External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G09600 379 / 1e-132
MYB
LCL5 (LHY-CCA1-LIKE5), REVEILLE 8 (RVE8)
REVEILLE 8, LHY-CCA1-LIKE5, Homeodomain-like superfamily protein (.1.2)
AT5G02840 363 / 2e-126
MYB
LCL1
LHY/CCA1-like 1 (.1.2.3)
AT5G52660 274 / 1e-90
MYB
Homeodomain-like superfamily protein (.1.2)
AT1G01520 261 / 2e-86
MYB
ASG4
ALTERED SEED GERMINATION 4, Homeodomain-like superfamily protein (.1)
AT4G01280 254 / 3e-83
MYB
Homeodomain-like superfamily protein (.1.2)
AT5G37260 108 / 2e-27
MYB
RVE2, CIR1
REVEILLE 2, CIRCADIAN 1, Homeodomain-like superfamily protein (.1)
AT5G17300 107 / 3e-26
MYB
RVE1
REVEILLE 1, Homeodomain-like superfamily protein (.1)
AT1G18330 102 / 2e-24
MYB
RVE7, EPR1
REVEILLE 7, EARLY-PHYTOCHROME-RESPONSIVE1, Homeodomain-like superfamily protein (.1.2)
AT3G10113 101 / 3e-24
MYB
Homeodomain-like superfamily protein (.1)
AT1G01060 101 / 2e-23
MYB
LHY, LHY1
LATE ELONGATED HYPOCOTYL 1, LATE ELONGATED HYPOCOTYL, Homeodomain-like superfamily protein (.1.2.3.4.5)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G083900
494 / 2e-177
AT3G09600 375 / 4e-131
REVEILLE 8, LHY-CCA1-LIKE5, Homeodomain-like superfamily protein (.1.2)
Potri.014G089300
290 / 2e-96
AT4G01280 293 / 2e-98
Homeodomain-like superfamily protein (.1.2)
Potri.004G073300
286 / 7e-96
AT5G52660 298 / 2e-100
Homeodomain-like superfamily protein (.1.2)
Potri.002G163100
288 / 1e-95
AT4G01280 266 / 1e-87
Homeodomain-like superfamily protein (.1.2)
Potri.017G146800
283 / 3e-94
AT5G52660 303 / 3e-102
Homeodomain-like superfamily protein (.1.2)
Potri.012G038300
105 / 5e-25
AT1G18330 158 / 5e-44
REVEILLE 7, EARLY-PHYTOCHROME-RESPONSIVE1, Homeodomain-like superfamily protein (.1.2)
Potri.002G180800
105 / 5e-25
AT1G01060 369 / 1e-117
LATE ELONGATED HYPOCOTYL 1, LATE ELONGATED HYPOCOTYL, Homeodomain-like superfamily protein (.1.2.3.4.5)
Potri.015G030400
104 / 9e-25
AT1G18330 170 / 1e-48
REVEILLE 7, EARLY-PHYTOCHROME-RESPONSIVE1, Homeodomain-like superfamily protein (.1.2)
Potri.014G106800
103 / 3e-24
AT1G01060 378 / 3e-121
LATE ELONGATED HYPOCOTYL 1, LATE ELONGATED HYPOCOTYL, Homeodomain-like superfamily protein (.1.2.3.4.5)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019902
377 / 1e-130
AT3G09600 351 / 9e-121
REVEILLE 8, LHY-CCA1-LIKE5, Homeodomain-like superfamily protein (.1.2)
Lus10014015
343 / 5e-116
AT3G09600 317 / 3e-106
REVEILLE 8, LHY-CCA1-LIKE5, Homeodomain-like superfamily protein (.1.2)
Lus10027521
277 / 3e-92
AT5G52660 353 / 1e-122
Homeodomain-like superfamily protein (.1.2)
