Potri.008G220200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G220200.2 pacid=42806079 polypeptide=Potri.008G220200.2.p locus=Potri.008G220200 ID=Potri.008G220200.2.v4.1 annot-version=v4.1
ATGGCTGCTTCTTATTCACGTACTACCACTCGGTGGAAATATGATGTCTTCCTCAGTTTCAGAGGTGAAGACACCCGGAAGAGTTTTACTGACCATCTTT
ATACTGCCTTGTGTCACAGAGGAGTCATCACTTTTAGAGATGATCAAGAACTTGAGAGAGGGAATGAAATTTCCCGTGAACTTCTTCAAGCAATACAAGA
TTCAAGGTTTTCAGTGATAGTTTTCTCAAGAAACTATACTTCTTCCACATGGTGCTTGAACGAACTAGTAAAAATTGTGGAATGCATGAAACAAGGTAGA
CAAACAGTTATTCCAGTTTTCTATGATGTTGATCCTTCTGAAGTAAGGAATCAGACAGGGAGACTTCAACAGGCCTTTGCTGATCATGAAGAGGTTTTCA
AAGACAACATTGAGAAGGTGCAGACCTGGAGGATTGCGATGAAACTAGTGGCCAATCTTTCTGGATGGGATTTGCAGGATAGGCACGAGTCAGAGTTTAT
TCAAGGCATTGTTGAAGAGATAGTATGCAAGTTGAGAAAATCAAGTTACAGCATGTCATGGGTAACCGAGAACTTAGTTGGAATGGATTGGCGTTTGGAG
GAAATGAGTTTATATTTAGGAGTAGAGCAGCTGAATGATGTTCGCGTTATAGGGATCTGTGGGATGGGTGGAATAGGTAAGACGACCATTGCTAGAGCTG
TCTATGAGAAGATGCTGGGTCATTTTGAAGGCAGCAGCTTTCTTGCAAATGTTAGAGAAGTTGAGGAGAAACATGGTTTAGTTCGCTTACAGGAACAACT
TCTTTCAGACACGTTGATGGATAGAAGGACCAAAATCTCTGATGTTCATAGAGGAATGAATGAAATAAGGGTTAGGCTGCGTTCTAGAATGGTTCTTGTT
GTTCTTGATGATGTTGATCAATTGGTCCAATTAGAATCTTTAGTTGGAGATCGTAATTGGTTTGACAATGGAAGTAGAGTCATAATAACCACAAGAGATG
AACTTCTTCTGAAACAATTTGGAGTTGATAAAATATATAGAGTTGCAAGTTTAAATAACATTGAAGCCGTTCAACTCTTTTGTCTGAAAGCCTTCAGAAG
TTATTGTCCCCCGGAAGATTATGTGCTTCAGACCATCCAAGTTGTAAAGTATGCAGATGGCCTTCCATTAGCTCTCCATGTTTTGGGTTCATTTTTTTCT
GGGATCAGATCTGTAGAATTGTGGAACCATTCATTAAAAAGACTGAAAGATATCCCAGATAAAGGAATTCTAGATAAACTTAAAATAAGTTTTGATGGAC
TTAACGAAGTAGAGAAGAAAATATTTCTAGATATTGCTTGTTTTTTCAACGGGTGGGAAGAAGATTGTGTGACAAAATTAATGGAAAGTTCTGGTTTCTA
TCCACAGATTGGAATAAGGATTCTCGTAGAAAAATTTCTCATAAATATTTCAGATAACAGAGTATGGATGCATGACTTGCTTCAAGAAATGGGTCGACAA
