Pt-CIP111.1 (Potri.006G035000) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-CIP111.1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G035000.1 pacid=42767116 polypeptide=Potri.006G035000.1.p locus=Potri.006G035000 ID=Potri.006G035000.1.v4.1 annot-version=v4.1
ATGCCATCAAAGACAAAGAAGAATTCAAAGACACCATCGAGGCTGTCAAATTCTGACCAGTCTGCTTCATCTCCTAGAACACCATCATTGGCATCTAGTA
TAGACTTAGAAGCTAGTCAACAAGAAAATATTGCGTTGTGTTTAGAAGAAGCATCGAGTAAATATCCGTATTTGATTGATAAATCTGCTTTCATTGGTCG
AATCACTGATGTTGAGGCGGAGTCTAGTACAACTGCAAGAGGTTGTAAGATTTGGCTTTCAGAATCTTCTATGGTTTCTTCTTCTCTTGCCCCTGGTTCC
ATTGTATCGGTGTCACTTGCTGCTGTGGAGAGGAGATTTTCGAGTAGTAGTTTTCCTCTCAGTTCGTTTTCCTATGAATGGTCTAGACAATGTGAAGTGG
AATCTGTGGATAAAATAACTAATGAAGCTGGGAATTACTTTGCTCTTGCGACGGTTTTTCCTTCTTGTAAGGTTTCCAAGAATGGAGCCCGGTTGTCATC
AAACCTTGCCTATATGATGGGATGCCCTGCTTCAGGCAAGGTTGTATTTGTTCACACTATTAGAAATAAACTCCTAACTGATATTGTAAATGGGAATGAT
ACGCCAGAAGGTGCAAATGCTGATGATCTTTCACTTCATAACTGCAATGAATTGTATCTGGAGCTAGTTCCTTTTATGGACAGAGTAAAAATGAAGAGTG
ATACTATGTCTGCAATGAAATTATCTGCAGAAAAAAGACATGACAGATCTGAAAATGGCATGATTTCATCCCCGAAGACACCGTTGTGTCAGCCAAAGCT
TAGCTCTCCTAGTCCCATTCACTTAACTTCACCGATATGTGAAGAAGCAGCATCCAATATATCCAATTCAAATGGCACAGATGTTGGTTTGCTGAATATC
AAAGAGGTATTAGAGGATGAAAGTGCCAAGAAACTTCTACAGGTCTGTGCTACGTCATGGTTGTATTCTCGTGTTCTAATATGTGGAAATCTTGTGGCCA
TCCCAGTGCTTTCAAATCTTTGCATTTTCCGTGTCAAAAGTGCAAATAAATTACCTGCAGATGGTAGTGATCAAGATTTGATGAAAGATAGAACTCATGG
GATGCAGCCTCAAGATTCTGAAGAATTGAGCCATATGAAAGATGCTTTTTCCATAAACCGTGAGACAAAAGTATATTTACATCAACATATGAACTCAACG
GCTGAAAGGCCACAAAAACAAGGTTTGCCATTAATGCAAAGTGAATGTATTAATGGTAAAACAATTATAGGAAATGAGAGATCAAAATTGGGTGGTCTAC
ATAAGGAATATACAGTTTTGAAGGACATAATTGTATCATCCACGAAGAACACTTTGTCATGTTTTGGCTTACGAACTACAAAGGGAGTGCTTCTTCACGG
