Potri.005G042400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G042400.3 pacid=42805148 polypeptide=Potri.005G042400.3.p locus=Potri.005G042400 ID=Potri.005G042400.3.v4.1 annot-version=v4.1
ATGGAATTTGTGATTTCTATTGTAGCCACAGTTGCTGAGCTGTTAGTTGTTCCCATTAAGAGACAGATTGGTTATGTACTTGATTGTAATACCAATATTC
AGAACTTAAAAAATGAAGTTGAGAAGCTGACAGATGCAAAAACAAGGGTGAATCATTCTATAGAGGAGGCTAGAAGGAATGGGGAAGAGATTGAAGTTGA
TGTTGAGAACTGGCTGACAAGTGTTAATGGGGTCATTGGAGGGGGAGGTGGTGTCGTTGTAGATGAATCAAGTAAGAAGTGCTTCATGGGTTTGTGTCCT
GATCTAAAGCTACGTTATCGGTTGGGTAAAGCAGCGAAGAAGGAGTTGACGGTTGTTGTTAATCTCCAGGAAAAAGGGAAATTTGATAGAGTTTCTTACC
GTGCTGCTCCATCGGGTATAGGGCCTGTCAAGGATTATGAGGCCTTTGAATCAAGAAATTCTGTATTAAATGATATTGTGGATGCATTGAAGGATTGTGA
TGTGAACATGGTTGGGGTATATGGGATGGGTGGTGTGGGGAAGACCACACTTGCGAAGAAGGTTGCTGAACAAGTCAAGGAGGGTAGGCTGTTCGATAAG
GTGGTTCTGGCAGTTGTGTCACATACTCCAGACATCAGAAGAATTCAAGGTGAAATAGCAGATGGGCTAGGACTCAAATTAAATGCAGAGACTGATAAAG
GAAGGGCAGATCAACTATGCGAGGGATTGAAGAAAGTGACTAGAGTACTTGTAATTCTTGATGATATTTGGAAAGAACTTAAATTGGAGGATGTCGGAAT
CCCTTCAGGGAGCGATCATGAAGGATGCAAGATACTCATGACTTCAAGAAATAAAAATGTACTGTCTCGGGAGATGGGCGCAAACAGAAATTTCCAGGTT
CAAGTTTTGCCTGTAAGAGAAGCATGGAATTTTTTCGAAAAGATGGTGGGGGTTACCGTTAAAAATCCTAGTGTACAACCTGTTGCTGCTGAAGTAGCCA
AAAGATGTGCAGGTTTGCCGATTTTACTTGCTACAGTTGCTAGGGCACTGAAAAACGAAGATTTATATGCTTGGAAGGACGCCTTGAAACAGCTAACAAG
GTTTGATAAAGATGAAATTGACAATCAAGTTTACTCGTGTCTGGAACTGAGTTACAAAGCTTTGAGAGGTGATGAAATCAAGTCATTGTTCTTGCTTTGT
GGGCAATTCCTAACTTATGATTCTTCAATCTCAGACTTGTTAAAGTATGCTATAGGCTTGGATTTGTTCAAGGGACGTTCCACATTGGAAGAAGCTAGAA
ACAGACTGCGTACATTGGTGGATGAACTTAAAGCCTCTTGTTTGTTGCTAGAAGGTGACAAGGATGGAAGAGTGAAAATGCATGATGTTGTTCAAAGTTT
TGCCTTCTCTGTTGCATCAAGGGACCACCATGTGCTTATCGTGGCAGATGAATTCAAAGAATGGCCAACCAGTGATGTGCTTCAACAGTACACTGCAATC
TCTTTGCCTTACCGGAAAATTCCTGATCTTCCTGCAATATTAGAATGTCCAAATCTCAATTCGTTTATACTGCTGAACAAGGATCCCTCACTGCAAATAC