ATTGTCAAGAGAGAGTCTCATGAAGAACCTGGAAAACGCACTAGGTTGTGGCTCTGTGAGGATGTCATTCACGTTTTGCTAAATAATACGGGAACCGACA
AAGTTGAAGGCATCGTCCTGAATTCAAATGACGAAGTAGATGGACTATACTTAAGTGCTGAATCAATAATGAAGATGAAAAGGTTAAGAATTCTCAAACT
CCAAAATATAAACCTTTCCCAGGAAATTAAATATCTCTCCAATGAGTTGAGATATCTTGAATGGTGCCGATATCCTTTCAAATCATTGCCATCAACTTTT
CAACCGGATAAACTTGTTGAGCTCCACATGCGTCATAGCAGCATTAAACAATTATGGGAGGGAGTGAGACCTTTAAAGTTGCTTAGAGCTATTGATCTCA
GACACTCCCGAAACTTAATAAAGACTCCAGACTTCAGACAAGTTCCAAATCTTGAGAAGCTGAATCTTGAGGGTTGCAGAAAACTGGTCAAGATTGACGA
CTCTATCGGAATTCTGAAAGGGCTTGTTTTCTTGAATTTGAAGGACTGCGTAAAGCTTGCCTGTCTTCCAACAAACATATGTGAGTTGAAGACCCTCAGA
ATTCTTAACTTGTATGGATGCTTCAAACTTGAAAAATTGCCTGAGATGTTGGGAAATGTAATAAATCTGGAGGAGCTTGATGTAGGCAGAACTGCTATAA
CACAACTGCCATCAACCTTTGGTCTCTGGAAAAAACTTAAGGTGCTATCTTTTGATGGATGCAAGGGACCAGCACCTAAATCATGGTATTCTCTCTTCTC
CTTCCGGTCACTTCCAAGAAATCCTTGTCCAATTACTTTGATGCTGTCCTCTCTGTCAACGCTATACTCTTTGACAAAGCTAAATCTAAGCAACTGCAAT
CTCATGGAAGGAGAACTTCCTGATGATATGAGCTGCTTCCCATCATTGGAAGAATTAGACCTTATTGGAAATAACTTTGTGAGAATACCTTCAAGCATCA
GCCGGCTCTCTAAACTTAAAAGCCTCAGGTTAGGCAATTGCAAGAAGCTTCAGTCACTTCCGGATCTTCCATCAAGGCTTGAATATCTCGGGGTGGACGG
TTGCGCTTCGTTGGGAACTCTTCCAAATCTATTTGAAGAATGTGCCCGCTCAAAGTTTTTGTCCTTGATTTTTATGAACTGCTCCGAGTTGACTGATTAT
CAAGGGAATATCAGCATGGGTTTGACATGGCTTAAATATTACCTCCATTTCTTGTTGGAAAGCGGCCATCAGGGTCATCCAGCCTCTTGGTTTTTTACCT
GTTTTCCAGGAAGTGAAATTCCTAGTTGGTTTCACCATAAGAGCGTGGGTCATTCTTTAACAATACGGCTACTTCCTTATGAGCATTGGTCCAGCAGTAA
GTGGATGGGATTAGCTGTTTGTGCATTCTTTGAAGAGCTTGATTGTGGTGATTCATGTCTTATTACATTGAACTTCGATATCAAAGGCTTCAAATCTAGG
AGCTACTTTCTAGAGTACCCTGAAGGTTCAACATTTACGTCAAATCAGGTTTTCTTTATCTTCTTTCCCAGAGGCAAGTTTCCGGAACCACTTGCTGTGT
CTAATACTACTTCTCAGCCCATTGAGGTGGAGTTCCGGTCATCTATTCAAGAAAGAAACACCAACAATGAGTTTCAGGTTCTTTCTGCCCGAGTCATGAA
