External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39250 109 / 1e-32
MYB
RSM2, ATRL1
RADIALIS-LIKE SANT/MYB 2, RAD-like 1 (.1)
AT1G75250 105 / 8e-31
MYB
RSM3, ATRL6
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
AT1G19510 101 / 2e-29
MYB
RSM4, ATRL5
RADIALIS-LIKE SANT/MYB 4, RAD-like 5 (.1)
AT2G21650 96 / 2e-27
MYB
RSM1, ATRL2, MEE3
RADIALIS-LIKE SANT/MYB 1, MATERNAL EFFECT EMBRYO ARREST 3, ARABIDOPSIS RAD-LIKE 2, Homeodomain-like superfamily protein (.1)
AT2G18328 92 / 4e-26
MYB
ATRL4
RAD-like 4 (.1)
AT4G36570 76 / 1e-19
MYB
ATRL3
RAD-like 3 (.1)
AT2G38090 73 / 1e-16
MYB
MYB-R
Duplicated homeodomain-like superfamily protein (.1)
AT3G11280 72 / 3e-16
MYB
Duplicated homeodomain-like superfamily protein (.1.2)
AT5G05790 71 / 4e-16
MYB
Duplicated homeodomain-like superfamily protein (.1)
AT5G04760 71 / 4e-16
MYB
Duplicated homeodomain-like superfamily protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014301
112 / 9e-34
AT1G75250 102 / 5e-30
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
Lus10010831
109 / 1e-32
AT1G75250 108 / 6e-32
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
Lus10026009
109 / 1e-32
AT1G75250 103 / 1e-30
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
Lus10040450
107 / 5e-32
AT1G75250 98 / 5e-28
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
Lus10041752
107 / 6e-32
AT1G75250 103 / 2e-30
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
Lus10028306
106 / 1e-31
AT1G75250 103 / 1e-30
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
Lus10023568
105 / 4e-31
AT4G39250 123 / 5e-38
RADIALIS-LIKE SANT/MYB 2, RAD-like 1 (.1)
Lus10007152
103 / 2e-30
AT1G75250 113 / 3e-34
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
Lus10040453
103 / 3e-30
AT2G21650 115 / 4e-35
RADIALIS-LIKE SANT/MYB 1, MATERNAL EFFECT EMBRYO ARREST 3, ARABIDOPSIS RAD-LIKE 2, Homeodomain-like superfamily protein (.1)
Lus10023564
102 / 5e-30
AT1G75250 101 / 2e-29
RADIALIS-LIKE SANT/MYB 3, RAD-like 6 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Potri.009G117200.2 pacid=42772400 polypeptide=Potri.009G117200.2.p locus=Potri.009G117200 ID=Potri.009G117200.2.v4.1 annot-version=v4.1
ATGTCTTCAAGCTATCAAGCTTCCAGAAATTCTAGATCCGCTTGGACACCAAGGGAAAACAAACTGTTTGAAAAGGCACTAGCCTTGTTTGATAAGGACA
CACCGGACAGGTGGCAAAATATAGCCAAAGCTGTTGGTGGTGTGAAATCTGCCGAGGAAGTGAAGAAACACTATGAAATCCTTATTGAAGATCTTCAACA
TATCGAGTCTGGCCGCATTCCTATTCCTAAGTACAAGTCTAGTGGAAGCTGCAACAACACAAACGAAGAGGAAAGGCTTCTGAAGTATCTCAACCTGCAG
TGA
AA sequence
>Potri.009G117200.2 pacid=42772400 polypeptide=Potri.009G117200.2.p locus=Potri.009G117200 ID=Potri.009G117200.2.v4.1 annot-version=v4.1
MSSSYQASRNSRSAWTPRENKLFEKALALFDKDTPDRWQNIAKAVGGVKSAEEVKKHYEILIEDLQHIESGRIPIPKYKSSGSCNNTNEEERLLKYLNLQ
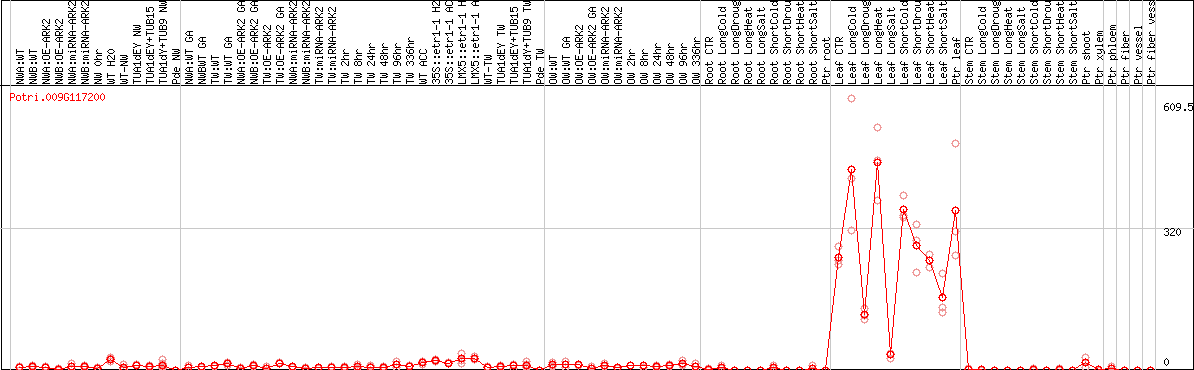
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G117200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.