External link
Symbol
DREB8,Pt-DREB2.4
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G40340 128 / 9e-35
AP2_ERF
AtERF48, DREB2C
Integrase-type DNA-binding superfamily protein (.1)
AT3G11020 120 / 6e-32
AP2_ERF
DREB2B
DEHYDRATION-RESPONSIVE ELEMENT BINDING PROTEIN 2, DRE/CRT-binding protein 2B (.1)
AT5G05410 120 / 6e-32
AP2_ERF
DREB2A
DEHYDRATION-RESPONSIVE ELEMENT BINDING PROTEIN 2, DRE-binding protein 2A (.1.2)
AT2G40350 113 / 5e-31
AP2_ERF
DREB2H
Integrase-type DNA-binding superfamily protein (.1)
AT1G75490 86 / 6e-20
AP2_ERF
DREB2D
Integrase-type DNA-binding superfamily protein (.1)
AT5G18450 84 / 2e-18
AP2_ERF
DREB2G
Integrase-type DNA-binding superfamily protein (.1)
AT3G57600 82 / 8e-18
AP2_ERF
DREB2F
Integrase-type DNA-binding superfamily protein (.1)
AT2G38340 77 / 2e-16
AP2_ERF
DREB2E, DREB19
dehydration response element-binding protein 19, Integrase-type DNA-binding superfamily protein (.1)
AT4G32800 72 / 5e-15
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT2G33710 72 / 1e-14
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G183700
329 / 1e-113
AT5G05410 124 / 1e-33
DEHYDRATION-RESPONSIVE ELEMENT BINDING PROTEIN 2, DRE-binding protein 2A (.1.2)
Potri.002G029400
88 / 2e-20
AT1G75490 163 / 3e-50
Integrase-type DNA-binding superfamily protein (.1)
Potri.005G233300
87 / 4e-20
AT1G75490 161 / 2e-49
Integrase-type DNA-binding superfamily protein (.1)
Potri.006G104200
88 / 2e-19
AT2G38340 91 / 1e-20
dehydration response element-binding protein 19, Integrase-type DNA-binding superfamily protein (.1)
Potri.016G126100
85 / 2e-18
AT2G38340 96 / 4e-22
dehydration response element-binding protein 19, Integrase-type DNA-binding superfamily protein (.1)
Potri.006G054500
82 / 4e-18
AT3G57600 234 / 1e-76
Integrase-type DNA-binding superfamily protein (.1)
Potri.016G053200
79 / 9e-17
AT3G57600 256 / 5e-85
Integrase-type DNA-binding superfamily protein (.1)
Potri.006G238600
72 / 2e-14
AT5G11590 199 / 8e-64
TINY2, Integrase-type DNA-binding superfamily protein (.1)
Potri.001G155700
71 / 2e-14
AT2G44940 159 / 1e-47
Integrase-type DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023633
137 / 3e-38
AT2G40340 181 / 1e-54
Integrase-type DNA-binding superfamily protein (.1)
Lus10034902
129 / 6e-36
AT2G40340 172 / 7e-52
Integrase-type DNA-binding superfamily protein (.1)
Lus10010632
88 / 2e-20
AT1G75490 174 / 6e-55
Integrase-type DNA-binding superfamily protein (.1)