CAGACAACTTTTTCAGAGAAATGAAGGAGCTTAAAGTCTTGGATCTGACACGAGTTAATCTTTCACCACTGCCTTCGTCACTTCAATTTCTTGAGAACCT
TCAAACGCTGTGTTTGGATGGTTGTGTTTTGGAAGATATATCTATAGTTGGAGAGCTAAAGAAGTTGAAAGTTCTCAGCTTAATCAGTTCTGATATTGTT
TGTTTGCCAAGAGAAATAGGAAAATTGACTCGTCTGCTACTTCTAGATTTGAGCAATTGTGAAAGGCTTGAAGTAATATCACCAAATGTCCTTTCTTCCT
TGACCCGGCTAGAAGAATTATATATGGGAAACAGCTTCGTGAAATGGGAGACTGAAGGATCAAGCAGTCAAAGAAACAATGCTTGCCTGTCAGAATTGAA
GCGTCTGTCAAACTTAATTACCTTACACATGCAAATTACAGATGCAGATAATATGCTCAAAGACTTGTCTTTTCTCTTTCAAAAGCTGGAAAGGTTCAGA
ATATTTATTGGAGATGGGTGGGACTGGTCTGTTAAGTATGCAACCTCAAGAACTTTGAAACTTAAGCTCAATACAGTTATTCAGTTGGAGGAGTGGGTGA
ACACGCTGCTGAAGAGCACTGAAGAACTTCATTTGCAAGAACTGAAGGGTGTTAAGAGTATCCTCAATGACTTGGACGGGGAAGATTTCCCTCGGCTGAA
GCATCTTCACGTTCAAAATTGTCCTGGAGTTCAATATATCATCAACTCAATCAGGATGGGTCCTCGAACTGCTTTTCTGAACTTGGACTCACTGTTTCTT
GAAAATTTGGATAACTTGGAGAAGATCTGCCACGGCCAACTTATGGCAGAGTCGCTTGGAAAGTTGAGAATTTTGAAGGTAGAAAGCTGTCATAGGTTGA
AGAATTTATTTTCAGTCTCCATGGCAAGAAGACTTGTGCGACTTGAAGAAATCACAATAATTGATTGCAAGATCATGGAAGAGGTTGTCGCTGAGGAAAG
TGAGAATGATACTGCTGACGGTGAACCGATAGAGTTCGCTCAATTACGTAGACTGACACTGCAATGTCTGCCTCAGTTCACAAGCTTTCACTCTAATAGA
AGGCAGAAGCTATTAGCAAGTGATGTGAGGTCAAAAGAAATTGTTGCGGGGAATGAGCTTGGGACTTCCATGTCACTTTTCAATACAAAGATTTTATTCC
CCAATTTGGAGGACCTGAAGCTCTCCTCCATCAAAGTGGAAAAGATATGGCACGACCAACCTGCTGTGCAGCCTCCCTGTGTTAAAAACTTAGCAAGCAT
GGTTGTGGAGAGTTGTAGTAATTTAAACTATTTATTGACATCTTCTATGGTCGAAAGTCTTGCTCAACTCGAAAGGCTTGAAATATGTAATTGCGAGTCC
ATGGAGGAAATAGTTGTTCCAGAAGGAATCGGAGAAGGCAAAATGATGAGCAAGATGCTATTCCCCAAACTACACCTCCTAGAGCTCAGTGGTTTGCCAA
AACTCACAAGATTCTGCACCAGTAACTTGCTTGAATGCCACTCCCTGAAAGTGCTGATGGTAGGGAATTGCCCTGAACTGAAGGAATTTATTTCCATCCC
ATCAAGTGCAGATGTCCCTGTCATGAGCAAACCTGACAACACAAAATCAGCTTTCTTCGATGACAAGGTTGCATTCCCTGACTTGGAGGTATTTCTAATT
