVTVEKCPEMKFFSNGALSTPKLRRVNLT
KVEKKNLNTTIQQLSTQTKAQIAESSSS
|
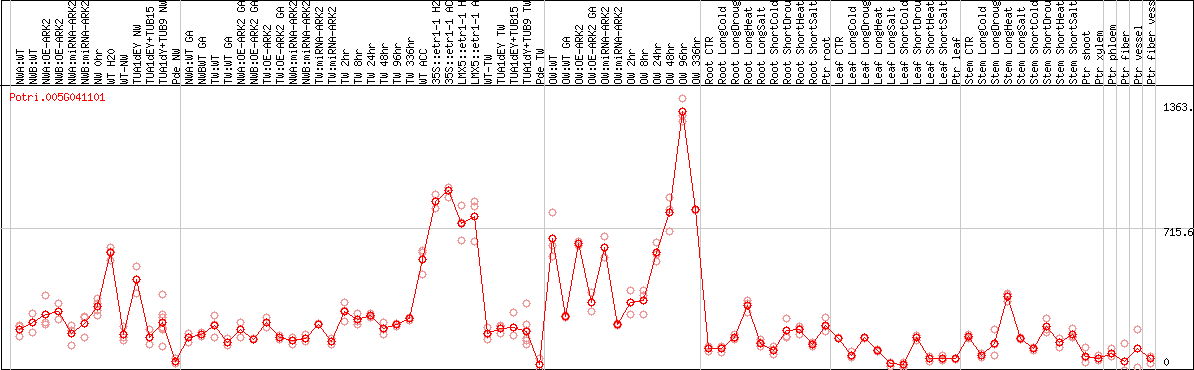
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G041101 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
