External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G23230 81 / 2e-20
AP2_ERF
ERF98
Integrase-type DNA-binding superfamily protein (.1)
AT3G23220 65 / 4e-14
AP2_ERF
ESE1
ethylene and salt inducible 1, Integrase-type DNA-binding superfamily protein (.1)
AT5G43410 65 / 5e-14
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT1G06160 67 / 7e-14
AP2_ERF
ORA59
octadecanoid-responsive Arabidopsis AP2/ERF 59 (.1)
AT5G47230 67 / 7e-14
AP2_ERF
ATERF5, ATERF-5, ERF5
ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR- 5, ethylene responsive element binding factor 5 (.1)
AT5G51190 66 / 7e-14
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT4G17490 66 / 2e-13
AP2_ERF
ERF-6-6, ATERF6
ethylene responsive element binding factor 6 (.1)
AT5G61600 64 / 5e-13
AP2_ERF
ERF104
ethylene response factor 104 (.1)
AT5G47220 64 / 1e-12
AP2_ERF
ATERF-2, ERF2, ATERF2
ETHYLENE RESPONSE FACTOR- 2, ethylene responsive element binding factor 2 (.1)
AT4G17500 64 / 1e-12
AP2_ERF
ATERF-1, AtERF1
ethylene responsive element binding factor 1 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021873
81 / 7e-20
AT3G23230 143 / 4e-44
Integrase-type DNA-binding superfamily protein (.1)
Lus10011830
75 / 1e-17
AT3G23230 134 / 4e-41
Integrase-type DNA-binding superfamily protein (.1)
Lus10022936
69 / 8e-16
AT3G23230 114 / 9e-34
Integrase-type DNA-binding superfamily protein (.1)
Lus10004752
72 / 9e-16
AT5G07580 122 / 4e-34
Integrase-type DNA-binding superfamily protein (.1)
Lus10007841
71 / 1e-15
AT5G47230 121 / 4e-33
ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR- 5, ethylene responsive element binding factor 5 (.1)
Lus10024883
70 / 1e-15
AT3G23230 130 / 3e-39
Integrase-type DNA-binding superfamily protein (.1)
Lus10033884
67 / 8e-15
AT3G23230 125 / 9e-38
Integrase-type DNA-binding superfamily protein (.1)
Lus10032499
69 / 2e-14
AT5G51190 186 / 8e-59
Integrase-type DNA-binding superfamily protein (.1)
Lus10042996
68 / 3e-14
AT5G51190 189 / 6e-60
Integrase-type DNA-binding superfamily protein (.1)
Lus10011831
66 / 3e-14
AT3G23230 132 / 2e-40
Integrase-type DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0081
MBD-like
PF00847
AP2
AP2 domain
Representative CDS sequence
>Potri.008G166100.1 pacid=42806683 polypeptide=Potri.008G166100.1.p locus=Potri.008G166100 ID=Potri.008G166100.1.v4.1 annot-version=v4.1
ATGGAAGAGCCTCATAAAGGGAAGGGAAATGAGGAGAAAGGGAGAGGAGAGGTTCGATACCGGGGTGTTCGAAGGAGACCTTGGGGAAAATTTGCAGCGG
AGATACGAGACCCATCAAGGCAAGGAGGAAGGTTATGGCTAGGGACATTTGATACGGCAGAAGAGGCAGCTCGTGCTTATGATAAAGCTGCATTTAACTT
GAGACGTCATCTTGCAATCCTCAACTTTCCTAGTGAATATATCTCTCAAGTGACGGAGTTTTCTCCTCGTCCTCGATCATTTTCAGCATCTAGCAGTAAT
GTTGGTGCATCTGAGAGTTTTGAGAGGGGAAGCTCATCAAGTACCGGACAAGGAAAGCAAGTTATCGAGTTTGAGTACTTGGATGATAAGATTTTGGAGG
AGCTTCTTGAAACAGAAGAGGAGAAGAAGAAGCGGCGGCAAGATTGA
AA sequence
>Potri.008G166100.1 pacid=42806683 polypeptide=Potri.008G166100.1.p locus=Potri.008G166100 ID=Potri.008G166100.1.v4.1 annot-version=v4.1
MEEPHKGKGNEEKGRGEVRYRGVRRRPWGKFAAEIRDPSRQGGRLWLGTFDTAEEAARAYDKAAFNLRRHLAILNFPSEYISQVTEFSPRPRSFSASSSN
VGASESFERGSSSSTGQGKQVIEFEYLDDKILEELLETEEEKKKRRQD
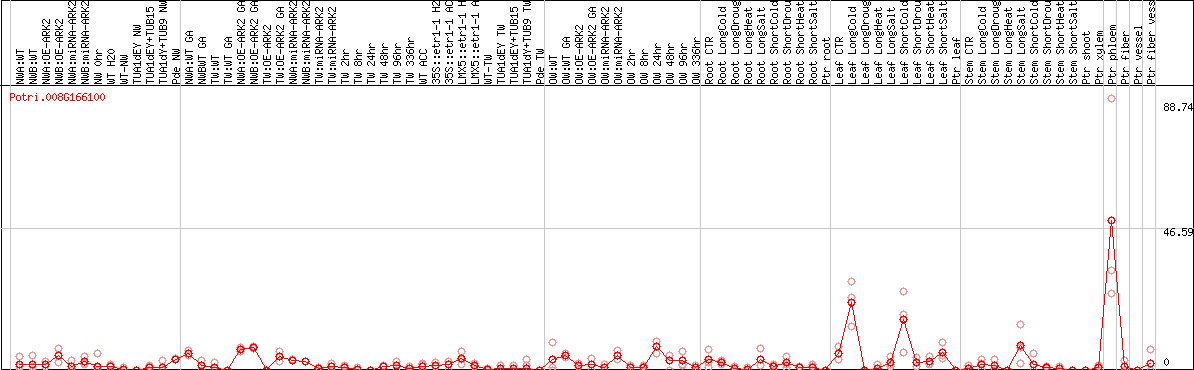
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G166100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.