CCCACCTGGAACTGGAAAGACTTCTTTGGCTCGATTATGTGTCATTGATGCTGGTGTCAACCTTTTCTCAGTAAACGGACCTGAGATATTTAGTCAGTAT
TATGGAGAAAGTGAACAAGCAATGCATAAAGTTTTTGATTCAGCCTGTCAATCTGCACCTGCCGTGGTGTTCATTGATGAGTTGGATGCAATTGCACCTG
CACGGAAAGATGGAGGTGAAGAACTATCTCAAAGAATGGTAGCTACATTATTGAATTTGATGGATGGGATTGCTAGGACTGATGGACTGCTTGTTATTGC
TGCTACAAACAGGCCTGATAGTATCGAGCCTGCACTGAGGCGGCCTGGAAGACTTGACAGAGAAATTGAAATAGGTGTGCCTTCTCCCAGCCAACGATTG
GACATACTTCATACTCTTCTAAGTGAAATGGAGCACTCAGTTTCAGATATGCAACTTAAACAACTTGCCATGGCTACTCATGGTTTTGTGGGTGCTGATT
TAGCAGCCCTCTGTAATGAGGCAGCTTTGGTTTGTCTCAAGCGTCATGCTAGATCCAAAAAATCTGATTATAGTTCTCGTTCCAAGGGATCATCTATTGC
ATATGAAGGCCATTCTGACAGCATGGTGAAAGGATCTGATTGCTCAACAGGGGCAAGAGATATGTTGAGGGATGGTGCAGATTCTGCATCTTCAAGCACC
TCACATTTGCCTGTTTCCTTGGAGAACCTGTCATCTTCTTGCTCGGACGGAGATGTCTCAGAGATCACAGATAATACAGAGAAAGGAATTATTGCTTGTC
CTAGAGAGGAATTTTTGGTGGAAGAAGAAGCCTTGTTGAACATTGTTTCTGAAGATTTTGAAATGGCCAGGATGAAAGTGAGGCCTAGTGCCATGAGAGA
GGTAATCCTTGAGGTTCCAAAAGTTAATTGGGAAGATGTGGGCGGCCAGGGGGAAATAAAAACCCAATTAATGGAAGCAGTGCTATGGCCTCAAACACAC
CAGGATGCATTCAAGCGCATTGGTACCCGTCCCCCAACGGGGATATTGATGTTTGGTCCCCCTGGTTGTAGCAAAACTCTTATGGCCCGTGCAGTGGCTT
CTAAAGCAGGATTGAATTTCCTTGCAGTGAAGGGTCCAGAACTTTTTAGCAAATGGGTTGGTGAATCCGAGAAGGCTGTAAGATCCCTCTTTGCAAAGGC
AAGGGCAAATGCTCCATCAATTATATTCTTTGATGAAATTGATGGCCTTGCTGTTATTCGTGGGAAGGAAAGTGATGGGGTTTCGGTTTCAGATAGGGTT
ATGAGTCAACTCCTTATTGAATTGGATGGCTTGCAACAAAGAGTTAATGTTACTGTAATTGCTGCTACAAATCGGCCAGACAAGATTGATCCTGCCCTTT
TGAGACCAGGGCGCTTTGATCGACTGTTGTACGTGGGACCTCCAAATCAGAATGACAGGGAAGACATATTTCGTATCCATTTGCACAAAGTTCCATGCAG
TTCTGATGTCAATATAAAAGAACTAGCTTGTCTCACTGATGGGTGCACTGGAGCTGATATAGCATTAATCTGCCGGGAGGCAGCTGTTGCAGCCATTGAG
