Potri.011G012475 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G012475.1 pacid=42781726 polypeptide=Potri.011G012475.1.p locus=Potri.011G012475 ID=Potri.011G012475.1.v4.1 annot-version=v4.1
ATGGCTGCTGGGAAGTATCAAGAATCCTACTCTTCACGGTTTCCTAATTGTAAATATCAAGTGTTCTTGAGTTTTAGAGGTGAAGACACCCGCAAGAACT
TTACCGATCACCTCTACACGGCCCTGGTTCAAGCAGGGATTCACACATTTAGAGATGATAATGAAATTCGGAGAGGAGAGAATATAGATTTCGAGCTCCA
GAAGGCAATACAACAATCAAAAATATCGATAATCGTGTTCTCCAAAGACTATGCCTCGTCGAGATGGTGCCTGGATGAACTTGTAATGATCATGGAACGT
AAAAGGAATCCTGATCACTGCATAGTTTTGCCAGTATTCTATGATGTGGATCCATCCCAAGTCGGACGTCAAACAGGGAGCTTCTCTGCAGCATTTGTGG
AACATGAAAAGAGCTTCAACGAGGAGATGGAGCGGGTGAATGGGTGGAGGATTGCTTTGAAGGAAGTTGCAGATTTAGCTGGAATGGTTCTAGGAGATGG
GTACGAGGCACCGTTTGTCCAATCTATTGTGGAGAAGGTCTTAAAGAATTTGGATCAAAAAATGTTTCATGTACCCCCTCATTTCATTGGAAGAGATCCT
CTAGTACAAGATATCAACTCATGGTTGCAAGATGGATCCCATGCTGCTGCCATTGCTTTACTCTATGGAATTGGTGGAGTTGGAAAGACAGCCATAGCTA
AGAGTGTTTATAACCAAAACTCTTATAAATTTGAAGGAAAGAGCTTCCTGTCAAATTTTAGGTCAAAGGATATAGTTTGCCTACAGAGACAACTTCTTTC
CGACATCCTAAAAAAGACTGTTGATGAGATAAATGATGAAGATGAAGGAATTCTGAAGATTAAGGATGCCTTATGTTGCAGAAGAACTCTTATTGTTCTA
GATGATGTGGACAAAAGGGACCAATTCAATAAAATCGTTGGCATGCAAAATTGGCTTTGTAAAGGAAGTAAAATAATTGTAACAACCAGAAATAAGGGTC
TGTTTTCAGCTAATGATATTGAGTGGGTCCGGTGCGAAGTCGAACCGCTAGATATTGAAAAATCTTTTGAGCTTTTCAGTTGGAATGTCTTTGGACAACC
TGACCCTGTTGATGGTTTTGTAGAAGACTCTTGGAGAATAGTACATCATTGTAGTGGACTTCCATTAGCTCTTCGAGTTATTGGCTCCTCATTGTCCGGA
AAAGGAAGAGAAATATGGGAAAGCGCATTACAACAATTGGAAGTGATTCCTAATTTTGAAGTTCTAAAGGTTCTTCGAATAAGTTACGACTTTCTTGATG
GTGATTGTCTGAAGAACTTATTCCTTGATATCGCATGTTTCTTCAATGGAATGGATGTGGATGATGCAGTTAGGATACTGGATGGGCTCGCTAAAGGTGC
AAGATTTGGGATTGACAATCTCATCGATAAATGTCTTGCTGAAATCAACATTGATCAAAGGTTGTGGATGCATCAACTAGTAAGAGCTATGGGAAGGGAA