TTGGGGGTTCCGCATGGTGTACGAGGAAGACACACTGCAGTTCAACGAACCGAATACAGGTGTCGACTCTATCAATGGATTGCATTCTGAAGAGTTGAGG
ATATTGGATGAGAACAACTCCATCATCCCATGCAAAAATATTGAGGGCAAATGCCTCCCAGAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G220200.2 pacid=42806079 polypeptide=Potri.008G220200.2.p locus=Potri.008G220200 ID=Potri.008G220200.2.v4.1 annot-version=v4.1
MAASYSRTTTRWKYDVFLSFRGEDTRKSFTDHLYTALCHRGVITFRDDQELERGNEISRELLQAIQDSRFSVIVFSRNYTSSTWCLNELVKIVECMKQGR
QTVIPVFYDVDPSEVRNQTGRLQQAFADHEEVFKDNIEKVQTWRIAMKLVANLSGWDLQDRHESEFIQGIVEEIVCKLRKSSYSMSWVTENLVGMDWRLE
EMSLYLGVEQLNDVRVIGICGMGGIGKTTIARAVYEKMLGHFEGSSFLANVREVEEKHGLVRLQEQLLSDTLMDRRTKISDVHRGMNEIRVRLRSRMVLV
VLDDVDQLVQLESLVGDRNWFDNGSRVIITTRDELLLKQFGVDKIYRVASLNNIEAVQLFCLKAFRSYCPPEDYVLQTIQVVKYADGLPLALHVLGSFFS
GIRSVELWNHSLKRLKDIPDKGILDKLKISFDGLNEVEKKIFLDIACFFNGWEEDCVTKLMESSGFYPQIGIRILVEKFLINISDNRVWMHDLLQEMGRQ
IVKRESHEEPGKRTRLWLCEDVIHVLLNNTGTDKVEGIVLNSNDEVDGLYLSAESIMKMKRLRILKLQNINLSQEIKYLSNELRYLEWCRYPFKSLPSTF
QPDKLVELHMRHSSIKQLWEGVRPLKLLRAIDLRHSRNLIKTPDFRQVPNLEKLNLEGCRKLVKIDDSIGILKGLVFLNLKDCVKLACLPTNICELKTLR
ILNLYGCFKLEKLPEMLGNVINLEELDVGRTAITQLPSTFGLWKKLKVLSFDGCKGPAPKSWYSLFSFRSLPRNPCPITLMLSSLSTLYSLTKLNLSNCN
LMEGELPDDMSCFPSLEELDLIGNNFVRIPSSISRLSKLKSLRLGNCKKLQSLPDLPSRLEYLGVDGCASLGTLPNLFEECARSKFLSLIFMNCSELTDY
QGNISMGLTWLKYYLHFLLESGHQGHPASWFFTCFPGSEIPSWFHHKSVGHSLTIRLLPYEHWSSSKWMGLAVCAFFEELDCGDSCLITLNFDIKGFKSR
SYFLEYPEGSTFTSNQVFFIFFPRGKFPEPLAVSNTTSQPIEVEFRSSIQERNTNNEFQVLSARVMNWGFRMVYEEDTLQFNEPNTGVDSINGLHSEELR
ILDENNSIIPCKNIEGKCLPE
|
DESeq2's median of ratios [POPLAR]
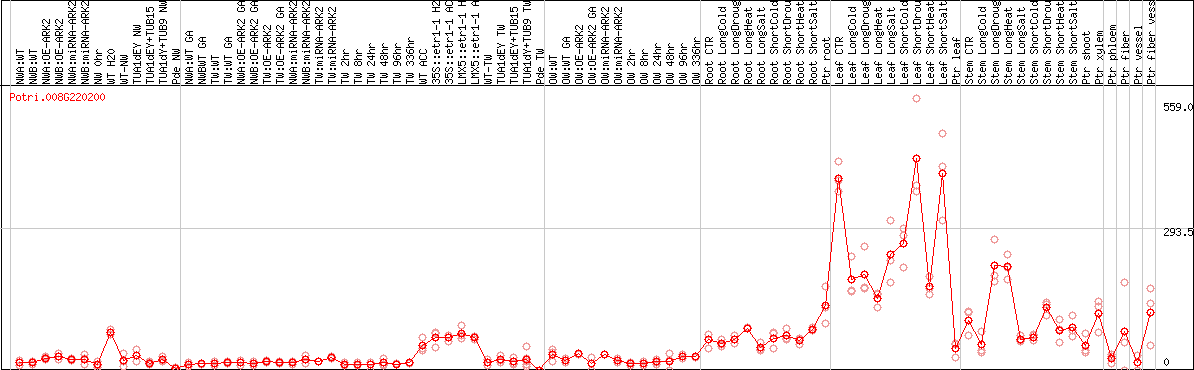
Coexpressed genes
Potri.008G220200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
