External link
Symbol
ERF44
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G28360 149 / 5e-46
AP2_ERF
AtERF12
ERF domain protein 12 (.1)
AT5G44210 110 / 1e-30
AP2_ERF
AtERF9, ATERF-9
ERF DOMAIN PROTEIN- 9, erf domain protein 9 (.1)
AT3G15210 109 / 4e-30
AP2_ERF
ATERF4, RAP2.5, ATERF-4
RELATED TO AP2 5, ethylene responsive element binding factor 4 (.1)
AT1G28370 104 / 8e-29
AP2_ERF
AtERF11
ERF domain protein 11 (.1)
AT1G50640 105 / 1e-28
AP2_ERF
ATERF3
ethylene responsive element binding factor 3 (.1)
AT3G20310 101 / 1e-26
AP2_ERF
ATERF7, ATERF-7
ethylene response factor 7 (.1)
AT1G12980 100 / 1e-25
AP2_ERF
DRN, ESR1
ENHANCER OF SHOOT REGENERATION 1, DORNROSCHEN, Integrase-type DNA-binding superfamily protein (.1)
AT1G03800 99 / 2e-25
AP2_ERF
AtERF10
ARABIDOPSIS THALIANA RF DOMAIN PROTEIN 10, ERF domain protein 10 (.1)
AT2G33710 98 / 2e-25
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1.2)
AT1G24590 99 / 5e-25
AP2_ERF
ESR2, DRNL, SOB2, DRN-LIKE
FOR SUPPRESSOR OF PHYTOCHROMEB-4 2, ENHANCER OF SHOOT REGENERATION 2, DORNROSCHEN-like (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033664
112 / 3e-31
AT3G15210 119 / 2e-33
RELATED TO AP2 5, ethylene responsive element binding factor 4 (.1)
Lus10039857
107 / 4e-29
AT5G44210 138 / 8e-41
ERF DOMAIN PROTEIN- 9, erf domain protein 9 (.1)
Lus10031526
102 / 5e-27
AT1G50640 112 / 2e-30
ethylene responsive element binding factor 3 (.1)
Lus10014345
98 / 2e-25
AT1G12980 97 / 1e-24
ENHANCER OF SHOOT REGENERATION 1, DORNROSCHEN, Integrase-type DNA-binding superfamily protein (.1)
Lus10035129
95 / 4e-24
AT5G13910 152 / 7e-46
LEAFY PETIOLE, Integrase-type DNA-binding superfamily protein (.1)
Lus10016827
95 / 7e-24
AT3G16770 183 / 4e-57
RELATED TO AP2 3, ETHYLENE RESPONSE FACTOR 72, ethylene-responsive element binding protein (.1)
Lus10026053
93 / 1e-23
AT5G18560 97 / 7e-24
Integrase-type DNA-binding superfamily protein (.1)
Lus10018623
91 / 1e-23
AT3G15210 104 / 9e-29
RELATED TO AP2 5, ethylene responsive element binding factor 4 (.1)
Lus10037448
96 / 2e-23
AT1G53910 244 / 4e-78
related to AP2 12 (.1.2.3)
Lus10017907
94 / 2e-23
AT1G24590 134 / 1e-37
FOR SUPPRESSOR OF PHYTOCHROMEB-4 2, ENHANCER OF SHOOT REGENERATION 2, DORNROSCHEN-like (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0081
MBD-like
PF00847
AP2
AP2 domain
Representative CDS sequence
>Potri.011G056900.1 pacid=42782401 polypeptide=Potri.011G056900.1.p locus=Potri.011G056900 ID=Potri.011G056900.1.v4.1 annot-version=v4.1
ATGGCTTCTTCGAGGGAAGGACACTACAGGGGGGTTCGAAAGAGACCATGGGGTCGCTATGCTGCAGAGATACGTGACCCATGGAAGAAAACACGAGTTT
GGTTAGGCACATTTGACACACCTGAAGAAGCTGCGCTTGCTTATGATGGTGCAGCTCGTTCTCTTCGTGGAGCTAAAGCAAAGACTAACTTCCCATCACC
TCCCTCCACTTCTGGTCTCTCTTTTGACCTCAATCTTCCATCGGATCCTCACCACCACCATCTTCATTGGAGTTCATGTCCTCATTTGGGGTCTCACAGG
TTTGGTGGTTTTGGTGAGTTCTTGCAGACTGGAGTAGTTTTTAATGAAATGAACCTCCATGCTACCGAGGCTGCAGCAGCTTCTGGGTCAGTAGCGAAGA
ATGAGGGTCCTGGTGCTGTTGCTGGAACTCCGGCGCCGGAAAATGTTGCTCCAGTTTCATTTTTGGGGATGGTGCGACGTGGGCTGCCAATAGATTTGAA
CGAGCCCCCGCCTTTGTGGCTGTGA
AA sequence
>Potri.011G056900.1 pacid=42782401 polypeptide=Potri.011G056900.1.p locus=Potri.011G056900 ID=Potri.011G056900.1.v4.1 annot-version=v4.1
MASSREGHYRGVRKRPWGRYAAEIRDPWKKTRVWLGTFDTPEEAALAYDGAARSLRGAKAKTNFPSPPSTSGLSFDLNLPSDPHHHHLHWSSCPHLGSHR
FGGFGEFLQTGVVFNEMNLHATEAAAASGSVAKNEGPGAVAGTPAPENVAPVSFLGMVRRGLPIDLNEPPPLWL
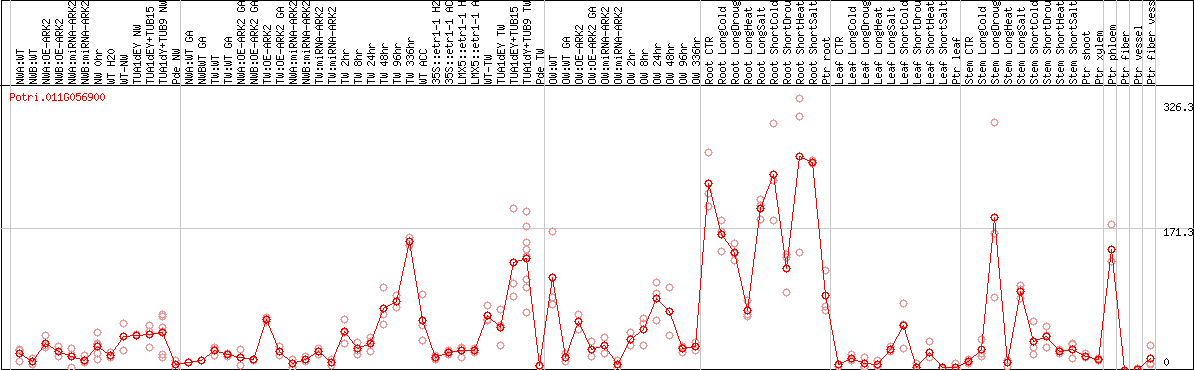
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G056900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.