Lus10036392
274 / 2e-90
AT5G52660 306 / 5e-103
Homeodomain-like superfamily protein (.1.2)
Lus10039284
275 / 5e-90
AT5G52660 350 / 9e-120
Homeodomain-like superfamily protein (.1.2)
Lus10007914
271 / 5e-89
AT5G52660 301 / 8e-101
Homeodomain-like superfamily protein (.1.2)
Lus10038846
240 / 2e-76
AT5G52660 292 / 2e-96
Homeodomain-like superfamily protein (.1.2)
Lus10031893
105 / 2e-25
AT3G10113 207 / 2e-63
Homeodomain-like superfamily protein (.1)
Lus10031322
105 / 2e-25
AT1G18330 199 / 4e-60
REVEILLE 7, EARLY-PHYTOCHROME-RESPONSIVE1, Homeodomain-like superfamily protein (.1.2)
Lus10042292
103 / 1e-24
AT1G18330 198 / 5e-60
REVEILLE 7, EARLY-PHYTOCHROME-RESPONSIVE1, Homeodomain-like superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Potri.006G133000.7 pacid=42768255 polypeptide=Potri.006G133000.7.p locus=Potri.006G133000 ID=Potri.006G133000.7.v4.1 annot-version=v4.1
ATGATTATGAACGCCAACTCCTCCTCTTCAACTTCAACTTCGACAGCAGGAGCAGGAGCAGAAACAGGTGCGAATCCTGCTGGTCCTTCTCAATCAATGG
CTACACCTACATCATCATCAACACCACCAGCTACAGGTACAGGTACAGGTACAACAGCAGCAGCTAGCAGCGATGGGTCAGGCAAGAAAGTTAGAAAACC
TTATACAATCACCAAGTCTCGTGAGAGTTGGACTGAAGAAGAGCATGACAAGTTCCTTGAAGCCCTTCAATTGTTTGATCGTGACTGGAAAAAGATTGAA
GATTTTGTGGGTTCAAAGACGGTGATCCAGATTCGGAGTCATGCCCAGAAATACTTTTTGAAGGTCCAAAAGAATGGTACAAGTGCGCATGTTCCTCCTC
CTCGCCCTAAACGCAAAGCCTCTCATCCTTACCCACAGAAGGCGTCGAAAAATGTTTTAGTGCCGCTCCCAGCATCCATGGCTTACGCTTCATCGATGAA
TACCTTTGCACCAGGATATGCTCTATGGGATGAAACTTCAGTGTTAATAAACTCCGCAACAAGCAAAATCATGCCATCACAAGATGAACTTCCAAATCTT
CACGGAGCTGAAGCTGATATTGGACCAAAGTGTGTATCAAGTAGTAACAATACTGTCAGTGGCCTTGGGACCTCAAGTAGAACATTGCCAAGTGCTGAGA
TGCCAAAACAAGGGAAGCAAGCGCCTGTGCTTCATGGTATACCAGATTTTGCTGAAGTATATAGCTTCATTGGAAGTGTCTTTGATCCAGACACGAAAGG
GCATGTGGAAAAGCTACAGGAAATGGACCCTATAAATTTTGAAACTGTTTTGTTGTTGATGCGGAACCTTACTGTTAACTTGTCTAGCCCTGATTTTGAG
CCATATAGGAAGGTTATGTCATCATATGATGTCAATACCAAAACAGTGGGAGTTTCTGCCCTGAAACAGACCAATGATATAGCATGTTGA
AA sequence
>Potri.006G133000.7 pacid=42768255 polypeptide=Potri.006G133000.7.p locus=Potri.006G133000 ID=Potri.006G133000.7.v4.1 annot-version=v4.1
MIMNANSSSSTSTSTAGAGAETGANPAGPSQSMATPTSSSTPPATGTGTGTTAAASSDGSGKKVRKPYTITKSRESWTEEEHDKFLEALQLFDRDWKKIE
DFVGSKTVIQIRSHAQKYFLKVQKNGTSAHVPPPRPKRKASHPYPQKASKNVLVPLPASMAYASSMNTFAPGYALWDETSVLINSATSKIMPSQDELPNL
HGAEADIGPKCVSSSNNTVSGLGTSSRTLPSAEMPKQGKQAPVLHGIPDFAEVYSFIGSVFDPDTKGHVEKLQEMDPINFETVLLLMRNLTVNLSSPDFE
PYRKVMSSYDVNTKTVGVSALKQTNDIAC
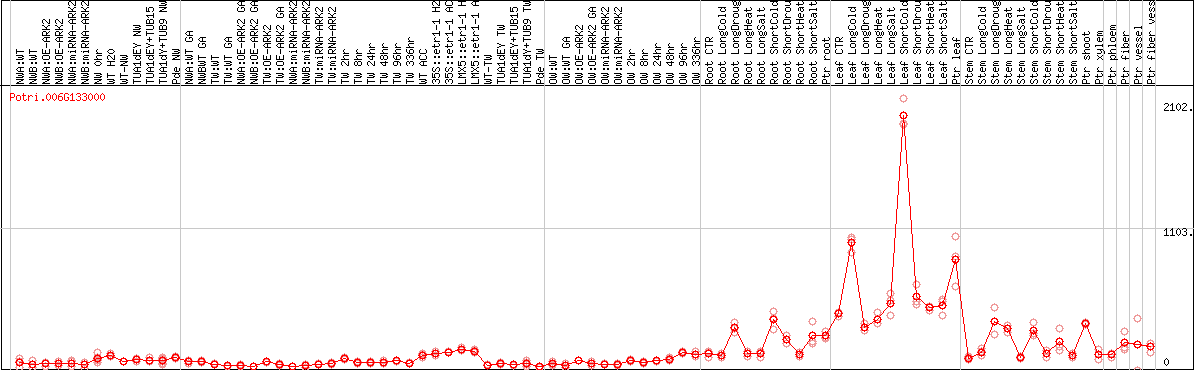
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G133000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.