External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G27680 322 / 4e-109
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G53540 308 / 1e-103
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G50140 133 / 1e-34
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
AT1G64110 130 / 1e-33
DAA1
DUO1-activated ATPase 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
AT3G19740 128 / 6e-33
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G02890 128 / 9e-33
AAA-type ATPase family protein (.1.2)
AT4G02480 127 / 1e-32
AAA-type ATPase family protein (.1)
AT4G28000 123 / 3e-31
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G52882 122 / 4e-31
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT4G24860 121 / 2e-30
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G007700
359 / 9e-124
AT4G27680 615 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.012G027901
137 / 1e-40
AT5G53540 182 / 2e-57
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.012G027101
130 / 4e-38
AT4G27680 166 / 3e-51
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.012G017760
127 / 1e-36
AT4G27680 164 / 1e-50
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.007G069800
132 / 2e-34
AT1G50140 1215 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
Potri.001G097600
129 / 4e-33
AT1G64110 1159 / 0.0
DUO1-activated ATPase 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Potri.012G096300
128 / 5e-33
AT4G02480 1153 / 0.0
AAA-type ATPase family protein (.1)
Potri.003G133700
128 / 7e-33
AT1G64110 1145 / 0.0
DUO1-activated ATPase 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Potri.015G094100
127 / 2e-32
AT4G02480 1128 / 0.0
AAA-type ATPase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008835
329 / 1e-111
AT4G27680 649 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10022361
316 / 3e-106
AT4G27680 632 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10024692
134 / 6e-35
AT1G64110 1176 / 0.0
DUO1-activated ATPase 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Lus10010204
134 / 1e-34
AT3G19740 1242 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10028752
132 / 3e-34
AT1G64110 1165 / 0.0
DUO1-activated ATPase 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Lus10017535
131 / 8e-34
AT1G64110 1157 / 0.0
DUO1-activated ATPase 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Lus10040976
130 / 1e-33
AT3G19740 472 / 2e-154
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10027551
126 / 3e-32
AT5G52882 1136 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10039309
126 / 3e-32
AT5G52882 1140 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10017405
125 / 1e-31
AT3G19740 1223 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00004
AAA
ATPase family associated with various cellular activities (AAA)
Representative CDS sequence
>Potri.012G025351.1 pacid=42783442 polypeptide=Potri.012G025351.1.p locus=Potri.012G025351 ID=Potri.012G025351.1.v4.1 annot-version=v4.1
ATGAATCAAATGATGCGGTGGAATCACGGTTACAATTGCCCGAGCCAGCCGCACGTGTCGCCAAAAGGTGGTGGAGCGACTAACTTAGACCAAGTTTTTG
CAAGAACCAGTGCAGCTTTAACTTGTTTGGTATTATTTGCTAGGCTCCGGAAACCTGACCCGAATCATGAAGCATCAAAGAAAGCCCTTGAACATAAGAA
GGAAATCGATAAGCGTTTGGGTCATTCCCTTATCCAAACAAACCCTTATGAGGGATGTATGTGTGTCAAATCAATTGGAGGATTGGAATCTATCAAGCAA
GCTTTGTATGAATTGGTGATTCTTCCTCTACGAAAACCTGAGCTCTTCTCACATGGGAAACTTCTTGGCACCGGGAAGACCATGCTTGCAAAAGCTATTG
TCAGAGAGTCTGGAGTTATTTTTATAAATACGAGGATCTCCAATCTGATAAGCAAGTGGTTCGATGATGCACAAAAGCTCGGTGATGCTGTTGTTTTTAG
CCTGGCTTATAAACTCCAGCCTGCTTTTATATTAATTGACGAGGTTGACAGTTTTCTGGGGCAGCGTCGTACTACAGACCATGAGGCATTAACCAATATG
AAGACTAAGTTCATGGCATTATGGAATGGGTTTACTATAGATCAGAATGCACAAGAGATGAAAATCGAAAGCAAAATTGACTTTGACTACATAGCAAGTT
TGTGTGAGAGCTATACTAGTTCAGATTTTCTTGAACCTTGCAAGAAAGCACCAAGGCCATTGTCCCAGTCAGACTTGCAGAGAGTTCTTGGCACATCAAC
TAAGACCAGGGTTGCTGCAAATGAGTACAGTAGTTCAAGCTCACATTCACCTAGGTGGCCAGTATCTCAAATTTTGAATCTTCAGTCGGATAATCAGGAC
CCTTGA
AA sequence
>Potri.012G025351.1 pacid=42783442 polypeptide=Potri.012G025351.1.p locus=Potri.012G025351 ID=Potri.012G025351.1.v4.1 annot-version=v4.1
MNQMMRWNHGYNCPSQPHVSPKGGGATNLDQVFARTSAALTCLVLFARLRKPDPNHEASKKALEHKKEIDKRLGHSLIQTNPYEGCMCVKSIGGLESIKQ
ALYELVILPLRKPELFSHGKLLGTGKTMLAKAIVRESGVIFINTRISNLISKWFDDAQKLGDAVVFSLAYKLQPAFILIDEVDSFLGQRRTTDHEALTNM
KTKFMALWNGFTIDQNAQEMKIESKIDFDYIASLCESYTSSDFLEPCKKAPRPLSQSDLQRVLGTSTKTRVAANEYSSSSSHSPRWPVSQILNLQSDNQD
P
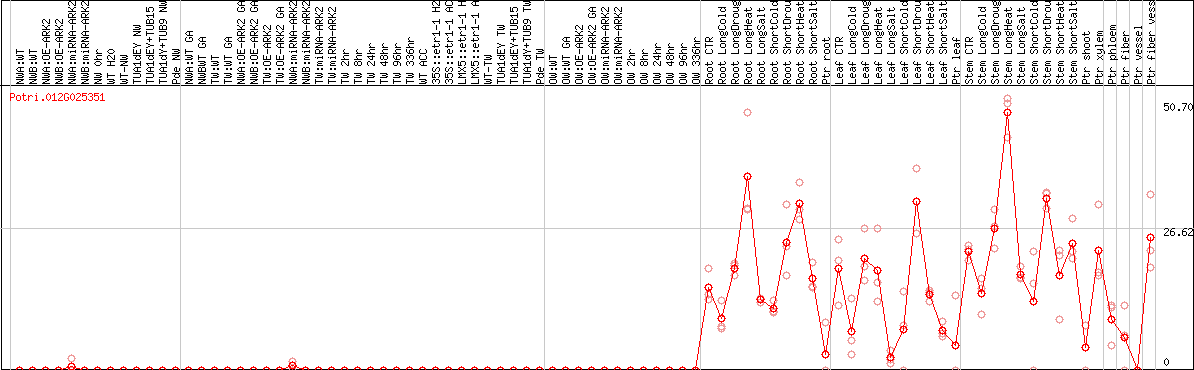
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G025351 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.