TTCGAAATGGATAATTTGAAGGCGATATGGCACAATGAACTCCATTCTGATTCCTTTTGCGAACTGAAGATTCTGCATGTTGGGCATGGAAAGAATTTAC
TGAATATTTTCCCTTCCAGTATGCTGGGAAGATTACATAATCTAGAGAATCTGATTATAAATGATTGTGATTCAGTAGAAGAGATTTTTGATCTCCAAGT
GCTCATAAATGTGGAACAAAGACTTGCTGATACAGCCACTCAATTGAGAGTTGTGCGTCTAAGGAATCTTCCACATCTGAAGCATGTTTGGAATAGGGAT
CCACAAGGAATTCTTTCCTTCCATAACCTATGTACTGTGCATGTCCGGGGATGTCCTGGTCTAAGAAGTCTCTTCCCTGCCTCTATAGCCCTAAACCTTC
TACAACTTGAAGAGCTTCTGATAGAAAATTGTGGAGTGGAGGAAATTGTTGCAAAGGATGAAGGACTAGAAGAAGGTCCTTCATCCTTTCGGTTTTCGTT
TCCTAAAGTTACCTATCTGCATCTTGTGGAAGTACCAGAACTAAAGAGATTCTATCCAGGAGTACATGTATCAGAATGGCCGAGGTTGAAAAAGTTTTGG
GTGTATCACTGTAAGAAAATTGAAATATTTCCTTCAGAAATCAAGTGCTCTCACGAACCATGCAGGGAGGACCACGTGGACATCCAGGGTCAGCAACCAC
TTCTCTCATTTAGGAAGATCATCCCCAACTTGGAGGATTTATATTTGGAAAGTAAGGATGCATCGGCATTATTGAAAAGCCTGTGCCCTCAAGACTTCTA
TTACAAGCTCAAAGTTCTCAATCTGGTTTGCTTCCATGATGCACATGCAACTTTTCCAATTGATCTCCTTCCAAGGTTCCCTAAATTGGAAAAACTTATT
GCGGGATGTAGTGAGTTCAAAGAGTTGCTCCCGTCTAGACTTGATTGCATGGAGAAGCATGCAAGAGTTCTTTCATCGATACGATACTTAGAGTTGCAAA
TTCTTCCTTGCCTAGAGCATTTATGGAAGAGCAATTCCCAACTGGACCAAGCTCTGCAGACTCTTGAAACTCTAGTGGTACAGAATTGCAGCAGTTTGAT
TTATTTAGCTCCATCCCGTGCTTCTTTCCAAAATCTAACAAATTTGGATGTGCGGGATTGCAAGAGATTAGTAAAATTAGTGACATCCACAACAGCAAAA
AGTCTGGCGCAACTCACAAGAATGAGTATAAAGGACTGCGGTATGGTTACAGAAATAGTAGCAAACGAGGGAGAGGGAATAAAAGATGAAATTGTCTTCA
GCAAATTAGAAATATTAGAGCTTCATCTTTTACCAAGCCTAACTAGCTTTTGCTCAGAAAAACACAGCTTTGATTTCCCATCTCTGGTAGAAGTAACTGT
CGAACAATGTCCTGAGATGAAGTTTTTCTCCAACGGAGCACTGAGCACTCCCAAGCTGCGGAGAGTTAAATTGACTGAAGAGGACAAGAAAGGATCCTGG
GTTGGAAATCTTAACACCACAATACAACAGTTGTCTACACAAACGAAAGCTCAAATTGCTGGAAGTTCATCATCGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G042400.3 pacid=42805148 polypeptide=Potri.005G042400.3.p locus=Potri.005G042400 ID=Potri.005G042400.3.v4.1 annot-version=v4.1
