External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G02840 152 / 3e-49
Small nuclear ribonucleoprotein family protein (.1.2)
AT3G07590 151 / 8e-49
Small nuclear ribonucleoprotein family protein (.1.2)
AT1G20580 41 / 1e-05
Small nuclear ribonucleoprotein family protein (.1)
AT1G76300 40 / 5e-05
SMD3
snRNP core protein SMD3 (.1)
AT5G27720 39 / 0.0001
LSM4, EMB1644
SM-like protein 4, embryo defective 1644, Small nuclear ribonucleoprotein family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G054800
158 / 1e-51
AT3G07590 183 / 1e-61
Small nuclear ribonucleoprotein family protein (.1.2)
Potri.002G010200
42 / 7e-06
AT1G20580 214 / 5e-73
Small nuclear ribonucleoprotein family protein (.1)
Potri.005G251100
41 / 2e-05
AT1G20580 211 / 6e-72
Small nuclear ribonucleoprotein family protein (.1)
Potri.002G205700
39 / 0.0001
AT5G27720 179 / 5e-59
SM-like protein 4, embryo defective 1644, Small nuclear ribonucleoprotein family protein (.1)
Potri.014G130700
37 / 0.0006
AT5G27720 176 / 7e-58
SM-like protein 4, embryo defective 1644, Small nuclear ribonucleoprotein family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028022
149 / 3e-48
AT3G07590 181 / 1e-60
Small nuclear ribonucleoprotein family protein (.1.2)
Lus10003727
150 / 2e-47
AT3G07590 180 / 1e-58
Small nuclear ribonucleoprotein family protein (.1.2)
Lus10025186
136 / 8e-43
AT3G07590 169 / 6e-56
Small nuclear ribonucleoprotein family protein (.1.2)
Lus10016068
136 / 8e-43
AT3G07590 169 / 6e-56
Small nuclear ribonucleoprotein family protein (.1.2)
Lus10030747
42 / 1e-05
AT1G20580 214 / 2e-73
Small nuclear ribonucleoprotein family protein (.1)
Lus10013227
42 / 1e-05
AT1G20580 214 / 2e-73
Small nuclear ribonucleoprotein family protein (.1)
Lus10012263
42 / 2e-05
AT1G20580 219 / 3e-75
Small nuclear ribonucleoprotein family protein (.1)
Lus10016013
42 / 2e-05
AT1G20580 219 / 3e-75
Small nuclear ribonucleoprotein family protein (.1)
Lus10004221
38 / 0.0005
AT5G27720 182 / 2e-57
SM-like protein 4, embryo defective 1644, Small nuclear ribonucleoprotein family protein (.1)
Lus10029426
38 / 0.0005
AT5G27720 190 / 2e-58
SM-like protein 4, embryo defective 1644, Small nuclear ribonucleoprotein family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0527
Sm-like
PF01423
LSM
LSM domain
Representative CDS sequence
>Potri.005G207900.2 pacid=42803819 polypeptide=Potri.005G207900.2.p locus=Potri.005G207900 ID=Potri.005G207900.2.v4.1 annot-version=v4.1
ATGAAGCTCGTCAGGTTTTTGATGAAGCTGAACAATGAGACCGTCTCAATTGAACTGAAGAACGGAACTGTTGTTCACGGAACCATCACAGGTGTTGATA
TCAGTATGAACACTCATCTGAAAACAGTGAAAGGGAAAAATCCAGTAACTTTGGATCATCTCAGTGTGAGGGGCAACAACATTCGTTATTACATCCTTCC
TGACAGCTTGAATCTTGAGACTTTGCTGATCGAGGAGACACCTAAGGTCAAACCTAAGAAGCCAACAGCTGGGAGGCCTCTGGGTCGCGGTAGAGGCCGA
GGCCGTGGACGTGGCCGCGGCCGCTAA
AA sequence
>Potri.005G207900.2 pacid=42803819 polypeptide=Potri.005G207900.2.p locus=Potri.005G207900 ID=Potri.005G207900.2.v4.1 annot-version=v4.1
MKLVRFLMKLNNETVSIELKNGTVVHGTITGVDISMNTHLKTVKGKNPVTLDHLSVRGNNIRYYILPDSLNLETLLIEETPKVKPKKPTAGRPLGRGRGR
GRGRGRGR
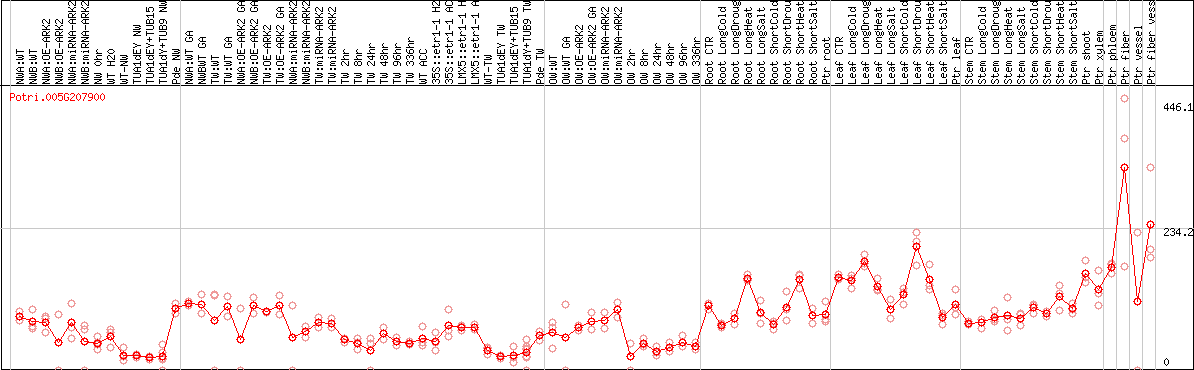
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G207900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.