GAGAATATTGATGCTTCGGAAGTACCCATGCAACATTTAAAGACTGCAATTCAACAAGTCCAGCCAACGGAGATCAATTCTTATCAAGATCTATCAGCAA
AATTTCAGAGGCTCGTTCACTCCAGTGACAAAGATGAATTGGGAAACCAAGAGTGCTCGAGCAGGGCAAACTCGTTTTCCATCAGGACCATAATAAAATC
CGCGATGCAGTTCTTGAATCGTTCCTCTGCCTCAGGCTCTGGGCAATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G035000.1 pacid=42767116 polypeptide=Potri.006G035000.1.p locus=Potri.006G035000 ID=Potri.006G035000.1.v4.1 annot-version=v4.1
MPSKTKKNSKTPSRLSNSDQSASSPRTPSLASSIDLEASQQENIALCLEEASSKYPYLIDKSAFIGRITDVEAESSTTARGCKIWLSESSMVSSSLAPGS
IVSVSLAAVERRFSSSSFPLSSFSYEWSRQCEVESVDKITNEAGNYFALATVFPSCKVSKNGARLSSNLAYMMGCPASGKVVFVHTIRNKLLTDIVNGND
TPEGANADDLSLHNCNELYLELVPFMDRVKMKSDTMSAMKLSAEKRHDRSENGMISSPKTPLCQPKLSSPSPIHLTSPICEEAASNISNSNGTDVGLLNI
KEVLEDESAKKLLQVCATSWLYSRVLICGNLVAIPVLSNLCIFRVKSANKLPADGSDQDLMKDRTHGMQPQDSEELSHMKDAFSINRETKVYLHQHMNST
AERPQKQGLPLMQSECINGKTIIGNERSKLGGLHKEYTVLKDIIVSSTKNTLSCFGLRTTKGVLLHGPPGTGKTSLARLCVIDAGVNLFSVNGPEIFSQY
YGESEQAMHKVFDSACQSAPAVVFIDELDAIAPARKDGGEELSQRMVATLLNLMDGIARTDGLLVIAATNRPDSIEPALRRPGRLDREIEIGVPSPSQRL
DILHTLLSEMEHSVSDMQLKQLAMATHGFVGADLAALCNEAALVCLKRHARSKKSDYSSRSKGSSIAYEGHSDSMVKGSDCSTGARDMLRDGADSASSST
SHLPVSLENLSSSCSDGDVSEITDNTEKGIIACPREEFLVEEEALLNIVSEDFEMARMKVRPSAMREVILEVPKVNWEDVGGQGEIKTQLMEAVLWPQTH
QDAFKRIGTRPPTGILMFGPPGCSKTLMARAVASKAGLNFLAVKGPELFSKWVGESEKAVRSLFAKARANAPSIIFFDEIDGLAVIRGKESDGVSVSDRV
MSQLLIELDGLQQRVNVTVIAATNRPDKIDPALLRPGRFDRLLYVGPPNQNDREDIFRIHLHKVPCSSDVNIKELACLTDGCTGADIALICREAAVAAIE
ENIDASEVPMQHLKTAIQQVQPTEINSYQDLSAKFQRLVHSSDKDELGNQECSSRANSFSIRTIIKSAMQFLNRSSASGSGQ
|
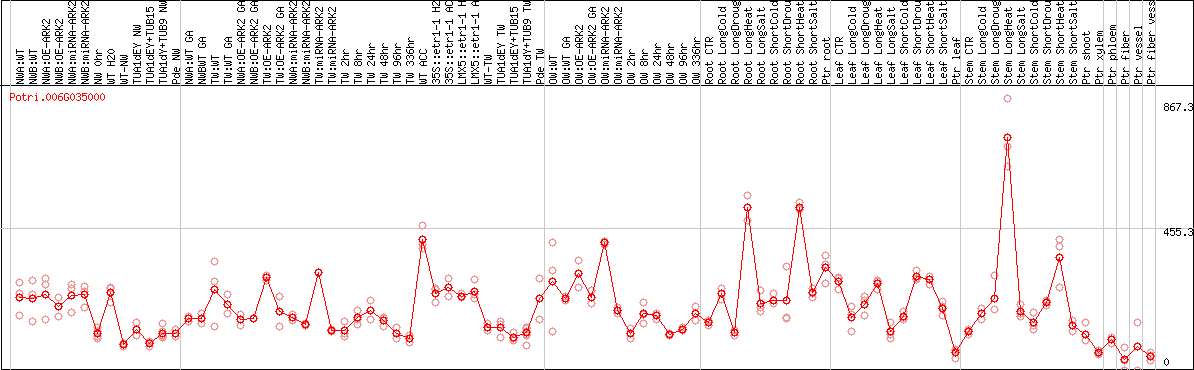
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G035000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