MEFVISIVATVAELLVVPIKRQIGYVLDCNTNIQNLKNEVEKLTDAKTRVNHSIEEARRNGEEIEVDVENWLTSVNGVIGGGGGVVVDESSKKCFMGLCP
DLKLRYRLGKAAKKELTVVVNLQEKGKFDRVSYRAAPSGIGPVKDYEAFESRNSVLNDIVDALKDCDVNMVGVYGMGGVGKTTLAKKVAEQVKEGRLFDK
VVLAVVSHTPDIRRIQGEIADGLGLKLNAETDKGRADQLCEGLKKVTRVLVILDDIWKELKLEDVGIPSGSDHEGCKILMTSRNKNVLSREMGANRNFQV
QVLPVREAWNFFEKMVGVTVKNPSVQPVAAEVAKRCAGLPILLATVARALKNEDLYAWKDALKQLTRFDKDEIDNQVYSCLELSYKALRGDEIKSLFLLC
GQFLTYDSSISDLLKYAIGLDLFKGRSTLEEARNRLRTLVDELKASCLLLEGDKDGRVKMHDVVQSFAFSVASRDHHVLIVADEFKEWPTSDVLQQYTAI
SLPYRKIPDLPAILECPNLNSFILLNKDPSLQIPDNFFREMKELKVLDLTRVNLSPLPSSLQFLENLQTLCLDGCVLEDISIVGELKKLKVLSLISSDIV
CLPREIGKLTRLLLLDLSNCERLEVISPNVLSSLTRLEELYMGNSFVKWETEGSSSQRNNACLSELKRLSNLITLHMQITDADNMLKDLSFLFQKLERFR
IFIGDGWDWSVKYATSRTLKLKLNTVIQLEEWVNTLLKSTEELHLQELKGVKSILNDLDGEDFPRLKHLHVQNCPGVQYIINSIRMGPRTAFLNLDSLFL
ENLDNLEKICHGQLMAESLGKLRILKVESCHRLKNLFSVSMARRLVRLEEITIIDCKIMEEVVAEESENDTADGEPIEFAQLRRLTLQCLPQFTSFHSNR
RQKLLASDVRSKEIVAGNELGTSMSLFNTKILFPNLEDLKLSSIKVEKIWHDQPAVQPPCVKNLASMVVESCSNLNYLLTSSMVESLAQLERLEICNCES
MEEIVVPEGIGEGKMMSKMLFPKLHLLELSGLPKLTRFCTSNLLECHSLKVLMVGNCPELKEFISIPSSADVPVMSKPDNTKSAFFDDKVAFPDLEVFLI
FEMDNLKAIWHNELHSDSFCELKILHVGHGKNLLNIFPSSMLGRLHNLENLIINDCDSVEEIFDLQVLINVEQRLADTATQLRVVRLRNLPHLKHVWNRD
PQGILSFHNLCTVHVRGCPGLRSLFPASIALNLLQLEELLIENCGVEEIVAKDEGLEEGPSSFRFSFPKVTYLHLVEVPELKRFYPGVHVSEWPRLKKFW
VYHCKKIEIFPSEIKCSHEPCREDHVDIQGQQPLLSFRKIIPNLEDLYLESKDASALLKSLCPQDFYYKLKVLNLVCFHDAHATFPIDLLPRFPKLEKLI
AGCSEFKELLPSRLDCMEKHARVLSSIRYLELQILPCLEHLWKSNSQLDQALQTLETLVVQNCSSLIYLAPSRASFQNLTNLDVRDCKRLVKLVTSTTAK
SLAQLTRMSIKDCGMVTEIVANEGEGIKDEIVFSKLEILELHLLPSLTSFCSEKHSFDFPSLVEVTVEQCPEMKFFSNGALSTPKLRRVKLTEEDKKGSW
VGNLNTTIQQLSTQTKAQIAGSSSS
|
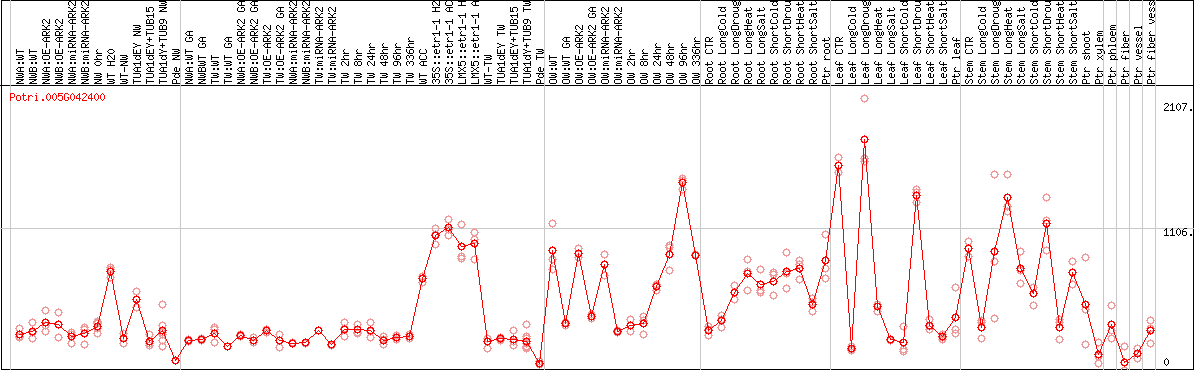
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G042400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