ATTGCTCGTCAAGAATCTGCCAAATGTCAAAGAATATGGCGTCACGAGGATGCTTTTACCGTTTTGAAAGGAACTACTGATGCTGAAAATTTGTGTGGCC
TTACCATTGATATGCATGCATTAATGGAAGATAATTATGCAGAAGTTGTCTGTACTGATTCAATGGTTCTCCGCAGTCTTAACTTCTTTCAACAATGGCT
TTCCAATTTTTCCATTGAGGGAAAATTACAAACTAGCCAAACAAGTTTGTTTCCCATCCTCAGCACGGATGCTTTTAGAAAGATGGCAGATGTAAAATTT
CTCCAGCTAAACTACACTAATTTTTGTGGAAGTTTTGAGCACTTTCCCAAGAAGTTGATATGGTTATGTTGGCATGGATTGTCTTTGAAATCCATACCAA
ATCACGTATGCTTGGAGAAGCTGGTGGTTCTTGATCTATCCAGAAGCTGTCTGGTTGATGCTTGGAAAGGCAAATTGTTTCTTCCAAAATTGAAAATTCT
TGATCTCCGTCACTCTCGTGATCTCATTAGAACCCCAGACTTCTCGGGTCTCCCATCCCTTGAAAAGCTAATACTTGAGGACTGCATCCGTTTGGTTCAA
ATTCACGAATCTATTGGTGATTTACAAAGATTGTTGATCTTAAATCTAAGAAATTGTACAAGTCTTGTGGAGCTTCCAGAAGAAATGAGTAGACTGAATT
CACTTCAAGAGCTGGTTTTAGATGGTTGTTCAAATCTTAACAACCTGAATATGGAGTTAGAGCATGATCAGGGGCGCAAGTTGCTTCAAAGTGATGGAAT
TGTTGCAAGTACATCATTCATTTCATCTCTTCCATTGAAGCTATTCTTTCCCTCTAGGTTTTCAACGAGGAAAATGTTGAGATTTACCTTGTTTTCACTG
CCACGCTTCTTGGAGAGTCTAGATTTAAGTGGAACTCCAATTCGTTTTCTTCCGGAAAGCATCAAGGATCTTGGTCTACTCAGAGCCCTATATTTAAGAA
ATTGCAAAATGCTTCAGGCACTCCCAGAGCTTCCATTCCTTTTGGATTTGTTAGATGTGTCCTTGTGCTATTCACTGCAAGGACTTGCAAATCCAAATAG
TTGGACTGAAGGAGATGGTTGCGATCACTTAGTCGAGTTCCAAGATCGGATAAAGCAAGAATTAATCCAAAAGTTAGACTCTCAAATGTTCAGAATAATG
GAAACGGTTAGTGCTCAAATACAGCCATCGAGATTTCAGATATTATTTGGTCATGGCATATTCAACCTTGTCGTATCTGTATTTGATGATGAAGAGAAGT
CGAGGTGGTTTCATGGGGAGGAAGAAGAGGATAAATGGCTAATTCAGAATGAGTTTGTAGATTACTTTTCATTCAAAATATCCTCACCTCCTCCTGCGCA
CCGGATATGTGGCTTTAATTTATTCACAGGTTGTTGTGTGAGTAAATTATTGAAATCAGAGTACACTTTCTATGATGAATTTTTTATTGAAATCAGAAAC
AATACCAGTGGTCGATCCTTGATTTGTCAAGCCAAAATCTTCCCTGCGAGTTACGCGCGTGGTCTTCGTGAAATCCAATTGCTAAGCCACTGGAAATTAG
GGGTCGATGATCCTACATTTGATAATGGTGATGACGTGAGTATTTCAGTGCGTCCACAGGGTCCAGCTATTCAAATAAGGACGGTTGGTGTACAATGGTT