Lus10003581
85 / 8e-20
AT2G40340 139 / 6e-42
Integrase-type DNA-binding superfamily protein (.1)
Lus10006127
85 / 5e-19
AT1G75490 155 / 8e-47
Integrase-type DNA-binding superfamily protein (.1)
Lus10024298
85 / 5e-19
AT1G75490 159 / 1e-48
Integrase-type DNA-binding superfamily protein (.1)
Lus10012226
77 / 7e-16
AT2G40340 137 / 2e-37
Integrase-type DNA-binding superfamily protein (.1)
Lus10002855
77 / 8e-16
AT2G40340 132 / 2e-37
Integrase-type DNA-binding superfamily protein (.1)
Lus10011319
75 / 2e-15
AT3G57600 207 / 5e-66
Integrase-type DNA-binding superfamily protein (.1)
Lus10010043
69 / 6e-14
AT2G23340 170 / 1e-54
DREB and EAR motif protein 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0081
MBD-like
PF00847
AP2
AP2 domain
Representative CDS sequence
>Potri.008G073600.2 pacid=42808361 polypeptide=Potri.008G073600.2.p locus=Potri.008G073600 ID=Potri.008G073600.2.v4.1 annot-version=v4.1
ATGGGTACTCTTATTCAAGGTTCTAACGCCACTTCCATGTCCATGGATTCTACTAAGAAGAGGAAAAGGGCAAGTGATAAATCTGTAGCTGAGACCCTTC
AAAAGTGGAAGGAATACAATGAGCATCTTGATGCTCAAGGTGATGGAAATAATAAACCAGTTCGTAAAGTTCCTGCCAAGGGCTCAAAGAAGGGGTGCAT
GAAAGGTAAAGGAGGGCCAGAGAATTCAGTCTGCAATTACAGAGGTGTGAGACAGAGGACATGGGGGAAGTGGGTTGCTGAGATTCGGGAGCCAAACAGA
GGGCCTAGGCTATGGCTTGGTACTTTTCCTACTGCCTATGAAGCTGCTCTTGCCTATGATAATGCTGCACGAGCTATGTATGGTTCTTGTGCTCGTTTAA
ACATTCCAGAAGTGGTGAATTCAACAAGCTCATCGAAGGATAACTTCTCTGCTGTGACACCATCATACTATTCTTCAGCTGCAAGTCCTGCTGACTCCGT
CACCACATCAACCCACTCTGAGGTTTGTGCGTACGAGGATCCTAACCAGAATGTATTAAGCCAAGCTGAGGATTGGATGACAAATATATCAAGCCAAGCT
GAGGTTTGCGAGCAGAATGTATCAAGCCAAGCTGAGGTTTACGAACAGAATGTATCAAGCCAACATATTGAGGATTGTTCTCGGGGAGTTGAAAAAAATA
GCAAGCTGAGTCAAGATGAGCTGAAGATTCAATCAGAAAATCCTTCGTGGACCAATGACTGGCACAGCTACGGCTGGGATGAGATCTTTAGCGTAGAAGA
GCTGCTAGGGGATATTGATTCTGGGATGACAGGAGCAGAGGGGTACTTCAGCTTAGGATTTTGA
AA sequence
>Potri.008G073600.2 pacid=42808361 polypeptide=Potri.008G073600.2.p locus=Potri.008G073600 ID=Potri.008G073600.2.v4.1 annot-version=v4.1
MGTLIQGSNATSMSMDSTKKRKRASDKSVAETLQKWKEYNEHLDAQGDGNNKPVRKVPAKGSKKGCMKGKGGPENSVCNYRGVRQRTWGKWVAEIREPNR
GPRLWLGTFPTAYEAALAYDNAARAMYGSCARLNIPEVVNSTSSSKDNFSAVTPSYYSSAASPADSVTTSTHSEVCAYEDPNQNVLSQAEDWMTNISSQA
EVCEQNVSSQAEVYEQNVSSQHIEDCSRGVEKNSKLSQDELKIQSENPSWTNDWHSYGWDEIFSVEELLGDIDSGMTGAEGYFSLGF
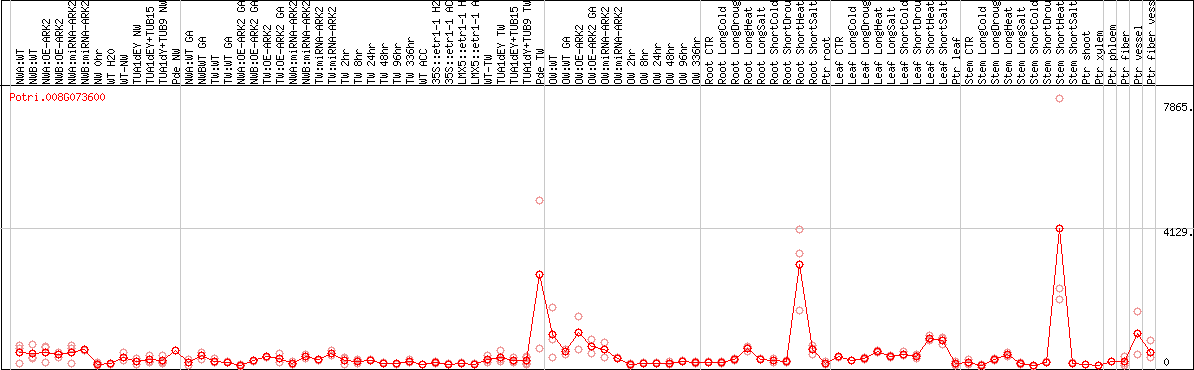
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G073600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.