GCATGAAGAGGAAGGAGATGATGATGATATCCAATCAAAGGATGAAGTTATCAATGCCCACAACAGTAGCGATGATGATGATGCACACGTAGCCAAAGTA
CAAATAGCTTCTCATATTTTTAGAAATTATTATTGTGCTTTCCGTACCAATGGCGGTTTTGACAGGTCGTATTATTTTAGGCTTCGTGATGCCTCGGTTT
CTTTACTGCAAAGTAGATTTTATTTTAAAAAATTAATTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G012475.1 pacid=42781726 polypeptide=Potri.011G012475.1.p locus=Potri.011G012475 ID=Potri.011G012475.1.v4.1 annot-version=v4.1
MAAGKYQESYSSRFPNCKYQVFLSFRGEDTRKNFTDHLYTALVQAGIHTFRDDNEIRRGENIDFELQKAIQQSKISIIVFSKDYASSRWCLDELVMIMER
KRNPDHCIVLPVFYDVDPSQVGRQTGSFSAAFVEHEKSFNEEMERVNGWRIALKEVADLAGMVLGDGYEAPFVQSIVEKVLKNLDQKMFHVPPHFIGRDP
LVQDINSWLQDGSHAAAIALLYGIGGVGKTAIAKSVYNQNSYKFEGKSFLSNFRSKDIVCLQRQLLSDILKKTVDEINDEDEGILKIKDALCCRRTLIVL
DDVDKRDQFNKIVGMQNWLCKGSKIIVTTRNKGLFSANDIEWVRCEVEPLDIEKSFELFSWNVFGQPDPVDGFVEDSWRIVHHCSGLPLALRVIGSSLSG
KGREIWESALQQLEVIPNFEVLKVLRISYDFLDGDCLKNLFLDIACFFNGMDVDDAVRILDGLAKGARFGIDNLIDKCLAEINIDQRLWMHQLVRAMGRE
IARQESAKCQRIWRHEDAFTVLKGTTDAENLCGLTIDMHALMEDNYAEVVCTDSMVLRSLNFFQQWLSNFSIEGKLQTSQTSLFPILSTDAFRKMADVKF
LQLNYTNFCGSFEHFPKKLIWLCWHGLSLKSIPNHVCLEKLVVLDLSRSCLVDAWKGKLFLPKLKILDLRHSRDLIRTPDFSGLPSLEKLILEDCIRLVQ
IHESIGDLQRLLILNLRNCTSLVELPEEMSRLNSLQELVLDGCSNLNNLNMELEHDQGRKLLQSDGIVASTSFISSLPLKLFFPSRFSTRKMLRFTLFSL
PRFLESLDLSGTPIRFLPESIKDLGLLRALYLRNCKMLQALPELPFLLDLLDVSLCYSLQGLANPNSWTEGDGCDHLVEFQDRIKQELIQKLDSQMFRIM
ETVSAQIQPSRFQILFGHGIFNLVVSVFDDEEKSRWFHGEEEEDKWLIQNEFVDYFSFKISSPPPAHRICGFNLFTGCCVSKLLKSEYTFYDEFFIEIRN
NTSGRSLICQAKIFPASYARGLREIQLLSHWKLGVDDPTFDNGDDVSISVRPQGPAIQIRTVGVQWLHEEEGDDDDIQSKDEVINAHNSSDDDDAHVAKV
QIASHIFRNYYCAFRTNGGFDRSYYFRLRDASVSLLQSRFYFKKLI
|
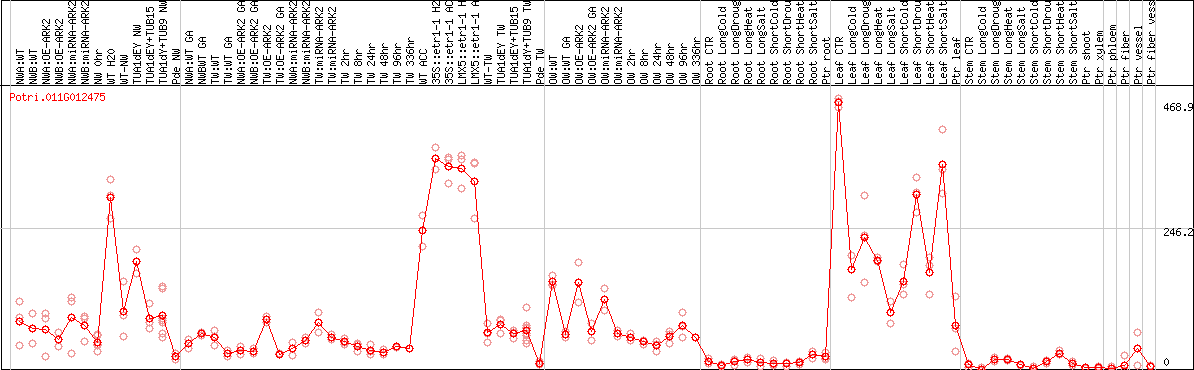
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G012475 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
